Abstract
The Yes-associated protein (YAP) and WW domain-containing transcription regulator 1 (WWTR1, also known as TAZ) are two transcription co-activators that act downstream of the Hippo tumor suppressor pathway. YAP/TAZ regulate expression of a large number of genes that are important in controlling organ size, tumorigenesis, and stem cell functions. The activity of YAP/TAZ is mainly inhibited by Lats kinases of the Hippo pathway. Upon phosphorylation by Lats kinases, YAP/TAZ are sequestered in the cytoplasm and undergo ubiquitination-mediated degradation. YAP/TAZ are also inhibited by interaction with cell junction proteins including angiomotin and α-catenin. Moreover, as transcription co-activators, YAP/TAZ need to associate with DNA-binding proteins such as TEAD family transcription factors to induce gene expression. Hence, the activity and specificity of YAP/TAZ in gene expression is also dependent on their nuclear partners.
Access provided by Autonomous University of Puebla. Download chapter PDF
Similar content being viewed by others
Keywords
Organ size regulation is fundamental in biology and is critical not only during development but also in adulthood. A key determinant of organ size is the number of cells. Thus, organ size control is largely dependent on modulation of cell numbers. The Hippo signaling pathway, initially identified in Drosophila, has important roles in regulating cell proliferation and cell death, which consequently determines cell numbers, tissue growth, and organ size.
Hippo (Hpo), a Drosophila serine/threonine kinase, has been named after a massive overgrowth phenotype resulting from its genetic inactivation (Harvey et al. 2003; Jia et al. 2003; Pantalacci et al. 2003; Wu et al. 2003). Additional core components of the Hippo pathway, such as Sav, Wts, and Mats, were defined similarly by genetic screens in Drosophila (Justice et al. 1995; Kango-Singh et al. 2002; Lai et al. 2005; Tapon et al. 2002; Xu et al. 1995). The transcription co-activator Yki mediates the biological functions of the Hippo pathway by regulating a broad transcription program (Goulev et al. 2008; Huang et al. 2005; Zhang et al. 2008; Zhao et al. 2008).
Organ size regulation by the Hippo pathway is evolutionarily conserved in mammals. Furthermore, dysregulation of this pathway leads to hyperplasia and tumorigenesis (reviewed by Zhao et al. (2010a)). The mammalian Hippo pathway is composed of a kinase cascade consisting of mammalian STE20-like protein kinase 1/2 (MST1/2, Hpo ortholog) and large tumor suppressor homolog 1/2 (Lats1/2). MST1/2, in complex with its regulatory protein Salvador (Sav), phosphorylates and activates Lats1/2 kinases (Callus et al. 2006; Chan et al. 2005). Lats1/2 also forms a complex with regulatory protein Mobkl1A/Mobkl1B (Mats ortholog, collectively referred to as Mob1 below) and phosphorylates the transcription co-activators Yes-associated protein (YAP, Yki ortholog) and WW domain containing transcription regulator 1 (WWTR1, also known as TAZ, a YAP paralog) (Chow et al. 2010; Hao et al. 2008; Lei et al. 2008; Zhao et al. 2007). YAP/TAZ are two major downstream effectors mediating functions of the mammalian Hippo pathway in development, organ size control, and tumorigenesis.
1 Biological Functions of YAP/TAZ
Functions of YAP and TAZ overlap but are not completely redundant, as revealed by the different phenotypes in YAP or TAZ knockout mice. YAP-null mice die at embryonic day 8.5 (E8.5), with defects in yolk sac vasculogenesis, chorioallantoic fusion, and body axis elongation (Morin-Kensicki et al. 2006), suggesting that YAP plays an important function in development. In contrast, TAZ knockout mice are viable but predisposed to renal and pulmonary diseases (Hossain et al. 2007; Makita et al. 2008; Tian et al. 2007). Furthermore, YAP/TAZ double knockout mice die before the morula stage (16–32 cells), prior to embryo implantation, indicating essential roles of YAP/TAZ in early embryonic development (Nishioka et al. 2009).
YAP/TAZ also play important roles in stem cell self-renewal and differentiation (Liu et al. 2012a). YAP activity declines when stem cells undergo differentiation (Lian et al. 2010; Tamm et al. 2011), and YAP and TAZ are required for maintaining the pluripotency of mouse and human stem cells, respectively (Alarcon et al. 2009; Lian et al. 2010; Varelas et al. 2010b). In addition, overexpression of YAP or knockdown of Lats2 increases the induction efficiency of induced pluripotent stem (iPS) cells (Lian et al. 2010; Qin et al. 2012). In transgenic animals, enhanced YAP activity expands tissue-specific stem cells in liver, intestine, skin, and neural tube (Benhamouche et al. 2010; Camargo et al. 2007; Cao et al. 2008; Lee et al. 2010; Lu et al. 2010; Schlegelmilch et al. 2011; Song et al. 2010; Zhang et al. 2011; Zhou et al. 2011). These observations collectively demonstrate an important function of YAP/TAZ in both embryonic and tissue-specific stem cells.
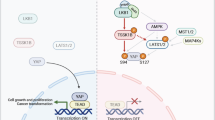
Given the importance of the Hippo pathway in cell number control, it is no surprise that alteration of this pathway contributes to tumor development. Indeed, the YAP gene locus is amplified in hepatocellular carcinoma and mammary tumors (Overholtzer et al. 2006; Zender et al. 2006), and elevated YAP or TAZ expression and nuclear localization have been frequently observed in human cancers (Chan et al. 2008; Dong et al. 2007; Steinhardt et al. 2008; Zender et al. 2006; Zhao et al. 2007). On the other hand, MST1/2 and Lats1/2 are downregulated in different type of cancers (reviewed by Zhao et al. (2010a)). In YAP transgenic mice, hyperplasia and tumors are frequently observed (Camargo et al. 2007; Dong et al. 2007). Similarly, inactivation of Hippo pathway components leads to tumor development (Lee et al. 2008, 2010; Lu et al. 2010; Song et al. 2010; Zhou et al. 2009). Moreover, neurofibromin 2 (NF2), which acts upstream of MST1/2, is a well-known human tumor suppressor (Rouleau et al. 1993; Ruttledge et al. 1994). These observations suggest that YAP/TAZ and the Hippo pathway play critical roles in cancer development.
YAP contains multiple domains, such as a proline-rich domain, TEAD-binding domain, two WW domains (or one in a shorter splicing variant), an SH3-binding motif, a transcription activation domain, a coiled-coil domain, and a PDZ-binding motif. TAZ comprises similar domains, although it lacks the proline-rich domain, the second WW domain, and the SH3-binding motif. The Drosophila Yki is more divergent as it lacks the proline-rich domain, SH3-binding motif, coiled-coil domain, and PDZ-binding motif (Fig. 5.1). These domains set up a platform for YAP/TAZ to form an extensive interactions with their upstream regulators and downstream effectors (reviewed by Mauviel et al. (2012)).
Domain organization and Lats targeting sites on YAP/TAZ/Yki. Protein domains are illustrated using gray boxes. Lats1/2 phosphorylation sites (HXRXXS) on YAP, TAZ, and Yki are depicted by blue circles with a “P” label, and the numbers below them represent their exact positions. TEAD- or Sd-binding domains, transactivation domains, SH3-binding motif, and PDZ-binding motif are also indicated. Pro proline-rich domain; WW WW domain; CC coiled-coil domain
The activity of YAP/TAZ is tightly controlled to maintain tissue homeostasis and to prevent tumorigenesis as well as other diseases. In this chapter, we will review molecular mechanisms that regulate YAP and TAZ functions.
2 Regulation by Phosphorylation
Phosphorylation is the most important mechanism known to regulate YAP and TAZ activity. YAP/TAZ are phosphorylated at multiple sites, as indicated by mass spectrometry and mutagenesis analysis, and YAP/TAZ phosphorylation is subjected to dynamic regulation mainly by the Hippo pathway (Zhao et al. 2010b).
Lats1/2 have been shown to directly phosphorylate YAP at Serine 127 (S127) and TAZ at Serine 89 (S89) (Hao et al. 2008; Lei et al. 2008; Zhao et al. 2010b, 2007). Phosphorylation of YAP S127 or TAZ S89 creates a binding site for 14-3-3 (Kanai et al. 2000; Lei et al. 2008; Zhao et al. 2007). The interaction of 14-3-3 with YAP/TAZ sequesters YAP/TAZ in the cytoplasm, which results in the inactivation of YAP/TAZ transcription co-activators (Kanai et al. 2000; Lei et al. 2008; Zhao et al. 2007). In Drosophila, Yki is similarly repressed by wts (lats kinase ortholog) through phosphorylation and 14-3-3 binding (Oh and Irvine 2008; Ren et al. 2010). Consistently, mutation of S127 in YAP or the corresponding serine in TAZ or Yki to alanine increased YAP/TAZ/Yki nuclear localization and activity (Camargo et al. 2007; Lei et al. 2008; Oh and Irvine 2008; Zhao et al. 2007). In search of Drosophila mutants resistant to hpo overexpression, which reduces organ size, Yki mutations were isolated. Interestingly, these gain-of-function Yki mutations were due to the abolishment of either the wts phosphorylation motif or the 14-3-3 binding motif in Yki (Zhao et al. 2007), thus providing convincing genetic evidence for the mechanism of Yki inhibition by wts-dependent phosphorylation.
The substrate specificity of the Lats kinases is defined as an HXRXXS consensus motif (Zhao et al. 2010b). In addition to S127, YAP has four additional HXRXXS sites being phosphorylated by Lats kinases (Fig. 5.1) (Zhao et al. 2010b). Phosphorylation of the other four sites may further inactivate YAP, because a YAP mutant with all five Lats targeting sites mutated (YAP-5SA) is more active in inducing gene expression and promoting cell growth than the S127A single mutant. The 5SA mutant is able to potently transform NIH-3T3 cells when overexpressed (Zhao et al. 2010b). Among the five Lats target sites in YAP, S127 and S381 are most critical, and a YAP with S127A and S381A double mutations is sufficient to transform NIH3T3 cells (Zhao et al. 2010b). There are additional Lats phosphorylation sites on TAZ and Yki as well (Fig. 5.1), and these sites are also important in regulating the activities of TAZ and Yki, respectively (Lei et al. 2008; Ren et al. 2010). Unlike S127 phosphorylation, which mainly exerts its inhibitory effect through 14-3-3 binding, the functions of phosphorylation at additional Lats target sites are less clear (also see below).
Inactivation of YAP or TAZ can be achieved by upregulating the activity of Lats kinases (reviewed by Zhao et al. (2010a)). In addition, phosphatases may antagonize the function of Lats kinases on YAP/TAZ by dephosphorylation. It has been shown that protein phosphatase 1 (PP1) physically interacts with TAZ (Liu et al. 2011). PP1 can promote dephosphorylation of TAZ at S89 and S311, stabilize TAZ, and induce TAZ nuclear localization, which in turn induces transcriptional activity (Liu et al. 2011). The interaction between PP1 and TAZ is strengthened by ASPP2, a known phosphatase regulatory subunit (Liu et al. 2011). Recently, another study has demonstrated a similar role of PP1 in YAP dephosphorylation (Wang et al. 2011a). In addition to PP1, PP2A has also been shown to dephosphorylate YAP in vitro (Schlegelmilch et al. 2011). Although phosphatases have been implicated in YAP/TAZ regulation, it is generally unknown how YAP/TAZ dephosphorylation is regulated and how the action of phosphatases coordinates with Lats kinases.
3 Regulation by Protein Stability
The protein turnover of YAP/TAZ in cells is dependent on both protein synthesis and degradation. The half-life of TAZ is about 1–2 h, while YAP is significantly more stable (Liu et al. 2010; Vigneron et al. 2010). YAP is more stable in low density cells than in high density cultures. When cells are cultured at low density, YAP/TAZ are hypophosphorylated, more stable, and tend to accumulate (Liu et al. 2010; Zhao et al. 2010b), indicating that phosphorylation may play a role in regulating the protein stability of YAP/TAZ.
Phosphorylation on YAP S381 is one of the key phosphorylation events necessary for triggering YAP degradation (Zhao et al. 2010b). Phosphorylation on S381 primes a subsequent phosphorylation on S384 and possibly S387 by another kinase, likely casein kinases 1 (CK1δ/ε, Fig. 5.2) (Zhao et al. 2010b). The amino acid sequence around S384 (DSGLS) is similar to the canonical phosphodegron DpSGXXpS recognized by β-transducin repeat-containing proteins (β-TRCP), a F-box protein which determines selectivity of SCF E3 ubiquitin ligase (Fuchs et al. 2004). Indeed, SCFβ-TRCP physically interacts with YAP, and the interaction is facilitated by phosphorylation on YAP S381, S384, and S387 (Zhao et al. 2010b). The interaction between SCFβ-TRCP and YAP induces YAP ubiquitination and eventually degradation (Zhao et al. 2010b). The phosphorylation of phosphodegron, the interaction between YAP and SCFβ-TRCP, and YAP degradation are all dependent on phosphorylation on S381, suggesting that the sequential posttranslational modifications on YAP are physiologically regulated by the Hippo pathway (Fig. 5.2). In addition, similar to S127A/S381A YAP double mutants, S127A/S384A and S127A/D383A double mutants can also transform NIH-3T3 cells (Zhao et al. 2010b), indicating that the phosphodegron is critical to the oncogenic activity of YAP, probably by regulating YAP protein stability.
Sequencing phosphorylation on YAP/TAZ and β-catenin. The amino acid sequences near phosphodegrons on YAP, TAZ, and β-catenin are aligned. Amino acids representing phosphorylation sites are given in bold face, with exact positions indicated by numbers above or below. Lats1/2, CK, and GSK-3 target sites are highlighted in green, blue, and red, respectively. CK1α phosphorylates S45 of β-catenin, and this phosphorylation will prime a subsequent phosphorylation on S33, S37, and T41 by GSK-3. Similarly, phosphorylation on S381 of YAP by Lats1/2 will prime phosphorylation on S384 and S387 by CK1δ/ε, and phosphorylation on S311 of TAZ by Lats1/2 will prime phosphorylation on S314 by CK1δ/ε. The priming relationships are depicted by dished blue arrows
The protein stability of TAZ is regulated in a manner similar to the regulation of YAP by Lats kinases, CK1, and SCFβ-TRCP (Liu et al. 2010). Phosphorylation of TAZ S311 by Lats kinases primes subsequent phosphorylation on S314 in the phosphodegron by CK1ε and recruitment of the SCFβ-TrCP E3 ubiquitin ligase, thus leading to TAZ ubiquitylation and degradation (Liu et al. 2010). TAZ has an additional phosphodegron at the N-terminal targeted by SCFβ-TrCP E3 ubiquitin ligase, which may contribute to the lower protein stability of TAZ compared to YAP (Huang et al. 2012b; Liu et al. 2010; Tian et al. 2007). Phosphorylation of the C-terminal phosphodegron in TAZ is regulated by Lats, supporting TAZ stability control by the Hippo pathway. However, the N-terminal phosphodegron in TAZ is primarily controlled by GSK-3, and the latter is inhibited by PI3K and AKT pathway (Huang et al. 2012b). Activation of PI3K or PTEN mutation frequently occurs in human cancers, which may lead to AKT activation, GSK-3 inhibition, and TAZ accumulation; upregulation of TAZ protein level may stimulate cell proliferation and contributes to cancer driven by PI3K and PTEN mutations.
The phosphodegron sequence around S381 of YAP is not conserved in Drosophila Yki, indicating a divergence between mammals and Drosophila.
4 Regulation by Subcellular Localization via Protein–Protein Interaction
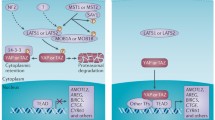
YAP and TAZ are transcription co-activators and are required to enter the nucleus and access target transcription factors and gene promoters to exert their role in gene expression. Due to their oncogenic potential, the nuclear localization of YAP and TAZ is restricted in vivo by multiple mechanisms. As mentioned above, the phosphorylation of S127 on YAP or S89 on TAZ creates a binding site for 14-3-3. Binding with 14-3-3 leads to cytoplasmic retention of YAP/TAZ and prevents their nuclear entry (Kanai et al. 2000; Lei et al. 2008; Zhao et al. 2007). Other than 14-3-3, additional binding partners of YAP/TAZ have been recently identified, and these YAP/TAZ interacting proteins can also modulate YAP/TAZ cellular localizations.
Angiomotin (AMOT) family proteins have recently been identified as a YAP/TAZ interacting protein (Chan et al. 2011; Wang et al. 2011b; Zhao et al. 2011). The interaction is mediated by PP×Y motifs of AMOT and WW domain(s) of YAP/TAZ, and is not directly dependent on YAP/TAZ phosphorylation by Lats kinases (Zhao et al. 2011). AMOT can recruit YAP to different subcellular compartments, such as tight junctions and/or the actin cytoskeleton through physical interaction, thus reducing the translocation of YAP into the nucleus and resulting in decreased YAP activity (Chan et al. 2011; Wang et al. 2011b; Zhao et al. 2011). In addition, AMOT also potentiates YAP/TAZ phosphorylation at Lats target sites (Zhao et al. 2011). Therefore, AMOT may inhibit YAP/TAZ function through a direct binding and an indirect increase of YAP phosphorylation. A recent report shows that AMOT can bind to MST2, Lats2, and YAP, function as a scaffold protein for the core components of the Hippo pathway, and result in increased Lats2 kinase activity thus YAP phosphorylation (Paramasivam et al. 2011). Hippo pathway kinases MST1/2 and Lats1/2 and regulatory protein mob have been shown to be activated at the cell membrane (Hergovich et al. 2006; Ho et al. 2010), and AMOT family proteins may induce the clustering of YAP/TAZ and Hippo pathway kinases at tight junctions in response to cell density to regulate YAP/TAZ phosphorylation and activity (Paramasivam et al. 2011; Zhao et al. 2011).
A role for α-catenin in YAP localization has been recently suggested (Schlegelmilch et al. 2011; Silvis et al. 2011). In keratinocytes, α-catenin strongly co-immunoprecipitates with YAP (Schlegelmilch et al. 2011). However, the interaction between α-catenin and YAP is not direct, and 14-3-3 functions as a mediator for this interaction (Schlegelmilch et al. 2011). Furthermore, only phosphorylated (S127) YAP can form a complex with α-catenin because 14-3-3 recognizes phosphorylated YAP (Schlegelmilch et al. 2011). It is known that α-catenin is a component of adherent junctions; therefore, a tripartite complex of α-catenin, 14-3-3, and YAP may sequester YAP at cell adherent junctions and prevent YAP dephosphorylation, nuclear translocation, and target gene expression. The inhibition of YAP by α-catenin may contribute to the tumor suppressor function of α-catenin (Schlegelmilch et al. 2011; Silvis et al. 2011). Phosphorylated TAZ also interacts with 14-3-3 (Kanai et al. 2000; Lei et al. 2008), but it is unclear if TAZ localization is also regulated by α-catenin. Moreover, whether the inhibitory role of α-catenin on YAP is conserved in tissues other than skin remains unknown.
Other components or regulators of cell junctions such as ZO-1, ZO-2, and PTPN14 have also been suggested as regulators of YAP/TAZ localization and activity (Huang et al. 2012a; Liu et al. 2012b; Oka et al. 2010, 2012), suggesting that sequestration of YAP/TAZ at cell junctions is a common mechanism to restrict the growth-promoting activity of YAP/TAZ.
Yki also physically interacts with upstream components of the Hippo pathway, such as Expanded, Wts, and Hpo; these interactions will restrict Yki activity by restraining Yki in the cytoplasm (Badouel et al. 2009; Oh et al. 2009). However, an ortholog of AMOT is not present in Drosophila and thus AMOT-dependent regulation of YAP/TAZ localization is not evolutionary conserved.
5 Regulation by Transcription Factor Target Selection
YAP/TAZ are both transcription co-activators without DNA-binding ability. In order to induce gene transcription, they must interact with specific transcription factors that bind to promoters of target genes. Target transcription factors selection therefore provides an additional layer of complexity to YAP/TAZ regulation.
TEAD family transcription factors (TEAD1-4) have been shown to serve as the major target transcription factors mediating the biological functions of YAP/TAZ (Vassilev et al. 2001; Zhao et al. 2008). In a functional screen of a human transcription factor library, TEADs were identified as transcriptional factors most potently activated by YAP (Zhao et al. 2008). Indeed, TEAD1/2 and YAP share a largely overlapping set of target genes (Ota and Sasaki 2008; Zhao et al. 2008). Downregulation of TEADs or disruption of the YAP–TEAD interaction blunts the expression of most YAP targeting genes and largely diminishes the ability of YAP to promote cell proliferation, cell transformation, EMT, cell contact inhibition, and maintenance of stem cell pluripotency (Lian et al. 2010; Ota and Sasaki 2008; Schlegelmilch et al. 2011; Zhao et al. 2008). In addition, TEAD1/2 double knockout mice exhibit reduced cell proliferation and enhanced apoptosis (Sawada et al. 2008), and these phenotypes are similar to those of YAP knockout mice (Zhang et al. 2010). Similarly, TEADs also interact with TAZ and mediate the function of TAZ on cell growth and EMT (Chan et al. 2009; Mahoney et al. 2005; Zhang et al. 2009). These findings indicate that TEADs serve as the major transcription factors mediating the function of YAP/TAZ in gene expression and organ size control.
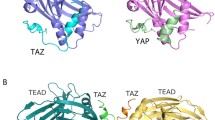
YAP and TEAD form a strong physical interaction, and the detailed molecular mechanism of YAP–TEAD interaction is revealed by structural studies. Three-dimensional structures of a human YAP and TEAD1 complex (Li et al. 2010) and a mouse YAP and TEAD4 complex (Chen et al. 2010) have been resolved, although both studies used the YAP-binding domain of TEAD and the TEAD-binding domain of YAP rather than full length proteins. The complex structures indicate that the C-terminal domain of TEAD forms a globular structure with a β-sandwich fold surrounded by four α-helices on one side, with the N-terminal domain of YAP wrapping around the TEAD to form extensive interactions (Chen et al. 2010; Li et al. 2010). The crystal structure of the YAP-binding domain of human TEAD2 has also been resolved, and it adopts an immunoglobulin-like β-sandwich fold with two extra helix-turn-helix inserts (Tian et al. 2010).
The YAP–TEAD complex structures clearly show that YAP S94 directly forms a hydrogen bond with TEAD1 Y406 (Chen et al. 2010; Li et al. 2010). This provides a beautiful molecular explanation of the disrupted interaction between YAP and TEAD1 by either YAP S94 or TEAD1 Y406 mutations (Zhao et al. 2008). The TEAD1 Y406H mutation is causal to Sveinsson’s chorioretinal atrophy, also referred to as helicoid peripapillary chorioretinal degeneration (Fossdal et al. 2004). Therefore, the structural and biochemical studies revealed that a disruption of YAP–TEAD1 interaction due to mutations in TEAD1 might be the underlying molecular basis for this human genetic disorder. The YAP–TEAD interaction requires only a small region of YAP; short peptides and small molecules have been shown to disrupt the interaction and reduce YAP activity in vivo (Liu-Chittenden et al. 2012; von Gise et al. 2012), and inhibitors targeting YAP/TAZ–TEAD interactions provide new therapeutic strategies to fight diseases caused by dysregulated YAP/TAZ activity, such as cancers.
Smad1, a transcription factor in the BMP signaling pathway, has been reported as YAP-interacting protein (Alarcon et al. 2009). The interaction between Smad1 and YAP is dependent on BMP signaling; following BMP stimulation, the linker region of Smad1 undergoes phosphorylation, and phosphorylated Smad1 then interacts with YAP via WW domains (Alarcon et al. 2009). The phosphorylation at the linker region of Smad1 is required for Smad1–YAP interaction, as the interaction is decreased when the phosphorylation sites are mutated (Alarcon et al. 2009). YAP has also been shown to mediate BMP target gene expression in mouse embryonic stem cells, which rely on BMP signaling for maintenance of pluripotency (Alarcon et al. 2009). In consistence, a critical role of YAP in maintaining pluripotency of mouse embryonic stem cells has been reported (Lian et al. 2010).
Smad2 and Smad3, two transcription factors in the TGFβ signaling pathway, can bind to the coiled-coil domain of TAZ (Varelas et al. 2008). TAZ also interacts with MED15, a component of the mediator complex important for gene transcription (Varelas et al. 2008). In a TGFβ signaling sensitive manner, TAZ recruits both the Smad2/3/4 complex and mediator complex to promoters of TGFβ target genes to induce transcription (Varelas et al. 2008). Knockdown of TAZ not only impairs TGFβ-induced gene expression but also promotes human stem cell differentiation (Varelas et al. 2008), suggesting that TAZ is required for the TGFβ signaling to maintain stem cell pluripotency.
YAP can also interact with Smad7, a Smad protein that inhibits both TGFβ and BMP signaling, and the interaction is mediated by the PPxY motif on Smad7 and WW domain on YAP (Ferrigno et al. 2002). Though both WW and coiled-coil domains are largely conserved in YAP and TAZ, it would be interesting to know how YAP and TAZ can interact with different Smad proteins regulated by BMP or TGFβ. A more detailed study on the relationship among YAP, TAZ, and different Smad proteins is required to address this issue. Nevertheless, these studies suggest YAP and TAZ may function in a distinct manner in human and mouse stem cells.
Besides TEADs and Smad proteins, RUNX1/2 has also been shown to interact with YAP/TAZ (Yagi et al. 1999). In addition, YAP has been shown to interact with p63, p73, and ErbB4 (Komuro et al. 2003; Omerovic et al. 2004; Strano et al. 2001). These interactions may regulate transcription of diverse genes related to cell proliferation and development.
In Drosophila, Sd (TEAD ortholog) genetically and physically interacts with Yki and is required for Yki-induced gene expression and tissue overgrowth (Goulev et al. 2008; Wu et al. 2008; Zhang et al. 2008; Zhao et al. 2008). Although Sd is a major transcription factor mediating Yki function, the Yki mutant has a more dramatic growth defect than the Sd mutant and regulates expression of a broader range of genes than Sd (Huang et al. 2005; Wu et al. 2008), suggesting additional transcription factors acting downstream of Yki. Indeed, other transcription factors such as Mad or a complex of homothrorax (Hth) and Teashirt (Tsh) have been shown to mediate part of Yki activity in inducing microRNA bantam (Nolo et al. 2006; Oh and Irvine 2011; Peng et al. 2009; Thompson and Cohen 2006). It is clear that TEAD/Sd are key downstream transcription factors of YAP/TAZ and Yki. However, whether other transcription factors, such as Smad and p63, truly mediate the biological functions of YAP/TAZ requires further investigation. In addition, whether mammalian homologs of Hth and Tsh are involved in YAP/TAZ biology also waits to be tested.
6 Similarities Between YAP and β-Catenin Regulation
Representing the primary downstream effector of the Wnt signaling pathway, β-catenin also functions as a transcription co-activator and plays key roles in normal development and malignant transformation, and its activity is regulated at multiple layers similar to those in YAP/TAZ regulations.
Similar to YAP/TAZ, β-catenin is mainly regulated by protein phosphorylation, stability, and localization (MacDonald et al. 2009). In the absence of upstream Wnt signals, β-catenin is phosphorylated by a protein complex containing GSK-3, axin, and adenomatous polyposis coli (APC), and this phosphorylation promotes proteolytic degradation of β-catenin (MacDonald et al. 2009). Under Wnt stimulation the kinase activity of GSK-3 is inhibited causing cytoplasmic β-catenin to be hypophosphorylated, stabilized, and translocated into the nucleus. At the nucleus β-catenin induces target gene transcription by interacting with TCF/LEF family transcription factors (MacDonald et al. 2009). The signal transduction from GSK-3 to β-catenin to TCF/LEF is highly homologous to the pathway from Lats1/2 to YAP/TAZ to TEAD.
YAP/TAZ and β-catenin also share the same E3 ubiquitin ligase SCFβ-TRCP for ubiquitination and degradation (Clevers 2006). Binding between β-catenin and SCFβ-TRCP depends strictly on multistep phosphorylation of the phosphodegron involving CK1α and GSK-3, in which CK1α phosphorylates S45 primes subsequent phosphorylation on S33, S37, and S41 by GSK-3 (Liu et al. 2002). Clearly, the sequential phosphorylation, ubiquitination, and protein degradation is a common strategy for regulating stability of YAP/TAZ and β-catenin.
YAP/TAZ are recruited to cell junction structures and exhibit extensive interactions with different cell junction proteins, especially at high cell densities (see above). β-catenin is well known as a structural component of adherent junctions, and is important for mediating cell adhesion and linking cadherins to the actin cytoskeleton (Gumbiner 1995). Retention at cell junctions might be a common mechanism for regulating functions of YAP/TAZ and β-catenin.
Accumulation of both YAP/TAZ and β-catenin oncoproteins has been reported in human cancers (reviewed in Clevers 2006; Zhao et al. 2010a). Upstream kinases and kinase-associated scaffolds of YAP/TAZ or β-catenin function as tumor suppressors, and downregulation or inactivation of these tumor suppressors may cause cancer via activation of YAP/TAZ or β-catenin. Indeed, downregulated MST1/2 and Lats1/2 expression, mutations of Sav1 or Mob1 (Chakraborty et al. 2007; Hisaoka et al. 2002; Jimenez-Velasco et al. 2005; Zhao et al. 2012), and mutations of APC and axin (Liu et al. 2000; Rubinfeld et al. 1996) have been reported in different types of human cancers. Therefore, there is an astonishingly high similarity between the regulation of YAP/TAZ and β-catenin. Experience from β-catenin research may help us study the regulatory mechanisms and functions of YAP/TAZ in the future.
The Hippo pathway also crosstalks with the Wnt pathway. Cytoplasmic TAZ can bind to and interfere with the phosphorylation of disheveled (Dvl), leading to β-catenin degradation (Varelas et al. 2010a). Another report shows that both YAP/TAZ can interact with β-catenin and prevent translocation of β-catenin into the nucleus (Imajo et al. 2012). In both cases, decreased Hippo signaling leads to nuclear accumulation and activation of β-catenin and YAP/TAZ. When YAP is overexpressed in mouse intestinal epithelium, total and nuclear β-catenin is increased (Camargo et al. 2007). In addition, heart-specific inactivation of Sav in mice increases heart size and at the same time enhances Wnt signaling (Heallen et al. 2011). These in vivo observations again support a positive role of YAP/TAZ on β-catenin activation.
Interesting genetic and biochemical studies within the last 10 years have revealed that the Hippo pathway plays a major role in organ size control, and that dysregulation of this pathway contributes to either tumor growth or atrophy. YAP/TAZ co-activator inhibition represents the primary outcome of the Hippo pathway, which is accomplished through a phosphorylation-dependent cytoplasmic retention and degradation. However, not much is known about upstream regulators of the Hippo pathway. Future studies on Hippo pathway signaling cascade will lead to a better understanding of organ size control and pathobiology of tumorigenesis.
References
Alarcon C, Zaromytidou AI, Xi Q, Gao S, Yu J, Fujisawa S, et al. Nuclear CDKs drive Smad transcriptional activation and turnover in BMP and TGF-beta pathways. Cell. 2009;139:757–69.
Badouel C, Gardano L, Amin N, Garg A, Rosenfeld R, Le Bihan T, et al. The FERM-domain protein expanded regulates Hippo pathway activity via direct interactions with the transcriptional activator Yorkie. Dev Cell. 2009;16:411–20.
Benhamouche S, Curto M, Saotome I, Gladden AB, Liu CH, Giovannini M, et al. Nf2/Merlin controls progenitor homeostasis and tumorigenesis in the liver. Genes Dev. 2010;24:1718–30.
Callus BA, Verhagen AM, Vaux DL. Association of mammalian sterile twenty kinases, Mst1 and Mst2, with hSalvador via C-terminal coiled-coil domains, leads to its stabilization and phosphorylation. FEBS J. 2006;273:4264–76.
Camargo FD, Gokhale S, Johnnidis JB, Fu D, Bell GW, Jaenisch R, et al. YAP1 increases organ size and expands undifferentiated progenitor cells. Curr Biol. 2007;17:2054–60.
Cao X, Pfaff SL, Gage FH. YAP regulates neural progenitor cell number via the TEA domain transcription factor. Genes Dev. 2008;22:3320–34.
Chakraborty S, Khare S, Dorairaj SK, Prabhakaran VC, Prakash DR, Kumar A. Identification of genes associated with tumorigenesis of retinoblastoma by microarray analysis. Genomics. 2007;90:344–53.
Chan EH, Nousiainen M, Chalamalasetty RB, Schafer A, Nigg EA, Sillje HH. The Ste20-like kinase Mst2 activates the human large tumor suppressor kinase Lats1. Oncogene. 2005;24:2076–86.
Chan SW, Lim CJ, Guo K, Ng CP, Lee I, Hunziker W, et al. A role for TAZ in migration, invasion, and tumorigenesis of breast cancer cells. Cancer Res. 2008;68:2592–8.
Chan SW, Lim CJ, Loo LS, Chong YF, Huang C, Hong W. TEADs mediate nuclear retention of TAZ to promote oncogenic transformation. J Biol Chem. 2009;284:14347–58.
Chan SW, Lim CJ, Chong YF, Pobbati AV, Huang C, Hong W. Hippo pathway-independent restriction of TAZ and YAP by angiomotin. J Biol Chem. 2011;286:7018–26.
Chen L, Chan SW, Zhang X, Walsh M, Lim CJ, Hong W, et al. Structural basis of YAP recognition by TEAD4 in the hippo pathway. Genes Dev. 2010;24:290–300.
Chow A, Hao Y, Yang X. Molecular characterization of human homologs of yeast MOB1. Int J Cancer. 2010;126:2079–89.
Clevers H. Wnt/beta-catenin signaling in development and disease. Cell. 2006;127:469–80.
Dong J, Feldmann G, Huang J, Wu S, Zhang N, Comerford SA, et al. Elucidation of a universal size-control mechanism in Drosophila and mammals. Cell. 2007;130:1120–33.
Ferrigno O, Lallemand F, Verrecchia F, L’Hoste S, Camonis J, Atfi A, et al. Yes-associated protein (YAP65) interacts with Smad7 and potentiates its inhibitory activity against TGF-beta/Smad signaling. Oncogene. 2002;21:4879–84.
Fossdal R, Jonasson F, Kristjansdottir GT, Kong A, Stefansson H, Gosh S, et al. A novel TEAD1 mutation is the causative allele in Sveinsson’s chorioretinal atrophy (helicoid peripapillary chorioretinal degeneration). Hum Mol Genet. 2004;13:975–81.
Fuchs SY, Spiegelman VS, Kumar KG. The many faces of beta-TrCP E3 ubiquitin ligases: reflections in the magic mirror of cancer. Oncogene. 2004;23:2028–36.
Goulev Y, Fauny JD, Gonzalez-Marti B, Flagiello D, Silber J, Zider A. SCALLOPED interacts with YORKIE, the nuclear effector of the hippo tumor-suppressor pathway in Drosophila. Curr Biol. 2008;18:435–41.
Gumbiner BM. Signal transduction of beta-catenin. Curr Opin Cell Biol. 1995;7:634–40.
Hao Y, Chun A, Cheung K, Rashidi B, Yang X. Tumor suppressor LATS1 is a negative regulator of oncogene YAP. J Biol Chem. 2008;283:5496–509.
Harvey KF, Pfleger CM, Hariharan IK. The Drosophila Mst ortholog, hippo, restricts growth and cell proliferation and promotes apoptosis. Cell. 2003;114:457–67.
Heallen T, Zhang M, Wang J, Bonilla-Claudio M, Klysik E, Johnson RL, et al. Hippo pathway inhibits Wnt signaling to restrain cardiomyocyte proliferation and heart size. Science. 2011;332:458–61.
Hergovich A, Stegert MR, Schmitz D, Hemmings BA. NDR kinases regulate essential cell processes from yeast to humans. Nat Rev Mol Cell Biol. 2006;7:253–64.
Hisaoka M, Tanaka A, Hashimoto H. Molecular alterations of h-warts/LATS1 tumor suppressor in human soft tissue sarcoma. Lab Invest. 2002;82:1427–35.
Ho LL, Wei X, Shimizu T, Lai ZC. Mob as tumor suppressor is activated at the cell membrane to control tissue growth and organ size in Drosophila. Dev Biol. 2010;337:274–83.
Hossain Z, Ali SM, Ko HL, Xu J, Ng CP, Guo K, et al. Glomerulocystic kidney disease in mice with a targeted inactivation of Wwtr1. Proc Natl Acad Sci U S A. 2007;104:1631–6.
Huang J, Wu S, Barrera J, Matthews K, Pan D. The Hippo signaling pathway coordinately regulates cell proliferation and apoptosis by inactivating Yorkie, the Drosophila Homolog of YAP. Cell. 2005;122:421–34.
Huang JM, Nagatomo I, Suzuki E, Mizuno T, Kumagai T, Berezov A, et al. YAP modifies cancer cell sensitivity to EGFR and survivin inhibitors and is negatively regulated by the non-receptor type protein tyrosine phosphatase 14. Oncogene. 2012a. doi:10.1038/onc.2012.231.
Huang W, Lv X, Liu C, Zha Z, Zhang H, Jiang Y, et al. The N-terminal phosphodegron targets TAZ/WWTR1 for SCFbeta-TrCP dependent degradation in response to PI3K inhibition. J Biol Chem. 2012b;287(31):26245–53.
Imajo M, Miyatake K, Iimura A, Miyamoto A, Nishida E. A molecular mechanism that links Hippo signalling to the inhibition of Wnt/beta-catenin signalling. EMBO J. 2012;31:1109–22.
Jia J, Zhang W, Wang B, Trinko R, Jiang J. The Drosophila Ste20 family kinase dMST functions as a tumor suppressor by restricting cell proliferation and promoting apoptosis. Genes Dev. 2003;17:2514–9.
Jimenez-Velasco A, Roman-Gomez J, Agirre X, Barrios M, Navarro G, Vazquez I, et al. Downregulation of the large tumor suppressor 2 (LATS2/KPM) gene is associated with poor prognosis in acute lymphoblastic leukemia. Leukemia. 2005;19:2347–50.
Justice RW, Zilian O, Woods DF, Noll M, Bryant PJ. The Drosophila tumor suppressor gene warts encodes a homolog of human myotonic dystrophy kinase and is required for the control of cell shape and proliferation. Genes Dev. 1995;9:534–46.
Kanai F, Marignani PA, Sarbassova D, Yagi R, Hall RA, Donowitz M, et al. TAZ: a novel transcriptional co-activator regulated by interactions with 14-3-3 and PDZ domain proteins. EMBO J. 2000;19:6778–91.
Kango-Singh M, Nolo R, Tao C, Verstreken P, Hiesinger PR, Bellen HJ, et al. Shar-pei mediates cell proliferation arrest during imaginal disc growth in Drosophila. Development. 2002;129:5719–30.
Komuro A, Nagai M, Navin NE, Sudol M. WW domain-containing protein YAP associates with ErbB-4 and acts as a co-transcriptional activator for the carboxyl-terminal fragment of ErbB-4 that translocates to the nucleus. J Biol Chem. 2003;278:33334–41.
Lai ZC, Wei X, Shimizu T, Ramos E, Rohrbaugh M, Nikolaidis N, et al. Control of cell proliferation and apoptosis by mob as tumor suppressor, mats. Cell. 2005;120:675–85.
Lee JH, Kim TS, Yang TH, Koo BK, Oh SP, Lee KP, et al. A crucial role of WW45 in developing epithelial tissues in the mouse. EMBO J. 2008;27:1231–42.
Lee KP, Lee JH, Kim TS, Kim TH, Park HD, Byun JS, et al. The Hippo-Salvador pathway restrains hepatic oval cell proliferation, liver size, and liver tumorigenesis. Proc Natl Acad Sci U S A. 2010;107:8248–53.
Lei QY, Zhang H, Zhao B, Zha ZY, Bai F, Pei XH, et al. TAZ promotes cell proliferation and epithelial-mesenchymal transition and is inhibited by the hippo pathway. Mol Cell Biol. 2008;28:2426–36.
Li Z, Zhao B, Wang P, Chen F, Dong Z, Yang H, et al. Structural insights into the YAP and TEAD complex. Genes Dev. 2010;24:235–40.
Lian I, Kim J, Okazawa H, Zhao J, Zhao B, Yu J, et al. The role of YAP transcription coactivator in regulating stem cell self-renewal and differentiation. Genes Dev. 2010;24:1106–18.
Liu W, Dong X, Mai M, Seelan RS, Taniguchi K, Krishnadath KK, et al. Mutations in AXIN2 cause colorectal cancer with defective mismatch repair by activating beta-catenin/TCF signalling. Nat Genet. 2000;26:146–7.
Liu C, Li Y, Semenov M, Han C, Baeg GH, Tan Y, et al. Control of beta-catenin phosphorylation/degradation by a dual-kinase mechanism. Cell. 2002;108:837–47.
Liu CY, Zha ZY, Zhou X, Zhang H, Huang W, Zhao D, et al. The hippo tumor pathway promotes TAZ degradation by phosphorylating a phosphodegron and recruiting the SCF{beta}-TrCP E3 ligase. J Biol Chem. 2010;285:37159–69.
Liu CY, Lv X, Li T, Xu Y, Zhou X, Zhao S, et al. PP1 cooperates with ASPP2 to dephosphorylate and activate TAZ. J Biol Chem. 2011;286:5558–66.
Liu H, Jiang D, Chi F, Zhao B. The Hippo pathway regulates stem cell proliferation, self-renewal, and differentiation. Protein Cell. 2012a;3:291–304.
Liu X, Yang N, Figel SA, Wilson KE, Morrison CD, Gelman IH, et al. PTPN14 interacts with and negatively regulates the oncogenic function of YAP. Oncogene. 2012b. doi:10.1038/onc.2012.147.
Liu-Chittenden Y, Huang B, Shim JS, Chen Q, Lee SJ, Anders RA, et al. Genetic and pharmacological disruption of the TEAD-YAP complex suppresses the oncogenic activity of YAP. Genes Dev. 2012;26(12):1300–5.
Lu L, Li Y, Kim SM, Bossuyt W, Liu P, Qiu Q, et al. Hippo signaling is a potent in vivo growth and tumor suppressor pathway in the mammalian liver. Proc Natl Acad Sci U S A. 2010;107:1437–42.
MacDonald BT, Tamai K, He X. Wnt/beta-catenin signaling: components, mechanisms, and diseases. Dev Cell. 2009;17:9–26.
Mahoney Jr WM, Hong JH, Yaffe MB, Farrance IK. The transcriptional co-activator TAZ interacts differentially with transcriptional enhancer factor-1 (TEF-1) family members. Biochem J. 2005;388:217–25.
Makita R, Uchijima Y, Nishiyama K, Amano T, Chen Q, Takeuchi T, et al. Multiple renal cysts, urinary concentration defects, and pulmonary emphysematous changes in mice lacking TAZ. Am J Physiol Renal Physiol. 2008;294:F542–53.
Mauviel A, Nallet-Staub F, Varelas X. Integrating developmental signals: a Hippo in the (path)way. Oncogene. 2012;31:1743–56.
Morin-Kensicki EM, Boone BN, Howell M, Stonebraker JR, Teed J, Alb JG, et al. Defects in yolk sac vasculogenesis, chorioallantoic fusion, and embryonic axis elongation in mice with targeted disruption of Yap65. Mol Cell Biol. 2006;26:77–87.
Nishioka N, Inoue K, Adachi K, Kiyonari H, Ota M, Ralston et al. The Hippo signaling pathway components Lats and Yap pattern Tead4 activity to distinguish mouse trophectoderm from inner cell mass. Dev Cell. 2009;16(3):398–410.
Nolo R, Morrison CM, Tao C, Zhang X, Halder G. The bantam microRNA is a target of the hippo tumor-suppressor pathway. Curr Biol. 2006;16:1895–904.
Oh H, Irvine KD. In vivo regulation of Yorkie phosphorylation and localization. Development. 2008;135:1081–8.
Oh H, Irvine KD. Cooperative regulation of growth by Yorkie and Mad through bantam. Dev Cell. 2011;20:109–22.
Oh H, Reddy BV, Irvine KD. Phosphorylation-independent repression of Yorkie in Fat-Hippo signaling. Dev Biol. 2009;335:188–97.
Oka T, Remue E, Meerschaert K, Vanloo B, Boucherie C, Gfeller D, et al. Functional complexes between YAP2 and ZO-2 are PDZ domain-dependent, and regulate YAP2 nuclear localization and signalling. Biochem J. 2010;432:461–72.
Oka T, Schmitt AP, Sudol M. Opposing roles of angiomotin-like-1 and zona occludens-2 on pro-apoptotic function of YAP. Oncogene. 2012;31:128–34.
Omerovic J, Puggioni EM, Napoletano S, Visco V, Fraioli R, Frati L, et al. Ligand-regulated association of ErbB-4 to the transcriptional co-activator YAP65 controls transcription at the nuclear level. Exp Cell Res. 2004;294:469–79.
Ota M, Sasaki H. Mammalian Tead proteins regulate cell proliferation and contact inhibition as transcriptional mediators of Hippo signaling. Development. 2008;135:4059–69.
Overholtzer M, Zhang J, Smolen GA, Muir B, Li W, Sgroi DC, et al. Transforming properties of YAP, a candidate oncogene on the chromosome 11q22 amplicon. Proc Natl Acad Sci U S A. 2006;103:12405–10.
Pantalacci S, Tapon N, Leopold P. The Salvador partner Hippo promotes apoptosis and cell-cycle exit in Drosophila. Nat Cell Biol. 2003;5:921–7.
Paramasivam M, Sarkeshik A, Yates III JR, Fernandes MJ, McCollum D. Angiomotin family proteins are novel activators of the LATS2 kinase tumor suppressor. Mol Biol Cell. 2011;22:3725–33.
Peng HW, Slattery M, Mann RS. Transcription factor choice in the Hippo signaling pathway: homothorax and yorkie regulation of the microRNA bantam in the progenitor domain of the Drosophila eye imaginal disc. Genes Dev. 2009;23:2307–19.
Qin H, Blaschke K, Wei G, Ohi Y, Blouin L, Qi Z, et al. Transcriptional analysis of pluripotency reveals the Hippo pathway as a barrier to reprogramming. Hum Mol Genet. 2012;21:2054–67.
Ren F, Zhang L, Jiang J. Hippo signaling regulates Yorkie nuclear localization and activity through 14-3-3 dependent and independent mechanisms. Dev Biol. 2010;337:303–12.
Rouleau GA, Merel P, Lutchman M, Sanson M, Zucman J, Marineau C, et al. Alteration in a new gene encoding a putative membrane-organizing protein causes neuro-fibromatosis type 2. Nature. 1993;363:515–21.
Rubinfeld B, Albert I, Porfiri E, Fiol C, Munemitsu S, Polakis P. Binding of GSK3beta to the APC-beta-catenin complex and regulation of complex assembly. Science. 1996;272:1023–6.
Ruttledge MH, Sarrazin J, Rangaratnam S, Phelan CM, Twist E, Merel P, et al. Evidence for the complete inactivation of the NF2 gene in the majority of sporadic meningiomas. Nat Genet. 1994;6:180–4.
Sawada A, Kiyonari H, Ukita K, Nishioka N, Imuta Y, Sasaki H. Redundant roles of Tead1 and Tead2 in notochord development and the regulation of cell proliferation and survival. Mol Cell Biol. 2008;28:3177–89.
Schlegelmilch K, Mohseni M, Kirak O, Pruszak J, Rodriguez JR, Zhou D, et al. Yap1 acts downstream of alpha-catenin to control epidermal proliferation. Cell. 2011;144:782–95.
Silvis MR, Kreger BT, Lien WH, Klezovitch O, Rudakova GM, Camargo FD, et al. Alpha-catenin is a tumor suppressor that controls cell1 accumulation by regulating the localization and activity of the transcriptional coactivator Yap1. Sci Signal. 2011;4:ra33.
Song H, Mak KK, Topol L, Yun K, Hu J, Garrett L, et al. Mammalian Mst1 and Mst2 kinases play essential roles in organ size control and tumor suppression. Proc Natl Acad Sci U S A. 2010;107:1431–6.
Steinhardt AA, Gayyed MF, Klein AP, Dong J, Maitra A, Pan D, et al. Expression of Yes-associated protein in common solid tumors. Hum Pathol. 2008;39:1582–9.
Strano S, Munarriz E, Rossi M, Castagnoli L, Shaul Y, Sacchi A, et al. Physical interaction with Yes-associated protein enhances p73 transcriptional activity. J Biol Chem. 2001;276:15164–73.
Tamm C, Bower N, Anneren C. Regulation of mouse embryonic stem cell self-renewal by a Yes-YAP-TEAD2 signaling pathway downstream of LIF. J Cell Sci. 2011;124:1136–44.
Tapon N, Harvey KF, Bell DW, Wahrer DC, Schiripo TA, Haber DA, et al. salvador Promotes both cell cycle exit and apoptosis in Drosophila and is mutated in human cancer cell lines. Cell. 2002;110:467–78.
Thompson BJ, Cohen SM. The Hippo pathway regulates the bantam microRNA to control cell proliferation and apoptosis in Drosophila. Cell. 2006;126:767–74.
Tian Y, Kolb R, Hong JH, Carroll J, Li D, You J, et al. TAZ promotes PC2 degradation through a SCFbeta-Trcp E3 ligase complex. Mol Cell Biol. 2007;27:6383–95.
Tian W, Yu J, Tomchick DR, Pan D, Luo X. Structural and functional analysis of the YAP-binding domain of human TEAD2. Proc Natl Acad Sci U S A. 2010;107:7293–8.
Varelas X, Sakuma R, Samavarchi-Tehrani P, Peerani R, Rao BM, Dembowy J, et al. TAZ controls Smad nucleocytoplasmic shuttling and regulates human embryonic stem-cell self-renewal. Nat Cell Biol. 2008;10:837–48.
Varelas X, Miller BW, Sopko R, Song S, Gregorieff A, Fellouse FA, et al. The Hippo pathway regulates Wnt/beta-catenin signaling. Dev Cell. 2010a;18:579–91.
Varelas X, Samavarchi-Tehrani P, Narimatsu M, Weiss A, Cockburn K, Larsen BG, et al. The Crumbs complex couples cell density sensing to Hippo-dependent control of the TGF-beta-SMAD pathway. Dev Cell. 2010b;19:831–44.
Vassilev A, Kaneko KJ, Shu H, Zhao Y, DePamphilis ML. TEAD/TEF transcription factors utilize the activation domain of YAP65, a Src/Yes-associated protein localized in the cytoplasm. Genes Dev. 2001;15:1229–41.
Vigneron AM, Ludwig RL, Vousden KH. Cytoplasmic ASPP1 inhibits apoptosis through the control of YAP. Genes Dev. 2010;24:2430–9.
von Gise A, Lin Z, Schlegelmilch K, Honor LB, Pan GM, Buck JN, et al. YAP1, the nuclear target of Hippo signaling, stimulates heart growth through cardiomyocyte proliferation but not hypertrophy. Proc Natl Acad Sci U S A. 2012;109:2394–9.
Wang P, Bai Y, Song B, Wang Y, Liu D, Lai Y, et al. PP1A-mediated dephosphorylation positively regulates YAP2 activity. PLoS One. 2011a;6:e24288.
Wang W, Huang J, Chen J. Angiomotin-like proteins associate with and negatively regulate YAP1. J Biol Chem. 2011b;286:4364–70.
Wu S, Huang J, Dong J, Pan D. Hippo encodes a Ste-20 family protein kinase that restricts cell proliferation and promotes apoptosis in conjunction with salvador and warts. Cell. 2003;114:445–56.
Wu S, Liu Y, Zheng Y, Dong J, Pan D. The TEAD/TEF family protein Scalloped mediates transcriptional output of the Hippo growth-regulatory pathway. Dev Cell. 2008;14:388–98.
Xu T, Wang W, Zhang S, Stewart RA, Yu W. Identifying tumor suppressors in genetic mosaics: the Drosophila lats gene encodes a putative protein kinase. Development. 1995;121:1053–63.
Yagi R, Chen LF, Shigesada K, Murakami Y, Ito Y. A WW domain-containing yes-associated protein (YAP) is a novel transcriptional co-activator. EMBO J. 1999;18:2551–62.
Zender L, Spector MS, Xue W, Flemming P, Cordon-Cardo C, Silke J, et al. Identification and validation of oncogenes in liver cancer using an integrative oncogenomic approach. Cell. 2006;125:1253–67.
Zhang L, Ren F, Zhang Q, Chen Y, Wang B, Jiang J. The TEAD/TEF family of transcription factor Scalloped mediates Hippo signaling in organ size control. Dev Cell. 2008;14:377–87.
Zhang H, Liu CY, Zha ZY, Zhao B, Yao J, Zhao S, et al. TEAD transcription factors mediate the function of TAZ in cell growth and epithelial-mesenchymal transition. J Biol Chem. 2009;284:13355–62.
Zhang N, Bai H, David KK, Dong J, Zheng Y, Cai J, et al. The Merlin/NF2 tumor suppressor functions through the YAP oncoprotein to regulate tissue homeostasis in mammals. Dev Cell. 2010;19:27–38.
Zhang H, Pasolli HA, Fuchs E. Yes-associated protein (YAP) transcriptional coactivator functions in balancing growth and differentiation in skin. Proc Natl Acad Sci U S A. 2011;108:2270–5.
Zhao B, Wei X, Li W, Udan RS, Yang Q, Kim J, et al. Inactivation of YAP oncoprotein by the Hippo pathway is involved in cell contact inhibition and tissue growth control. Genes Dev. 2007;21:2747–61.
Zhao B, Ye X, Yu J, Li L, Li W, Li S, et al. TEAD mediates YAP-dependent gene induction and growth control. Genes Dev. 2008;22:1962–71.
Zhao B, Li L, Lei Q, Guan KL. The Hippo-YAP pathway in organ size control and tumorigenesis: an updated version. Genes Dev. 2010a;24:862–74.
Zhao B, Li L, Tumaneng K, Wang CY, Guan KL. A coordinated phosphorylation by Lats and CK1 regulates YAP stability through SCF(beta-TRCP). Genes Dev. 2010b;24:72–85.
Zhao B, Li L, Lu Q, Wang LH, Liu CY, Lei Q, et al. Angiomotin is a novel Hippo pathway component that inhibits YAP oncoprotein. Genes Dev. 2011;25:51–63.
Zhao B, Li L, Wang L, Wang CY, Yu J, Guan KL. Cell detachment activates the Hippo pathway via cytoskeleton reorganization to induce anoikis. Genes Dev. 2012;26:54–68.
Zhou D, Conrad C, Xia F, Park JS, Payer B, Yin Y, et al. Mst1 and Mst2 maintain hepatocyte quiescence and suppress hepatocellular carcinoma development through inactivation of the Yap1 oncogene. Cancer Cell. 2009;16:425–38.
Zhou D, Zhang Y, Wu H, Barry E, Yin Y, Lawrence E, et al. Mst1 and Mst2 protein kinases restrain intestinal stem cell proliferation and colonic tumorigenesis by inhibition of Yes-associated protein (Yap) overabundance. Proc Natl Acad Sci U S A. 2011;108:E1312–20.
Acknowledgments
We apologize for being unable to cite some primary work due to space limitations. We would like to thank Jenna Jewell and Jessica Zhou for critical reading of this manuscript. This work was supported by grants from NIH and CIRM to K.-L.G.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2013 Springer Science+Business Media New York
About this chapter
Cite this chapter
Yu, FX., Zhao, B., Guan, KL. (2013). Regulation of YAP and TAZ Transcription Co-activators. In: Oren, M., Aylon, Y. (eds) The Hippo Signaling Pathway and Cancer. Springer, New York, NY. https://doi.org/10.1007/978-1-4614-6220-0_5
Download citation
DOI: https://doi.org/10.1007/978-1-4614-6220-0_5
Published:
Publisher Name: Springer, New York, NY
Print ISBN: 978-1-4614-6219-4
Online ISBN: 978-1-4614-6220-0
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)