Abstract
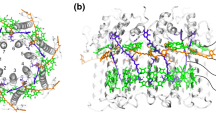
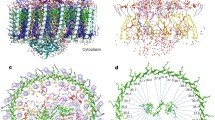
We illustrate in this chapter how one proceeds to predict the structure of integral membrane proteins when a highly homologous structure is unknown. We focus here on the prediction of the structure of the light-harvesting complex II (LH-II) of Rhodospirillum molischianum, an integral membrane protein of 16 polypeptides aggregating and binding to 24 bacteriochlorophyll a’s and 12 lycopenes. Hydropathy analysis was performed to identify the putative transmembrane segments, which were independently verified by multiple sequence alignment propensity analyses and homology modeling. A consensus assignment for secondary structure was derived from a combination of all the prediction methods used. Transmembrane helices were built by comparative modeling. The resulting tertiary structures were then aggregated into a quaternary structure through molecular dynamics simulations and energy minimization under constraints provided by site directed mutagenesis and FT Resonance Raman spectra, as well as conservation of residues. The structure of LH-II, so determined, was an octamer of αβ heterodimers forming a ring with a diameter of 70 Å. We discuss how the resulting structure may be used to solve the phase problem in X-ray crystallography in a procedure called molecular replacement. We will also discuss the exciton structure which results from the circular arrangement of chlorophyls in LH-II.
Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only
Preview
Unable to display preview. Download preview PDF.
Similar content being viewed by others
References
Allen J, Yeates T, Komiya H, Rees D (1987): Structure of the reaction center from Rhodobacter sphaeroides R-26: The protein subunits. Proc Natl Acad Sci USA 84:6162
Altschul SF, Gish W, Miller W, Myers EW, Lipman DJ (1990): Basic local alignment search tool. J Mol Biol 215:403–410
Argos P, Rao J, Hargrave P (1982): Structural prediction of membrane-bound proteins. European Journal of Biochemistry 128:565
Arkin I, Adams P, MacKenzie K, Lemmon M, Brünger A, Engelman D (1994): Structural organization of the pentameric transmembrane α-helices of phospholamban, a cardiac ion channel. Embo Journal 13:4757
Blundell TL, Sibanda BL, Sternberg MJ, Thornton JM (1987): Knowledge-based prediction of protein structures and the design of novel molecules. Nature 326:347
Boonstra AF, Visschers RW, Calkoen F, van Grondelle R, van Bruggen EF, Roekema EJ (1993): Structural characterization of the B800–850 and B875 light-harvesting antenna complexes from Rhodobacter-Sphaeroides by electron microscopy. Biochimica et Biophysica Acta 1142:181
Brooks BR, Bruccoleri RE, Olafson BD, States DJ, Swaminathan S, Karplus M (1983): CHARMm: a program for macromolecular energy, minimization, and dynamics calculations.J Comp Chem 4(2): 187–217
Brünger AT (1990): Extension of molecular replacement: A new search strategy based on Patterson correlation refinement. Acta Cryst A46:46–57
Briinger AT (1992): X-PLOR, Version 3.1 f A System for X-ray Crystallography and NMR. The Howard Hughes Medical Institute and Department of Molecular Biophysics and Biochemistry, Yale University
Brunisholz RA, Wiemken V, Suter F, Bachofen R, Zuber H (1984): The light-harvesting polypeptides of Rhodospirillum rubrum. IL localisation of the amino-terminal regions of the light-harvesting polypeptides B870-« and B870-/? and the reaction-centre subunit L at the cytoplasmic side of the photosynthetic membrane of Rhodospirillum rubrum G-9+. Hoppe-Seylers Zeitschrift fur Physiologische Chemie 365:689
Busetta B (1986): Examination of folding patterns for predicting protein topologies. Biochimica et Biophysica Acta 870:327
Bylina E, Robles S, Youvan D (1988): Directed mutations affecting the putative bacterio-chlorophyll-binding sites in the light-harvesting I antenna of Rhodobacter capsulatus. Israel Journal of Chemistry 28:73
Chou P, Fasman G (1978): Prediction of the secondary structure of proteins from their amino acid sequence. Advances in Enzymology and Related Areas of Molecular Biology 47:45
Cohen FE, Abarbanel RM, Kuntz ID, Fletterick RJ (1986): Turn prediction in proteins using a pattern-matching approach. Biochemistry 25:266
Cornette J, Cease K, Margalit H, Spouge J, Berzofsky J, DeLisi C (1987): Hydrophobicity scales and computational techniques for detecting amphipathic structures in proteins. Journal of Molecular Biology 195:659
Cowan SW, Schirmer T, Rummel G, Steiert M, Ghosh R, Pauptit RA, Jansonius, JN, Rosenbusch JP (1992): Crystal structures explain functional properties of 2 E. coli porins. Nature 358(6389):727–733
Cramer W, Engelman D, Heijne GV, Rees D (1992): Forces involved in the assembly and stabilization of membrane proteins. Faseb Journal 6:3397
Creighton T, ed. (1992): Protein folding. New York: WH Freeman
Crielaard W, Visschers R, Fowler G, van Grondelle R, Hellingwerf K, Hunter C (1994): Probing the B800 bacteriochlorophyll binding site of the accessory light-harvesting complex from Rhodobacter sphaeroides using site-directed mutants. I. Mutagenesis, effects on binding, function and electrochromic behaviour of its carotenoids. Biochim Biophys Acta 1183:473
Deisenhofer J, Michel H (1989): The photosynthetic reaction centre from the purple bacterium Rhodopseudomonas viridis. EMBO J 8:2149
Deisenhofer J, Epp O, Mikki K, Huber R, Michel H (1985): Structure of the protein subunits in the photosynthetic reaction center of Rhodopseudomonas viridis at 3 Å resolution. Nature 318:618–624
Devereux J, Haeberli P, Smithies O (1984): A comprehensive set of sequence analysis programs for the vax. Nucleic Acids Research 12:387
Donnelly D, Overington JP, Ruffle SV, Nugent JH, Blundell TL (1993): Modeling α-helical transmembrane domains: the calculation and use of substitution tables for lipid-facing residues. Protein Science 2:55
Eccles J, Honig B, Schulten K (1988): Spectroscopic determinants in the reaction center of Rhodo-pseudomonas viridis. Biophys J 53:137–144
Eisenberg D (1984): Three-dimensional structure of membrane and surface proteins. Annual Review of Biochemistry 53:595
Eisenberg D, Schwarz E, Komaromy M, Wall R (1984): Analysis of membrane and surface protein sequences with the hydrophobic moment plot. Journal of Molecular Biology 179:125
Eisenberg D, Weiss R, Terwilliger T, Wilcox W (1982): Hydrophobic moments and protein structure. Faraday Symposia of the Chemical Society 17:109
Engelman D (1982): An implication of the structure of bacteriorhodopsin: globular membrane proteins are stabilized by polar interactions. Biophys J 37:187
Engelman DM, Steitz TA, Goldman A (1986): Identifying nonpolar transbilayer helices in amino acid sequences of membrane proteins. Ann Rev Biophys Biophys Chem 15:321–353
Fasman G (1989a): Protein conformational prediction. Trends in Biochemical Sciences 14:295
Fasman G, ed. (1989b): Prediction of protein structure and the principles of protein conformation. New York: Plenum
Fowler G, Sockalingum G, Robert B, Hunter C (1994): Blue shifts in bacteriochlorophyll absorbance correlate with changed hydrogen bonding patterns in light-harvesting 2 mutants of Rhodobacter Sphaeroides with alterations at α-Tyr-44 and α-Tyr-45. Biochemical Journal 299:695
Gamier J, Osguthorpe D, Robson B (1978): Analysis of the accuracy and implications of simple methods for predicting the secondary structure of globular proteins. Journal of Molecular Biology 120:97
Geourjon C, Deleage G (1994): SOPM: a self optimised prediction method for protein secondary structure prediction. Protein Engineering 7:157
Germeroth L, Lottspeich F, Robert B, Michel H (1993): Unexpected similarities of the B800–850 light-harvesting complex from Rhodospirillum molischianum to the B870 light-harvesting complexes from other purple photosynthetic bacteria. Biochemistry 32:5615–5621
Hawthornthwaite AM, Cogdell RJ (1991): Bacteriochlorophyll binding proteins. In: Chlorophylls, (Scheer H, ed) pp. 493–528, Boca Raton: CRC Press
Henderson R, Baldwin JM, Ceska TA, Zemlin F, Beckmann E, Downing KH (1990): Model for the structure of Bacteriorhodopsin based on high-resolution electron cryo-microscopy.J Mol Biol 213:899–929
Holley LH, Karplus M (1991): Neural networks for protein structure prediction. Methods in Enzymology 202:204
Holley LH, Karplus M (1989): Protein secondary structure prediction with a neural network. Proc Natl Acad Sci USA 86:152–156
Hu X, Xu D, Hamer K, Schulten K, Koepke J, Michel H (1995): Predicting the structure of the light-harvesting complex II of Rhodospirillum molischianum. Protein Science 4:1670–1682
Jähnig F (1989): Structure prediction for membrane proteins. In: Prediction of protein structure and the principles of protein conformation, (Fasman G, ed) p. 707, New York: Plenum
Johnson M, Srinivasan N, Sowdhamini R, Blundell T (1994): Knowledge-based protein modeling. Critical Reviews in Biochemistry and Molecular Biology 29:1
Kabsch W, Sander C (1983): Dictionary of protein secondary structure: pattern recognition of hydrogen-bonded and geometrical features. Bioploymers 22:2577
Karrasch S, Bullough P, Ghosh R (1995): 8.5Å projection map of the light-harvesting complex I from Rhodospirillum rubrum reveals a ring composed of 16 subunits. EMBO J 14:631
Kleinekofortg W, Germeroth L, van der Broek J, Schubert D, Michel H (1992): The light-harvesting complex II (B800/850) from Rhodospirillum molischianum is an octamer. Biochimica et Biophysica Acta 1140:102–104
Koepke J, Hu X, Muenke C, Schulten K, Michel H (1996): The crystal structure of the light harvesting complex II (B800–850) from Rhodospirillum molischianum. Structure (submitted)
Kramer HJM, van Grondelle R, Hunter CN, Westerhuis WHJ, Amesz J (1984): Pigment organization of the B800–850 antenna complex of Rhodopseudomonas sphaeroides. Biochim Biophys Acta 765:156–165
Kraulis P (1991): MOLSCRIPT—a program to produce both detailed and schematic plots of protein structures. J Appl Cryst 24:946–950
Kühlbrandt W, Wang D-N, Fujiyoshi Y (1994): Atomic model of plant light-harvesting complex by electron crystallography. Nature 367:614
Kuhn L, Leigh J (1985): A statistical technique for predicting membrane protein structure. Biochimica et Biophysica Acta 828:351
Kyte J, Doolittle RF (1982): A simple method for displaying the hydropathic character of a protein.J Mol Biol 157:105
Lattman E (1985): Diffraction methods for biological macromolecules. Use of the rotation and translation functions. Methods in Enzymology 115:55
Levitt M (1978): Conformational preference of amino acids in globular proteins. Biochemistry 17:4277
Lipman D, Pearson W (1985): Rapid and sensitive protein similarity searches. Science 227:1435
Lohmann R, Schneider G, Behrens D, Wrede P (1994): A neural network model for the prediction of membrane-spanning amino acid sequences. Protein Science 3:1597
Mackerell A (1995): unpublished research
Mcdermott G, Prince S, Freer A, Hawthornthwalte-Lawless A, Paplz M, Cogdell R, Isaacs N (1995): Crystal structure of an integral membrane light-harvesting complex from photosynthetic bacteria. Nature 374:517
McRee D (1993): Practical Protein Crystallography. San Diego: Academic Press
Michel H (1991): General and practical aspects of membrane protein crystallization. In: Crystallization of membrane proteins, (Michel H, ed) p. 74, Boca Raton, Florida: CRC Press
Michel H, Weyer KA, Gruenberg H, Dunger I, Oesterhelt D, Lottspeich F (1986): The ‘light’ and ‘medium’ subunits of the photosynthetic reaction centre from Rhodopseudomonas viridis: Isolation of genes, nucleotide and amino acid sequence. EMBO J. 5:1149
Olsen JD, Hunter CN (1994): Protein structure modelling of the bacterial light-harvesting complex. Photochem Photobiol 60:521
Pearson W (1990): Rapid and sensitive sequence comparison with FASTP and FASTA. Methods in Enzymology 183:63
Persson B, Argos P (1994): Prediction of transmembrane segments in proteins utilising multiple sequence alignments. Journal of Molecular Biology 237:182
Popot J (1993): Integral membrane protein structure-transmembrane α-helices as autonomous folding domains. Current Opinion In Structural Biology 3:532
Popot J, de Vitry C (1990): On the microassembly of integral membrane proteins. Annual Review of Biophysics and Biophysical Chemistry 19:369
Popot J, Engelman D (1990): Membrane protein folding and oligomerization: the two-stage model. Biochemistry 29:4031
Popot J, de Vitry C, Atteia A (1994): Folding and assembly of integral membrane proteins: An introduction. In: Membrane protein structure: experimental approaches, (White, S., ed) p. 41, New York: Oxford University press
Presnell SR, Cohen BI, Cohen FE (1992): A segment-based approach to protein secondary structure prediction. Biochemistry 31:983
Rao JM, Argos P (1986): A conformational preference parameter to predict helices in integral membrane proteins. Biochimica et Biophysica Acta 869:197
Rees D, DeAntonio L, Eisenberg D (1989): Hydrophobic organization of membrane proteins. Science 245:510
Ring C, Cohen F (1993): Modeling protein structures: construction and their applications. Faseb Journal 7:783
Rooman M, Wodak S (1988): Identification of predictive sequence motifs limited by protein structure data base size. Nature 335:45
Rossmann M, ed (1972): The Molecular Replacement Method. New York: Gordon and Breach
Sali A, Blundell TL (1993): Comparative protein modelling by satisfaction of spatial restraints. Journal of Molecular Biology 234:779
Schuler G, Altschul S, Lipman D (1991): A workbench for multiple alignment construction and analysis. Proteins: Structure, Function, and Genetics 9:180
Segrest J, Loof HD, Dohlman J, Brouillette C, Anantharamaiah G (1990): Amphipathic helix motif: Classes and properties. Proteins, Struct Funct Genet 8:103
Sundstrom V, van Grondelle R (1991): Dynamics of excitation energy transfer in pho-tosynthetic bacteria. In: Chlorophylls, (Scheer H, ed) pp. 627–704, Boca Raton: CRC Press
Treutlein H, Schulten K, Deisenhofer J, Michel H, Brünger A, Karplus M (1988): Molecular dynamics simulation of the primary processes in the photosynthetic reaction center of Rhodopseudomonas viridis. In: The Photosynthetic Bacterial Reaction Center: Structure and Dynamics, (Breton J, Verméglio A, eds) volume 149 of NATO ASI Series A: Life Sciences pp. 139–150. Plenum New York
Tuffery P, Etchebest C, Popot J, Lavery R (1994): Prediction of the positioning of the seven transmembrane α-helices of bacteriorhodopsin. A molecular simulation study. Journal of Molecular Biology 236:1105
van Grondelle R, Sundstrom V (1988): Excitation energy transfer in photosynthesis. In: Photosynthetic Light-Harvesting Systems, (Scheer H, ed) pp. 403–438, Berlin, New York: Walter de Gruyter and Co
Visschers RW, Crielaard W, Fowler GJ, Hunter CN, van Grondelle R (1994) Probing the B800 bacteriochlorophyll binding site of the accessory light-harvesting complex from Rhodobacter sphaeroides using site-directed mutants. II. A low temperature spectroscopy study of structural aspects of the pigment-protein conformation. Biochim Biophys Acta 1183:483
von Heijne G (1994a): Decoding the signals of membrane protein sequence. In: Membrane protein structure: experimental approaches, (White S, ed) p. 27, New York: Oxford University press
von Heijne G (1994b): Membrane proteins: from sequence to structure. Annual Review of Biophysics and Biomolecular Structure 23:167
von Heijne G (1992): Membrane protein structure prediction—hydrophobicity analysis and the positive-inside rule. Journal of Molecular Biology 225:487
von Heijne G (1988): Transcending the impenetrable: how proteins come to terms with membranes. Biochimica et Biophysica Acta 947:307
von Heijne G, Manoil C (1990): Membrane proteins: from sequence to structure. Protein Engineering 4:109
Weiss M, Kreusch A, Nestel U, Weite W, Weckesser J, Schulz G (1991): The structure of porin from Rhodobacter capsulatus at 1.8 Å resolution. FEBS Lett 280:379
White SH (1994): Hydropathy plots and the prediction of membrane protein topology. In: Membrane protein structure: experimental approaches, (White SH, ed), New York: Oxford University press
Zuber H (1993): Structural features of photosynthetic light-harvesting systems. In: The Photo synthetic Reaction Center, (Deisenhofer J, Norris JR, eds) p. 43, San Diego: Academic Press
Zuber H (1986): Structure of light-harvesting antenna complexes of photosynthetic bacteria, cyanobacteria and red algae. Trends Biochem Sci 11:414
Zuber H (1985): Structure and function of light-harvesting complexes and their polypeptides. Photochem Photobiol 42:821
Zuber H, Brunisholz R (1991): Structure and function of antenna polypeptides and chlorophyll-protein complexes: Principles and variability. In: Chlorophylls, (Scheer H, ed) pp. 627–692, Boca Raton: CRC Press
Zvelebil MJ, Barton GJ, Taylor WR, Sternberg MJ (1987): Prediction of protein secondary structure and active sites using the alignment of homologous sequences. J Mol Biol 195:957
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 1996 Birkhäuser Boston
About this chapter
Cite this chapter
Hu, X., Xu, D., Hamer, K., Schulten, K., Koepke, J., Michel, H. (1996). Prediction of the Structure of an Integral Membrane Protein: The Light-Harvesting Complex II of Rhodospirillum molischianum . In: Merz, K.M., Roux, B. (eds) Biological Membranes. Birkhäuser Boston. https://doi.org/10.1007/978-1-4684-8580-6_15
Download citation
DOI: https://doi.org/10.1007/978-1-4684-8580-6_15
Publisher Name: Birkhäuser Boston
Print ISBN: 978-1-4684-8582-0
Online ISBN: 978-1-4684-8580-6
eBook Packages: Springer Book Archive