Abstract
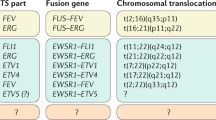
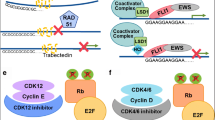
The Ewing’s sarcoma family of tumors (EFT) is a group of malignancies affecting bone and soft tissue in adolescents. It is characterized by a unique gene rearrangement between theEWSgene and anetstranscription factor gene. EFT can be cured with conventional multi modal treatment, however, about 40% of patients still succumb to the disease. Relapses can be observed more than 5 years after the end of primary treatment suggesting persistence of minimal residual disease (MRD). Due to the still enigmatic nature of EFT histogenesis the phenotype of EFT stem cells and of dormant tumor cells remains unknown. The most frequent fusion product associated with EFT, EWS-FLIT, is the founding member of a whole class of similarly structured chimeric proteins associated with a variety of human sarcomas and also specific leukemias. The corresponding gene rearrangement constitutes a rate limiting step in oncogenesis as implied by the high association of EFT withEWS-ETSfusions, strong selective pressure for maintenance of a correct reading frame in the tumors, and by experimental data confirming the transforming and tumorigenic potential of EWS-FLI1. Understanding the biology ofEWS-ETSgene fusions and its interplay with essential cellular pathways regulating cell growth, apoptosis, differentiation, genomic integrity, and treatment resistance may unravel specifically vulnerable sites for therapeutic targeting. This review summarizes the current knowledge about the EWS-FLI1 pathway in EFT and provides some ideas as to how this knowledge may be translated into innovative treatment approaches.
Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only
Preview
Unable to display preview. Download preview PDF.
Similar content being viewed by others
References
Turc Carel, C., Aurias, A., Mugneret, F., Lizard, S., Sidaner, I., Volk, C., Thiery, J. P., Olschwang, S., Philip, I., Berger, M. P., and et al Chromosomes in Ewing’s sarcoma. I. An evaluation of 85 cases of remarkable consistency of t(11;22)(g24;g12). Cancer Genet.Cytogenet.32:229–238, 1988.
Delattre, O., Zucman, J., Plougastel, B., Desmaze, C., Melot, T., Peter, M., Kovar, H., Joubert, I., De Jong, P., Rouleau, G., and et al Gene fusion with an ETS DNA-binding domain caused by chromosome translocation in human tumours. Nature, 359: 162–165, 1992.
Kovar, H., Aryee, D., and Zoubek, A. The Ewing family of tumors and the search for the Achilles’ heel. Curr.Opin.Oncol11:275–284, 1999.
Deneen, B. and Denny, C. T. Loss of p16 pathways stabilizes EWS/FLII expression and complements EWS/FLI1 mediated transformation. Oncogene20:6731–6741, 2001.
Lessnick, S. L., Braun, B. S., Denny, C. T., and May, W. A. Multiple domains mediate transformation by the Ewing’s sarcoma EWS/FLI-1 fusion gene. Oncogene10:423–431, 1995.
Toretsky, J. A., Kalebic, T., Blakesley, V., LeRoith, D., and Helman, L. J. The insulin-like growth factor-I receptor is required for EWS/FLI-1 transformation of fibroblasts [In Process Citation]. J.Biol.Chem.272:30822–30827, 1997.
Scotlandi, K., Benini, S., Nanni, P., Lollini, P. L., Nicoletti, G., Landuzzi, L., Serra, M., Manara, M. C., Picci, P., and Baldini, N. Blockage of insulin-like growth factor-I receptor inhibits the growth of Ewing’s sarcoma in athymic mice. Cancer Res.58:4127–4131, 1998.
May, W. A., Gishizky, M. L., Lessnick, S. L., Lunsford, L. B., Lewis, B. C., Delattre, O., Zucman, J., Thomas, G., and Denny, C. T. Ewing sarcoma 11;22 translocation produces a chimeric transcription factor that requires the DNA-binding domain encoded by FLIT for transformation. Proc.Natl.Acad.Sci.U.S.A.90:5752–5756, 1993.
Arvand, A., Welford, S. M., Teitell, M. A., and Denny, C. T. The COON-terminal domain of FLI-1 is necessary for full tumorigenesis and transcriptional modulation by EWS/FLI-1. Cancer Res.61:5311–5317, 2001.
Jaishankar, S., Zhang, J., Roussel, M. F., and Baker, S. J. Transforming activity of EWS/FLI is not strictly dependent upon DNA- binding activity [In Process Citation]. Oncogene18:5592–5597, 1999.
Welford, S. M., Hebert, S. P., Deneen, B., Arvand, A., and Denny, C. T. DNA binding domain-independent pathways are involved in EWS/FLI1-mediated oncogenesis. J.Biol.Chem.276:4197741984,2001.
Zucman-Rossi, J., Legoix, P., Victor, J. M., Lopez, B., and Thomas, G. Chromosome translocation based on illegitimate recombination in human tumors. Proc.Natl.Acad.Sci.U.S.A., 95: 11786–11791, 1998.
Zucman, J., Melot, T., Desmaze, C., Ghysdael, J., Plougastel, B., Peter, M., Zucker, J. M., Triche, T. J., Sheer, D., Turc Carel, C., and et al Combinatorial generation of variable fusion proteins in the Ewing family of tumours. EMBO J.12:4481–4487, 1993.
Kovar, H., Aryee, D. N., Jug, G., Henockl, C., Schemper, M., Delattre, O., Thomas, G., and Gadner, H. EWS/FLI-1 antagonists induce growth inhibition of Ewing tumor cells in vitro. Cell Growth Differ.7:429–437, 1996.
Ouchida, M., Ohno, T., Fujimura, Y., Rao, V. N., and Reddy, E. S. Loss of tumorigenicity of Ewing’s sarcoma cells expressing antisense RNA to EWS-fusion transcripts. Oncogene, 11: 1049–1054, 1995.
Tanaka, K., Iwakuma, T., Harimaya, K., Sato, H., and Iwamoto, Y. EWS-Flit antisense oligodeoxynucleotide inhibits proliferation of human Ewing’s sarcoma and primitive neuroectodermal tumor cells. J.CIin.Invest.99:239–247, 1997.
Toretsky, J. A., Connell, Y., Neckers, L., and Bhat, N. K. Inhibition of EWS-FLI-1 fusion protein with antisense oligodeoxynucleotides. J.Neurooncol.31:9–16, 1997.
Kovar, H., Auinger, A., Jug, G., Aryee, D., Zoubek, A., Salzer Kuntschik, M., and Gadner, H. Narrow spectrum of infrequent p53 mutations and absence of MDM2 amplification in Ewing tumours. Oncogene8:2683–2690, 1993.
Kovar, H., Jug, G., Aryee, D. N. T., Zoubek, A., Ambros, P., Gruber, B., Windhager, R., and Gadner, H. Among genes involved in the RB dependent cell cycle regulatory cascade, the p16 tumor suppressor gene is frequently lost in the Ewing family of tumors. Oncogene15:1997.
Arvand, A. and Denny, C. T. Biology of EWS/ETS fusions in Ewing’s family tumors. Oncogene20:5747–5754,2001.
Kovar, H., Dworzak, M., Strehl, S., Schnell, E., Ambros, I. M., Ambros, P. F., and Gadner, H. Overexpression of the pseudoautosomal gene MIC2 in Ewing’s sarcoma and peripheral primitive neuroectodermal tumor. Oncogene, 5: 1067–1070, 1990.
Lessnick, S. L., Dacwag, C. S., and Golub, T. R. The Ewing’s sarcoma oncoprotein EWS/FLI induces a p53-dependent growth arrest in primary human fibroblasts. Cancer Cell 1, 393–401. 2002. Ref Type: Generic
Matsumoto, Y., Tanaka, K., Nakatani, F., Matsunobu, T., Matsuda, S., and Iwamoto, Y. Downregulation and forced expression of EWS-Fli 1 fusion gene results in changes in the expression of G(1)regulatory genes. Br.J.Cancer84:768–775, 2001.
Girnita, L., Girnita, A., Wang, M., Meis-Kindblom, J. M., Kindblom, L. G., and Larsson, O. A link between basic fibroblast growth factor (bFGF) and EWS/FLI-1 in Ewing’s sarcoma cells. Oncogene19:4298–4301, 2000.
Thompson, A. D., Teitell, M. A., Arvand, A., and Denny, C. T. Divergent Ewing’s sarcoma EWS/ETS fusions confer a common tumorigenic phenotype on NIH3T3 cells [In Process Citation]. Oncogene18:5506–5513, 1999.
Bertolotti, A., Lutz, Y., Heard, D. J., Chambon, P., and Tora, L. hTAF(II)68, a novel RNA/ssDNAbinding protein with homology to the pro-oncoproteins TLS/FUS and EWS is associated with both TFIID and RNA polymerase II. EMBO J.15:5022–5031, 1996.
Bertolotti, A., Melot, T., Acker, J., Vigneron, M., Delattre, O., and Tora, L. EWS, but not EWS-FLI1, is associated with both TFIID and RNA polymerase II: interactions between two members of the TET family, EWS and hTAFII68, and subunits of TFIID and RNA polymerase II complexes [In Process Citation]. Mol.Cell Biol.18:1489–1497, 1998.
Petermann, R., Mossier, B. M., Aryee, D. N., Khazak, V., Golemis, E. A., and Kovar, H. Oncogenic EWS-Flil interacts with hsRPB7, a subunit of human RNA polymerase II [In Process Citation]. Oncogene17:603–610, 1998.
Chansky, H. A., Hu, M., Hickstein, D. D., and Yang, L. Oncogenic TLS/ERG and EWS/Fli-1 fusion proteins inhibit RNA splicing mediated by YB-1 protein. Cancer Res.61:3586–3590, 2001.
Knoop, L. L. and Baker, S. J. The splicing factor U1C represses EWS/FLI-mediated transactivation. J.Biol.Chem.275:24865–24871, 2000.
Yang, L., Embree, L. J., Tsai, S., and Hickstein, D. D. Oncoprotein TLS interacts with serine-arginine proteins involved in RNA splicing. J.Biol.Chem.273:27761–27764, 1998.
Yang, L., Chansky, H. A., and Hickstein, D. D. EWS[middle dot]Fli-1 fusion protein interacts with hyperphosphorylated RNA polymerase II and interferes with serine-arginine protein-mediated RNA splicing. J.Biol.Chem.275:37612–37618, 2000.
Zhang, D., Paley, A. J., and Childs, G. The transcriptional repressor ZFM1 interacts with and modulates the ability of EWS to activate transcription. J.Biol.Chem.273:18086–18091, 1998.
Calvio, C., Neubauer, G., Mann, M., and Lamond, A. I. Identification of hnRNP P2 as TLS/FUS using electrospray mass spectrometry. RNA.1:724–733, 1995.
Hahm, K. B., Cho, K., Lee, C., Im, Y. H., Chang, J., Choi, S. G., Sorensen, P. H., Thiele, C. J., and Kim, S. J. Repression of the gene encoding the TGF-beta type II receptor is a major target of the EWS-FLIT oncoprotein. Nat.Genet.23:222–227, 1999.
Yang, L., Embree, L. J., and Hickstein, D. D. TLS-ERG leukemia fusion protein inhibits RNA splicing mediated by serine-arginine proteins. Mol.Cell Biol.20:3345–3354, 2000.
Knoop, L. L. and Baker, S. J. EWS/FLI alters 5’-splice site selection. J.Biol.Chem.276:2231722322,2001.
Ohkura, N., Yaguchi, H., Tsukada, T., and Yamaguchi, K. The EWS/NOR1 fusion gene product gains a novel activity affecting pre-mRNA splicing. J.Biol.Chem.277:535–543, 2002.
Spahn, L., Petermann, R., Siligan, C., Schmid, J. A., Aryee, D. N., and Kovar, H. Interaction of the EWS NH2 terminus with BARD1 links the Ewing’s sarcoma gene to a common tumor suppressor pathway. Cancer Res.62:4583–4587, 2002.
Kleiman, F. E. and Manley, J. L. Functional interaction of BRCA1-associated BARD1 with polyadenylation factor CstF-50. Science285:1576–1579, 1999.
Kleiman, F. E. and Manley, J. L. The BARD1-CstF-50 Interaction Links mRNA 3’ End Formation to DNA Damage and Tumor Suppression. Cell104:743–753, 2001.
Venkitaraman, A. R. Functions of BRCA1 and BRCA2 in the biological response to DNA damage. J.Cell Sci.114:3591–3598, 2001.
Hicks, G. G., Singh, N., Nashabi, A., Mai, S., Bozek, G., Klewes, L., Arapovic, D., White, E. K., Koury, M. J., Oltz, E. M., Van Kaer, L., and Ruley, H. E. Fus deficiency in mice results in defective B-lymphocyte development and activation, high levels of chromosomal instability and perinatal death. Nat.Genet.24:175–179, 2000.
Kuroda, M., Sok, J., Webb, L., Baechtold, H., Urano, F., Yin, Y., Chung, P., de Rooij, D. G., Akhmedov, A., Ashley, T., and Ron, D. Male sterility and enhanced radiation sensitivity in TLS(-/-) mice. EMBO J.19:453–462, 2000.
Soldatenkov, V. A., Trofimova, I. N., Rouzaut, A., McDermott, F., Dritschilo, A., and Notario, V. Differential regulation of the response to DNA damage in Ewing’s sarcoma cells by ETS1 and EWS/FLI-1. Oncogene21:2890–2895, 2002.
Prasad, S. C., Thraves, P. J., Bhatia, K. G., Smulson, M. E., and Dritschilo, A. Enhanced poly(adenosine diphosphate ribose) polymerase activity and gene expression in Ewing’s sarcoma cells. Cancer Res.50:38–43, 1990.
Powers, C. A., Mathur, M., Raaka, B. M., Ron, D., and Samuels, H. H. TLS (translocated-inliposarcoma) is a high-affinity interactor for steroid, thyroid hormone, and retinoid receptors. Mol.Endocrinol.12:4–18, 1998.
Uranishi, H., Tetsuka, T., Yamashita, M., Asamitsu, K., Shimizu, M., Itoh, M., and Okamoto, T. Involvement of the pro-oncoprotein TLS (translocated in liposarcoma) in nuclear factor-kappa B p65-mediated transcription as a coactivator. J.Biol.Chem.276:13395–13401, 2001.
Deloulme, J. C., Prichard, L., Delattre, O., and Storm, D. R. The prooncoprotein EWS binds calmodulin and is phosphorylated by protein kinase C through an IQ domain [In Process Citation]. J.Biol.Chem.272:27369–27377, 1997.
Felsch, J. S., Lane, W. S., and Peralta, E. G. Tyrosine kinase Pyk2 mediates G-protein-coupled receptor regulation of the Ewing sarcoma RNA-binding protein EWS. Curr.Biol.9:485–488, 1999.
Guinamard, R., Fougereau, M., and Seckinger, P. The SH3 domain of Bruton’s tyrosine kinase interacts with Vav, Sam68 and EWS. Scand.J.Immunol., 45: 587–595, 1997.
Olsen, R. J. and Hinrichs, S. H. Phosphorylation of the EWS IQ domain regulates transcriptional activity of the EWS/ATF1 and EWS/FLI1 fusion proteins. Oncogene20:1756–1764, 2001.
Kim, J., Lee, J. M., Branton, P. E., and Pelletier, J. Modulation of EWS/WT1 activity by the v-Src protein tyrosine kinase. FEBS Lett.474:121–128, 2000.
Matsuoka, Y., Matsuoka, Y., Shibata, S., Yasuhara, N., and Yoneda, Y. Identification of Ewing’s Sarcoma Gene Product as a Glycoprotein Using a Monoclonal Antibody that Recognizes an Immunodeterminant Containing 0-Linked N-Acetylglucosamine Moiety. Hybrid.Hybridomics.21:233–236, 2002.
Belyanskaya, L. L., Gehrig, P. M., and Gehring, H. Exposure on cell surface and extensive arginine methylation of EWS protein. J.Biol.Chem.,.: 2001.
Kovar, H., Jug, G., Hattinger, C., Spahn, L., Aryee, D. N., Ambros, P. F., Zoubek, A., and Gadner, H. The ews protein is dispensable for ewing tumor growth. Cancer Res.61:5992–5997, 2001.
Khazak, V., Sadhale, P. P., Woychik, N. A., Brent, R., and Golemis, E. A. Human RNA polymerase II subunit hsRPB7 functions in yeast and influences stress survival and cell morphology. Mol.Biol.Cell6:759–775, 1995.
Li, K. K. and Lee, K. A. Transcriptional activation by the Ewing’s sarcoma (EWS) oncogene can be cis-repressed by the EWS RNA-binding domain. J.Biol.Chem.275:23053–23058, 2000.
Lambert, G., Bertrand, J. R., Fattal, E., Subra, F., Pinto-Alphandary, H., Malvy, C., Auclair, C., and Couvreur, P. EWS fli-1 antisense nanocapsules inhibits ewing sarcoma-related tumor in mice. Biochem.Biophys.Res.Commun.279:401–406, 2000.
McManus, M. T. and Sharp, P. A. Gene silencing in mammals by small interfering RNAs. Nat.Rev.Genet., 3: 737–747, 2002.
Capodici, J., Kariko, K., and Weissman, D. Inhibition of HIV-1 Infection by Small Interfering RNA-Mediated RNA Interference. J.Immunol.169:5196–5201, 2002.
Xia, H., Mao, Q., Paulson, H. L., and Davidson, B. L. siRNA-mediated gene silencing in vitro and in vivo. Nat.Biotechnol.20:1006–1010, 2002.
Lewis, D. L., Hagstrom, J. E., Loomis, A. G., Wolff, J. A., and Herweijer, H. Efficient delivery of siRNA for inhibition of gene expression in postnatal mice. Nat.Genet.32:107–108, 2002.
Brurnmelkamp, T., Bernards, R., and Agami, R. Stable suppression of tumorigenicity by virus-mediated RNA interference. Cancer Cell2:243, 2002.
Zeng, Y. and Cullen, B. R. RNA interference in human cells is restricted to the cytoplasm. RNA.8:855–860,2002.
Zoubek, A., Pfleiderer, C., Salzer-Kuntschik, M., A.mann, G., Windhager, R., Fink, F. M., Koscielniak, E., Delattre, 0., Strehl, S., and Ambros, P. F. Variability of EWS chimaeric transcripts in Ewing tumours: a comparison of clinical and molecular data. Br.J.Cancer70:908–913, 1994.
Yi, H., Fujimura, Y., Ouchida, M., Prasad, D. D., Rao, V. N., and Reddy, E. S. Inhibition of apoptosis by normal and aberrant Fli-1 and erg proteins involved in human solid tumors and leukemias. Oncogene14:1259–1268, 1997.
Wang, M., Xie, Y., Girnita, L., Nilsson, G., Dricu, A., Wejde, J., and Larsson, O. Regulatory Role of Mevalonate and N-Linked Glycosylation in Proliferation and Expression of the EWS/FLI-1 Fusion Protein in Ewing’s Sarcoma Cells. Exp.Cell Res.246:38–46, 1999.
Mackall, C., Berzofsky, J., and Hetman, L. J. Targeting tumor specific translocations in sarcomas in pediatric patients for immunotherapy. Clin.Orthop.275:25–31, 2000.
Author information
Authors and Affiliations
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2003 Springer Science+Business Media New York
About this chapter
Cite this chapter
Kovar, H. (2003). Ewing Tumor Biology: Perspectives for Innovative Treatment Approaches. In: Llombart-Bosch, A., Felipo, V. (eds) New Trends in Cancer for the 21st Century. Advances in Experimental Medicine and Biology, vol 532. Springer, Boston, MA. https://doi.org/10.1007/978-1-4615-0081-0_4
Download citation
DOI: https://doi.org/10.1007/978-1-4615-0081-0_4
Publisher Name: Springer, Boston, MA
Print ISBN: 978-1-4613-4914-3
Online ISBN: 978-1-4615-0081-0
eBook Packages: Springer Book Archive