Abstract
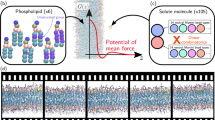
To assist in understanding the mechanism of membrane permeation, the movement of four different molecules within hydrated lipid bilayer membranes were studied via over 15 nanoseconds of atomic-level molecular dynamics simulation. In particular, the simulations were used to explain the anomolously high rate of permeation seen for small molecules. These simulations support the hypothesis that the rate of diffusion of small solutes is enhanced because they can move rapidly within and jump between spontaneously arising voids. The enhanced diffusion rate is greatest in the bilayer center where the voids are most frequently found and of the largest size. Molecules the volume of benzene or smaller experience this enhanced movement, however larger molecules, those the size of adamantane or larger, do not. The details of the diffusional mechanisms of these molecules are discussed. The role of hydrogen bonding for the interactions between drugs (a nifedipine analog) and membranes is discussed.
Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only
Preview
Unable to display preview. Download preview PDF.
Similar content being viewed by others
References
Alper HE, Stouch TR (1995): Orientation and diffusion of a drug analog in biomembranes: Molecular dynamics simulations. J Phys Chem 99:5724–5731
Alper HE, Bassolino DA, Stouch TR (1993a): Computer simulations of a phospholipid monolayer/water system: The effect of long range forces on water structure and dynamics. J Chem Phys 98:9798–9807
Alper HE, Bassolino D, Stouch TR (1993b): The limiting behavior of water hydrating a phospholipid monolayer: A computer simulation study. J Chem Phys 99:5547–5559
Anel A, Richieri GV, Kleinfeld AM (1993): Membrane partition of fatty acids and inhibition of T-cell function. Biochemistry 32:530–536
Bassolino D, Alper HE, Stouch TR (1995): Mechanism of solute diffusion through lipid bilayer membranes by molecular dynamics simulation. J Amer Chem Soc 117:4118–4129
Bassolino D, Alper HE, Stouch TR (1993): Solute diffusion in lipid bilayer membranes: An anatomic level study by molecular dynamics stimulation. Biochemistry 32:12624–12637
Bauerle H-D, Seelig J (1991): Interaction of charged and uncharged calcium channel antagonists with phospholipid membranes. Binding equilibrium, binding enthalpy and membrane location. Biochemistry 30:7023
Beschiaschvili G, Seelig J (1992): Peptide binding to lipid bilayers. Nonclassical hydrophobic effect and membrane-induced pK shifts. Biochemistry 31:10044–10053
Damodaran KV, Merz KM Jr, Gaber BP (1992): Structure and dynamics of the dilauroyl-phosphatidylethanolamine (DLPE) lipid bilayer. Biochemistry 31:1–20
De Loof H, Harvey SC, Segrest JP, Pastor RW (1991): Mean field stochastic boundary molecular dynamics simulation of a phospholipid in a membrane. Biochemistry 30:2099–2113
DeYoung LR, Dill KA (1988): Solute partitioning into lipid bilayer membranes. Biochemistry 27:5281–52S9
Diamond JM, Katz Y (1974): Interpretation of non-electrolyte partition coefficients between dimyristoyl lecithin and water. J Membr Biol 17:121–154
Dix JA, Diamond JM, Kivelson D (1974): Translational difusión coefficient and partition coefficient of a spin-labeled solute in lecithin bilayer membranes. Proc Nat Acad Sei USA 71:474–478
Dix JA, Kivelson D, Diamond JM (1978): Molecular motions of small molecules in lecithin bilayers. J Membr Biol 40:315–342
Edholm, O. and Johansson, J. (1987): Lipid bilayer polypeptide interactions studied by molecular dynamics simulation. Eur Biophys J 14:203–209
Egberts E, Berendsen HJC (1988b): Molecular dynamics simulation of a smectic liquid crystal with atomic detail. J Chem Phys 89:3718–3732
Franks NP (1976): Structural analysis of hydrated egg lecithin and cholesterol bilayers. J Mol Biol 100:345–358
Franks NP, Lieb WR (1979): The structure of lipid bilayers and the effects of general anesthetics. J Mol Biol 133:469–500
Gawrisch K, Ruston D, Zimmerberg J, Parsegian VA, Rand RP, Fuller N (1992): Membrane dipole potentials, hydration forces, and ordering of water at membrane surfaces. Biophys J 61:1213–1223
Gennis RB (1989): Biomembranes, molecular structure and function. In: Springer Advanced Texts in Chemistry Cantor CR, ed. New York: Springer-Verlag
Jacobs RE, White SH (1984): Behavior of hexane dissolved in DMPC bilayers: An NMR and calorimetric study. J Am Chem Soc 106:915–920
Jonsson B, Edholm O, Teleman O (1986): Molecular dynamics simulations of a sodium octanoate micelle in aqueous solution. J Chem Phys 85:22663–2271
Jorgensen K, Ipsen JH, Mouritsen OG, Bennett D, Zuckermann MJ (1991a): The effects of density fluctuations on the partitioning of foreign molecules into lipid bilayers: Application to anasthetics and insecticides. Biophys Acta 1067:241–253
Jorgensen K, Ipsen JH, Mouritsen OG, Bennett D, Zuckermann MJ (1991b): A general model for the interaction of foreign molecules with lipid membranes: Drug and anaesthetics. Biophys Acta 1062:227–238
Kamp F, Hamilton JA (1993): Movement of fatty acids, fatty acid analogues and bile acids across phospholipid bilayers. Biochemistry 32:1074–11086
Lieb WR (1986): J Membr Biol 92:111–119
Lieb WR, Stein WD (1971): The molecular basis of simple diffusion within biological membranes. Curr Top Membr Transp 2:1–39
Lieb WR, Stein WD (1969): Biological membranes behave as non-porous polymer sheets with respect to the diffusion of non-electrolytes. Nature 224:240–243
Marqussee JA, Dill KA (1986): Solute Partitioning into chain molecule interphases: Monolayers, bilayer membranes, and micelles. J Chem Phys 85:434–444
Marrink S-J, Berendsen HJC (1994): Simulation of water transport through a lipid membrane. J Phys Chem 98:4155
Mason PR, Moring J, Herbette LG (1990): A molecular model involving the membrane bilayer in the binding of lipid soluble drugs to their receptors in the heart and brain. Nucl Med Biol 17:13
Mason RP, Rhodes DG, Herbette LG (1991): Reevaluating equalibrium and kinetic binding parameters for the lipophilic drugs based on a structural model for drug interaction with biological membranes. J Med Chem 34:869–877
Miller DM (1986): Biochim Biophys Acta 856:27–35
Miiller-Plathe F, Rogers SC, van Gunsteren WF (1993): Gas sorption and transport in polyisobutylene. J Chem Phys 98:9895–9904
Nelander JC, Blaurock AE (1978): Disorder in nerve myelin: Phasing the higher order reflections by means of the diffuse scatter. J Mol Biol 118:497–532
Nicholas JB, Trouw FR, Mertz JE, Iton LE, Hopfinger AJ (1993): MD simulations of propane and methane in silicate. J Phys Chem 97:4149–4163
Overton E (1899): Vierteljahrsschr Naturforsch Ges Zuerich 44:88–135
Pastor R, Venable RM, Karplus M (1991): Model for the structure of the lipid bilayer. Proc Nat Acad Sci USA 88:892–896
Scott HL, Kalaskar S (1989): Lipid chains and cholesterol in model membranes: A Monte Carlo study. Biochemistry 28:3687–3691
Seelig A, Seelig J (1974): The dynamics structure of phopholipids measured by deuterium NMR. Biochemistry 13:4839
Stouch TR (1993): Lipid membrane structure and dynamics studied by all-atom molecular dynamics simulations of hydrated phospholipid bilayers. Mol Sim 10:317–345
Stouch TR, Alper HE, Bassolino D (1994): Supercomputing studies of biomembranes Internat J Supercomp App 8:6–23
Stouch TR, Ward KB, Altieri A, Hagler AT (1991): Simulations of lipid crystals: Characterization of the potential energy functions and parameters for lecithin molecules. J Comp Chem 12:1033–1046
Takeuchi H, Okazaki K (1990): Molecular dynamics simulation of diffusion of simple gas molecules in a short chain polymer. J Chem Phys 92:5643–5652
Trauble H (1971): The movement of molecules across lipid membranes: A molecular theory. J Membr Biol 4:193–208
Turner GL, Oldfield E (1979): Effect of a local anesthetic on hydrocarbon chain order in membranes. Nature 277:669–670
Van der Ploeg P, Berendsen HJC (1983): Molecular dynamics of a bilayer membrane. Mol Phys 49:233–248
Walter A, Gutknecht J (1986): Permeability of small molecules though lipid bilayer membranes. J Membr Biol 90:207–217
Watanabe K, Ferrado M, Klein M (1988): J Chem Phys 92:818–821
Williams DE, Stouch TR (1993): Characterization of force fields for lipid molecules: Applications to crystal structures. J Comp Chem 14:1066–1076
Xiang T, Anderson B (1994): Molecular distributions in interphase: Statistical mechanical theory combined with molecular dynamics simulation of a model lipid bilayer. Biophys J 66:561
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 1996 Birkhäuser Boston
About this chapter
Cite this chapter
Stouch, T.R., Bassolino, D. (1996). Movement of Small Molecules in Lipid Bilayers: Molecular Dynamics Simulation Studies. In: Merz, K.M., Roux, B. (eds) Biological Membranes. Birkhäuser Boston. https://doi.org/10.1007/978-1-4684-8580-6_8
Download citation
DOI: https://doi.org/10.1007/978-1-4684-8580-6_8
Publisher Name: Birkhäuser Boston
Print ISBN: 978-1-4684-8582-0
Online ISBN: 978-1-4684-8580-6
eBook Packages: Springer Book Archive