Abstract
Microorganisms produce a diverse range of carbohydrates, including cytoplasmic storage polymers (glycogens) and structural polymers (glycans) that make up a portion of the microbial envelope. Glycan polymers include capsular polysaccharides (if polymer tightly bound to the cell surface) or exopolysaccharides (if loosely attached to the extracellular surface). Many Gram-positive and Gram-negative bacteria have been found to have capsular polysaccharides as virulence features. Therefore, the bacterial isolates which produce capsules cannot be considered probiotics. The capsules can be detected by various microscopy methods. Moreover, polymerase chain reaction (PCR) based method can detect specific capsule gene harbored by the bacterial strain.
Access provided by Autonomous University of Puebla. Download chapter PDF
Similar content being viewed by others
Key words
1 Introduction
Bacterial capsular polysaccharides are a different group of high molecular weight polysaccharides that contribute to the virulence of many human pathogens in the gut, urinary tract, respiratory tract, and other host tissues by masking cell-surface components that would otherwise elicit an immune response from the host [1]. Pathogenic bacterial polysaccharide capsules have been shown to sterically shelter antigens from opsonizing antibodies, protect against antimicrobial peptides, and alter immunological responses. Capsules polysaccharide from probiotic bacterial species might have similar effects. The previous study indicated that biosynthesis of high-molecular-weight, galactose-rich surface polysaccharide molecules negatively impacts Lactobacillus rhamnosus GG ability to bind to intestinal epithelial cells, which may be due to adhesins on the bacterial cell surface is shielded [2]. Similarly, the purified capsular polysaccharide of Lactobacillus casei shirota, which was demonstrated to mediate the repression of pro-inflammatory responses in macrophages, has been postulated to play a direct role in host signaling [3]. Bacterial capsules have been detected by serological reactions, molecular genetic approaches, and electron microscopy but light microscopy is one of the most readily available and inexpensive techniques. After staining bacterial cells with nonspecific dyes, capsular polysaccharides can be seen using a light microscope. Standard dyes do not stain the capsules well, basic dyes are frequently combined with acidic dyes to stain the cells and background, respectively, while leaving the capsules transparent.
2 Materials
2.1 Microscopy Methods
-
1.
Anthony’s Method: 1% crystal violet and 20% copper sulfate.
-
2.
Hiss’s Method: 0.1% crystal violet and 20% copper sulfate.
-
3.
Maneval’s Method: Congo red and Maneval’s stain {(distilled water (89.26 mL): CH3COOH (4.68 mL): C6H6O (3.21 g): FeCl3∙6H2O (2.80 g): acid fuchsin (0.05 g)}.
-
4.
M’Fadyean’s Method: 96% ethanol/methanol and 1% methylene blue or Loeffler’s blue: {methylene blue (0.30 g): 95% ethanol (30 mL): 0.01% KOH (100 mL): K2CO3 (0.65 g)}.
-
5.
Duguid’s Method: India ink.
-
6.
India ink staining Method: India ink and Crystal violet.
-
7.
Negative–positive staining Method: Negative stains (India ink/Congo red/nigrosin) and Positive stains: (1% crystal violet/1% methylene blue).
-
8.
Bacterial culture.
-
9.
Microscope slide.
-
10.
Covers slips.
-
11.
Filter paper.
-
12.
Nichrome wire loop (for inoculation).
-
13.
Compound light microscope.
2.2 PCR Based Method
-
1.
Ethidium bromide.
-
2.
Agarose gel.
-
3.
Tris–HCl.
-
4.
KCl.
-
5.
MgCl2.
-
6.
dNTPs.
-
7.
Taq DNA polymerase.
-
8.
Primers for Capsule genes.
-
9.
Extracted bacterial DNA.
-
10.
Molecular size marker (50 or 100 bp DNA ladder).
-
11.
PCR tubes.
-
12.
Thermal cycler.
-
13.
Microtips and micropipettes.
-
14.
UV transilluminator.
3 Methods
3.1 Microscopy Methods
3.1.1 Anthony’s Method [4]
-
1.
Place three loopful cultures of test bacterium (probiotic) from broth culture on a glass slide.
-
2.
Obtain another clean microscope slide and place it at an angle to the end of the slide containing organisms. Spread out the drop into a thin film along with the first slide.
-
3.
Allow the smear to air dry without heat fixation.
-
4.
Apply 1% crystal violet to smear and allow it to react for 2 min.
-
5.
Wash the slide with 20% copper sulfate solution.
-
6.
Let the slide air dry and then examine under oil immersion.
-
7.
Bacterial cells will appear purple while the capsules will appear light blue.
3.1.2 Hiss’s Method [5]
-
1.
Place a drop of 1% crystal violet stain on a clean microscope slide.
-
2.
Using a sterile loop add a sufficient amount of bacteria from the culture tube or plate and mix it into the drop of crystal violet.
-
3.
Prepare a very thin smear over the entire surface of the slide with the help of another slide.
-
4.
Wait for 5–7 min to dry the slide, do not heat fix the slide.
-
5.
Wash the slide thoroughly with 20% CuSO4 solution.
-
6.
Allow it to air dry and examine under oil immersion.
-
7.
Bacterial cells will appear purple while the capsules will appear light blue.
3.1.3 Maneval’s Method [6]
-
1.
Place a drop of 1% congo red at one end of the microscope slide.
-
2.
Using a sterile loop, collect bacteria from the culture tube or plate and mix it into the drop of congo red.
-
3.
Then spread it gently on the slide to form a smear and allow it to air dry.
-
4.
Flood the Maneval’s solution to the smear and allow it to react for 5 min.
-
5.
Afterward, gently wash the slide with distilled water, allow it to air dry and examine under oil immersion.
-
6.
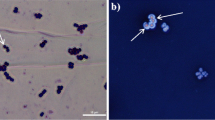
In the microscopic field, bacterial cells will appear red and capsules will be colorless against the red background (as shown in Fig. 1).
3.1.4 M’Fadyean’s Method [7]
-
1.
Make a thin smear by spreading a small drop of culture on a microscope slide using another slide.
-
2.
Allow it to air dry and then add 96% ethanol or methanol for 30–60 s.
-
3.
Allow the remaining solution to evaporate.
-
4.
Add methylene blue or Loeffler’s blue to smear for 30–60 s.
-
5.
Wash the slide with distilled water and leave it to air dry.
-
6.
Examine the slide under oil immersion.
-
7.
The pink capsules will appear surrounding the bluish bacterial cells.
3.1.5 Duguid’s Method [8]
-
1.
Place a drop of India ink on a microscope slide.
-
2.
Using a sterile loop, mix bacterial culture with an India ink and cover with a cover slip.
-
3.
Blot off the excess sample and then examine under 400× magnifications.
-
4.
The capsules will appear as brighter halos against dark bacterial cells and dark background of the slide.
3.1.6 India Ink Staining Method [9]
-
1.
Place a drop of India ink on a microscope slide.
-
2.
Using a sterile loop, mix bacterial culture with an India ink and spread as thin film with another slide.
-
3.
Allow it to air dry, and then stain with crystal violet for 1 min.
-
4.
Wash off the excess dye with distilled water.
-
5.
Allow it to air dry and examine under oil immersion.
-
6.
The bacterial cells will appear purple and the background black. The capsules will appear as clear zones around the cells.
3.1.7 The Negative–Positive Method [10]
-
1.
Place a drop of a negative stain such as nigrosin, India ink, congo red, or eosin on the microscope slide.
-
2.
Using a sterile loop, add a loopful of bacterial culture to slide and mix with stain.
-
3.
Use another slide to spread the mixture into a thin film along with the first slide.
-
4.
Allow it to air dry for 5–7 min. and do not heat fix it.
-
5.
Then, apply the positive dye such as crystal violet for 1 min.
-
6.
Remove the crystal violet by a positioned slide at 45° and then wash off with distilled water.
-
7.
Allow it to air dry and examine under oil immersion.
-
8.
If we use a combination of India ink as a negative dye and crystal violet or phenol fuchsin as a positive one then the cell will appear purple or pink against a blue background and capsules are revealed as a bright halo around bacterial cells.
3.2 PCR Based Method
The amplification of capsule gene can be carried out by using polymerase chain reaction. The steps for PCR based detection of capsule gene are given below.
-
1.
Extract the DNA from the bacterial strain by following any standard protocol.
-
2.
Analyze the extracted DNA on 1% agarose gel stained with ethidium bromide.
-
3.
Use genomic DNA isolated from bacterial strain as DNA template and select pairs of primers for PCR amplification to screen the genes involved in capsular polysaccharide formation (e.g. cps2A-J and cps4A-J in Lactobacillus plantarum WCFS1 [11]). Refer Table 1 for the list of capsule biosynthesis related genes primers.
-
4.
Make the final volume of 25 μL PCR mixture which include {10× (100 mM Tris–HCl pH 8.8 and 500 mM KCl), 1.7 mM MgCl2, 0.2 mM dNTPs, and 1.5 U of Taq DNA polymerase}, 10 pmol/μL of each primer, and 100 ng of target DNA.
-
5.
Setup the PCR conditions in a thermal cycler for the capsule gene amplification as follows: 5 min denaturation at 94 °C followed by 35 cycles of amplification with 1 min denaturation at 94 °C, 45 s of annealing at 45–55 °C (depending on capsule gene primers) and 2 min extension at 72 °C followed by final extension step 72 °C for 10 min.
-
6.
Analyze the PCR product using agarose gel (1%) electrophoresis and observe for specific bands by staining the gel with ethidium bromide and visualize under UV transilluminator.
-
7.
Compare the size of the amplified product with the molecular size marker (50 or 100 bp DNA ladder) for identification of the capsule gene.
References
Cress BF, Englaender JA, He W, Kasper D, Linhardt RJ, Koffas MA (2014) Masquerading microbial pathogens: capsular polysaccharides mimic host-tissue molecules. FEMS Microbiol Rev 38:660–697
Lebeer S, Verhoeven TL, Francius G, Schoofs G, Lambrichts I, Dufrêne Y, Vanderleyden J, De Keersmaecker SCJ (2009) Identification of a gene cluster for the biosynthesis of a long, galactose-rich exopolysaccharide in Lactobacillus rhamnosus GG and functional analysis of the priming glycosyltransferase. Appl Environ Microbiol 75:3554–3563
Yasuda E, Serata M, Sako T (2008) Suppressive effect on activation of macrophages by Lactobacillus casei strain Shirota genes determining the synthesis of cell wall-associated polysaccharides. Appl Environ Microbiol 74:4746–4755
Anthony E Jr (1931) A note on capsule staining. Science 73:319–320
Hiss PH Jr (1905) A contribution to the physiological differentiation of pneumococcus and streptococcus, and to methods of staining capsules. J Exp Med 6:317–345
Maneval W (1941) Staining bacteria and yeasts with acid dyes. Stain Technol 16:13–19
Owen MP, Kiernan JA (2004) The M’Fadyean reaction: a stain for anthrax bacilli. Biotech Histochem 79:107–108
Duguid J (1951) The demonstration of bacterial capsules and slime. J Pathol Bacteriol 63:673–685
Butt EM, Bonynge CW, Joyce RL (1936) The demonstration of capsules about hemolytic streptococci with India ink or azo blue. J Infect Dis 58:5–9
Oleksy M, Klewicka E (2017) Capsular polysaccharides of Lactobacillus spp.: theoretical and practical aspects of simple visualization methods. Probiotics Antimicrob Proteins 9:425–434
Remus DM, van Kranenburg R, van Swam II, Taverne N, Bongers RS, Wels M, Bron PA, Kleerebezem M (2012) Impact of 4 Lactobacillus plantarum capsular polysaccharide clusters on surface glycan composition and host cell signaling. Microb Cell Fact 11:1–10
Pan YJ, Fang HC, Yang HC, Lin TL, Hsieh PF, Tsai FC, Keynan Y, Wang JT (2008) Capsular polysaccharide synthesis regions in Klebsiella pneumoniae serotype K57 and a new capsular serotype. J Clin Microbiol 46:2231–2240
Chen Y, Dai J, Morris JG, Johnson JA (2010) Genetic analysis of the capsule polysaccharide (K antigen) and exopolysaccharide genes in pandemic Vibrio parahaemolyticus O3: K6. BMC Microbiol 10:1–10
Waite RD, Penfold DW, Struthers JK, Dowson CG (2003) Spontaneous sequence duplications within capsule genes cap8E and tts control phase variation in Streptococcus pneumoniae serotypes 8 and 37. Microbiology 149:497–504
Townsend KM, Boyce JD, Chung JY, Frost AJ, Adler B (2001) Genetic organization of Pasteurella multocida cap loci and development of a multiplex capsular PCR typing system. J Clin Microbiol 39:924–929
Author information
Authors and Affiliations
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2022 The Author(s), under exclusive license to Springer Science+Business Media, LLC, part of Springer Nature
About this chapter
Cite this chapter
Shah, R., Amaresan, N., Dwivedi, M.K. (2022). Assessment of Capsule Formation. In: Dwivedi, M.K., Amaresan, N., Sankaranarayanan, A., Begum, R. (eds) Biosafety Assessment of Probiotic Potential. Methods and Protocols in Food Science . Humana, New York, NY. https://doi.org/10.1007/978-1-0716-2509-5_17
Download citation
DOI: https://doi.org/10.1007/978-1-0716-2509-5_17
Published:
Publisher Name: Humana, New York, NY
Print ISBN: 978-1-0716-2508-8
Online ISBN: 978-1-0716-2509-5
eBook Packages: Springer Protocols