Abstract
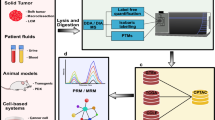
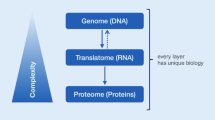
The field of proteomics holds promise for the discovery of new biomarkers for the early detection and diagnosis of disease, molecular targets for therapy and markers for therapeutic efficacy and toxicity. A variety of proteomics approaches may be used to address these goals. Two-dimensional gel electrophoresis (2D-PAGE) is the cornerstone of many discovery-based proteomics studies. Technologies such as laser capture microdissection (LCM) and highly sensitive MS methods are currently being used together to identify greater numbers of lower abundance proteins that are differentially expressed between defined cell populations. Newer technologies such as reverse phase protein arrays will enable the identification and profiling of target pathways in small biopsy specimens. Surface-enhanced laser desorption/ionization time-of-flight (SELDI-TOF) analysis enables the high throughput characterization of lysates from very few tumor cells or body fluids and may be best suited for diagnosis and monitoring of disease. Such technologies are expected to supplement our arsenal of mRNA-based assays, and we believe that in the future, entire cellular networks and not just a single deregulated protein will be the target of therapeutics and that we will soon be able to monitor the status of these pathways in diseased cells before, during and after therapy.
Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only
Preview
Unable to display preview. Download preview PDF.
Similar content being viewed by others
References
Steiner, S. and Witsmann, F. Proteomics: Applications and opportunities in preclinical drug development., Electrophoresis.21:2099–2104, 2000.
Celis, J., Kruhoffer, M., Gromova, I., Frederiksen, C., Ostergaard, M., Thykjaer, T., Gromov, P., Yu, J., Palsdottir, H., Magnusson, N., and Orntoft, T. Gene expression profiling: Monitoring transcription and translation products using DNA microarrays and proteomics., FEBS Lett.480:2–16, 2000.
Oda, Y., Nagasu, T., and Chait, B. Enrichment analysis of phosphorylated proteins as a tool for probing the phosphoproteome., Nat. Biotech.19:379–382, 2001.
Ducret, A., Desponts, C., Desmarais, S., Gresser, M., and Ramachandran, C. A general method for the rapid characterization of tyrosine-phosphorylated proteins by mini two-dimensional gel electrophoresis., Electrophoresis.21:2196–2208, 2000.
Larsson, T., Bergstrom, J., Nilsson, J., and Karlsson, K. Use of an affinity proteomics approach for the identification of low-abundant bacterial adherins as applied on the Lewis(b) binding adhesin of Helicobacter pylori., FEBS Lett.469:155–158, 2000.
Charlwood, J., Skehel, J., and Camilleri, P. Analysis of N-linked oligosaccharides released from glycoproteins separated by two-dimensional gel electrophoresis., Anal. Biochem.284:49–59, 2000.
Fivaz, M., Vilbois, F., Pasquali, C., and vanderGoot, F. Analysis of glycosyl phosphatidylinositol-anchored proteins by two-dimensional gel electrophoresis., Electrophoresis.21:3351–3356, 2000.
Lopez, M., Kristal, B., Chernokalskaya, E., Lazarev, A., Shestopalov, A., Bogdanove, A., and Robinson, M. High-throughput profiling of the mitochondrial proteome using affinity fractionation and automation., Electrophoresis.21:3427–3440, 2000.
Koe, E., Burkhart, W., Blackburn, K., Koc, H., Moseley, A., and Spremulli, L. Identification of four proteins from the small subunit of the mammalian mitochondrial ribosome using a proteomics approach., Protein Sci.10:471–481, 2001.
Godovac-Zimmerman, J., Sockic, V., Poznanovic, S., and Brianza, F. Functional proteomics of signal transduction by membrane receptors., Electorphoresis.20:952–961, 1999.
Kidd, D., Liu, Y., and Cravatt, B. Profiling serine hydrolase activities in complex proteomes., Biochem.40:4005–4015, 2001.
Adam, G., Cravatt, B., and Sorensen, E. Profiling the specific reactivity of the proteome with non-directed activity-based probes., Chemistry and Biol.8:81–95, 2001.
Lewis, T., Hunt, J., Aveline, L., Jonscher, K., Louie, D., Yeh, J., Nahreini, T., Resing, K., and Ahn, N. Identification of novel MAP kinase pathway signaling targets by functional proteomics and mass spectroscopy., Mol. Cell.6:1343–1354, 2000.
Ilag, L., Ng, J., and Jay, D. Chromophore-assisted laser inactivation (CALI) to validate drug targets and pharmacogenomic markers., Drug Dev’t. Res49:65–73, 2000.
Chambers, G., Lawrie, L., Cash, P., and Murray, G. Proteomics: A new approach to the study of disease., J. Pathol.192:280–288, 2000.
Rudert, F. Genomics and proteomics tools for the clinic., Curr. Opinion in Mol. Therapeutics.2:633–642, 2000.
Hanash, S. Opermics: Molecular analysis of tissues from DNA to RNA to protein., Clin. Chem. and Lab. Med.38:805–813, 2000.
Emmert-Buck, M., Bonner, R., Smith, P., Chuaqui, R., Zhuang, Z., Goldstein, S., Weiss, R., and Liotta, L. Laser capture microdissection., Science.274:998–1001., 1996.
Wulfkuhle, J., Sgroi, D., Krutzsch, H., McLean, K., McGarvey, K., Knowlton, M., Chen, S., Shu, H., Sahin, A.. Kurek, R., Wallwiener, D., Merino, M., Petricoin, E., Zhao, Y., and Steeg, P. Proteomics of human breast ductal carcinoma in situ (DCIS). In press., 2002.
Ornstein, D., Gillespie, J., Paweletz, C., Duray, P., Herring, J., Vocke, C., Topalian, S., Bostwick, D., Linehan, W., III, E. P., and Emmert-Buck, M. Proteomic analysis of laser capture microdissected human prostate cancer and in vitro prostate cell lines., Electrophoresis.21:2235–2242, 2000.
Banks, R., Dunn, M., Forbes, M., Stanley, A., Pappin, D., naven, T., Gough, M., Harnden, P., and Selby, P. The potential use of laser capture microdissection to selectively obtain distinct populations of cells forproteomic analysis - Preliminary findings., Electrophoresis.20:689–700, 1999.
Eggling, F. v., Davies, H., Loms, L., Fiedler, W., Junker, K., Claussen, U., and Ernst, G. Tissue specific microdissection coupled with ProteinChip(R)array technologies: Applications in cancer research., Biotechniques.29:1066–1070, 2000.
Gillespie. J., Ahram, M., Best, C., Swalwell, J., Krizman, D., Petricoin, E., Liotta, L., and Emmert-Buck, M. The role of tissue microdissection in cancer research, Cancer J.7:32–39, 2001.
Hutchens, T. and Yip, T. New desorption strategies for the mass spectrometric analysis of macromolecules., Rapid Commun. Mass Spectrom.7:576–580, 1993.
Merchant, M. and Weinberger, S. Recent advancements in surface-enhanced laser desorption/ionization time-of-flight mass spectometry., Electrophoresis.21:1164–1167, 2000.
Isaaq, H., Veenstra, T., Conrads, T. and Felschow, D. The SELDI-TOF MS approach to proteomics: Protein profiling and biomarker identification., Biochem. Biophys. Res. Commun.292:587–592, 2002.
Paweletz, C., Gillespie, J., Ornstein, D., Simone, N., Brown, M., Cole, K., Kohn, E., Linehan, W., Weber, T., Taylor, P., Emmert-Buck, M., Liotta, L., and Petricoin III, E. Rapid protein display profiling of cancer progression directly from human tissue using a protein biochip., Drug Dev’t. Res.49:34–42, 2000.
Vlahou, A., Schellhammer, P., Mendrinos, S., Patel, K., Kondlyis, F., Gong, L., Nasim, S. and Wright, Jr., G. Development of a novel proteomic approach for the detection of transitional cell carcinoma of the bladder in urine., Amer. J. Pathol.158:1491–1502, 2001.
Li, J., Zhang, Z., Rosenzweig, J., Wang, Y., and Chan, D. Proteomics and bioinformatics approaches for identification of serum biomarkers to detect breast cancer., Clin. Chem.48:1296–1304, 2002.
Petricoin III, E., Ardekani, A., Hitt, B., Levine, P., Fusaro, V., Steinberg, S., Mills, G., Simone. C., Fishman, D., Kohn, E. and Liotta, L. Use of proteomic patterns inserum to identify ovarian cancer., Lancet.359:572–577, 2002.
Ozols, R., Rubin, S., Thomas, G. and Robboy, S. Epithelial ovarian cancer. In: Principles and practice of gynecologic oncology. Hoskins, W., Perez, C. and Young, R., eds. Philadelphia: Lippincott Williams and Wilkins. 981–1058, 2000.
Petricoin III, E., Zoon, K., Kohn, E., Barrett, C. and Liotta, L. Clinical proteomics: Translating benchside promise into bedside reality. Nat. Drug Disc.1:683–695, 2002.
Paweletz, C., Charboneau, L., Bichsel, V., Simone, N., Chen, T., Gillespie, J., Emmert-Buck, M., Roth, M., III, E. P., and Liotta, L. Reverse phase protein microarrays which capture disease progression show activation of pro-survival pathways at the cancer invasion front., Oncogene.20:1981–1989, 2001.
Torhorst, J., Bucher, C., Kononen, J., Haas, P., Zuber, M., Kochli, O., Mross, F., Dieterich, H., Moch, H., Mihatsch, M., Kallioneimi, O. and Sauter, G. Tissue microarrays for rapid linking of molecular changes to clinical endpoints., Amer. J. Pathol.159:2249–2256, 2001.
Knezevic, V., Leethanakul, C., Bichsel, VI, Worth, J., Prabhu, V., Gutkind, J., Liotta, L., Munson, P., Petricoin III, E. and Krizman, D. Proteomic profiling of the cancer micorenvironment by antibody arrays., Proteomics.1:1271–1278, 2001.
Sreekumar, A., Nyati, M., Varambally, S., Barrette, T., Ghosh, D., Lawrence, T. and Chinniyan, A. Profiling of cancer cells using protein microarrays: Discovery of novel radiation-regulated proteins., Cancer Res.61:7585–93, 2001.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2003 Springer Science+Business Media New York
About this chapter
Cite this chapter
Wulfkuhle, J.D., Paweletz, C.P., Steeg, P.S., Petricoin, E.F., Liotta, L. (2003). Proteomic Approaches to the Diagnosis, Treatment, and Monitoring of Cancer. In: Llombart-Bosch, A., Felipo, V. (eds) New Trends in Cancer for the 21st Century. Advances in Experimental Medicine and Biology, vol 532. Springer, Boston, MA. https://doi.org/10.1007/978-1-4615-0081-0_7
Download citation
DOI: https://doi.org/10.1007/978-1-4615-0081-0_7
Publisher Name: Springer, Boston, MA
Print ISBN: 978-1-4613-4914-3
Online ISBN: 978-1-4615-0081-0
eBook Packages: Springer Book Archive