Abstract
The bacterial flagellum is a supramolecular motility machine consisting of the basal body, the hook, and the filament. For construction of the flagellum beyond the cellular membranes, a type III protein export apparatus uses ATP and proton-motive force (PMF) across the cytoplasmic membrane as the energy sources to transport flagellar component proteins from the cytoplasm to the distal end of the growing flagellar structure. The protein export apparatus consists of a PMF-driven transmembrane export gate complex and a cytoplasmic ATPase complex. In addition, the basal body C ring acts as a sorting platform for the cytoplasmic ATPase complex that efficiently brings export substrates and type III export chaperone–substrate complexes from the cytoplasm to the export gate complex. In this book chapter, we will summarize our current understanding of molecular organization and assembly of the flagellar type III protein export apparatus.
Access provided by Autonomous University of Puebla. Download chapter PDF
Similar content being viewed by others
Keywords
- Assembly
- ATPase
- Bacterial flagellum
- Chaperone
- Crystal structure
- Electron cryotomography
- Electron cryomicroscopy
- Injectisome
- Protein export apparatus
1 Introduction
Bacteria swim in liquid environments and move on solid surfaces by rotating a very long filamentous assembly called the bacterial flagellum. The flagellum consists of at least three parts: the basal body as a bidirectional rotary motor, the hook as a universal joint, and the filament as a helical propeller. Flagellar assembly begins with the basal body, followed by the hook and finally the filament. Fourteen flagellar proteins are transported via a type III protein export apparatus into the central channel inside the growing structure and assemble at the distal end (Macnab 2004; Minamino et al. 2008; Minamino 2014).
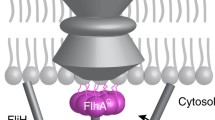
The flagellar type III protein export apparatus is composed of a transmembrane export gate complex made of FlhA, FlhB, FliP, FliQ, and FliR and a cytoplasmic ATPase complex consisting of FliH, FliI, and FliJ. These proteins are evolutionarily related to components of the virulence-associated type III secretion system (T3SS) of pathogenic bacteria, also known as the injectisome (Fig. 1) (Galán et al. 2014; Wagner et al. 2018).
Structural comparison between the flagellum and the injectisome. In situ structures of the basal bodies of the Salmonella flagellum (a) and injectisome (b) are visualized by electron cryotomography and subtomogram averaging. The central section maps of the flagellum (EMDB-2521) (a) and the injectisome (EMDB-8544) (b) after subtomogram averaging are shown. Upper panels, side view, lower panels, bottom view corresponding a cross section at height indicated by the dashed yellow line. c Schematic diagrams of cytoplasmic portions of the Salmonella flagellum (left panel) and injectisome (right panel). Name of each part of the basal body and component protein(s) are shown
FliG, FliM, and FliN form the C ring on the cytoplasmic face of the basal body MS ring made of a transmembrane protein, FliF. The C ring acts not only as the rotor of the flagellar motor but also as the switch for bidirectional motor rotation, allowing the flagellar motor to rotate both in the counterclockwise and in the clockwise directions (Berg 2003; Morimoto and Minamino 2014). FliM and FliN, which are well conserved in virulence-associated T3SS families, provide the binding sites for the cytoplasmic ATPase complex in complex with export substrates and export chaperone–substrate complexes (González-Pedrajo et al. 2006; Minamino et al. 2009; Lara-Tejero et al. 2011). In this book chapter, we will describe the structure and assembly of the flagellar type III protein export apparatus in Salmonella enterica.
2 Structure of the Transmembrane Export Gate Complex
FlhA, FlhB, FliP, FliQ, and FliR form the transmembrane export gate complex inside the MS ring (Minamino and Macnab 1999; Fukumura et al. 2017). A transmembrane protein, FliO, which is not conserved in virulence-associated T3SS families of pathogenic bacteria, is required for efficient assembly of the export gate complex inside the MS ring although it is not essential for flagellar protein export (Barker et al. 2010; Morimoto et al. 2014; Fukumura et al. 2017; Fabiani et al. 2017). The transmembrane export gate complex is powered by the proton-motive force (PMF) across the cytoplasmic membrane and facilitates unfolding and protein translocation across the cytoplasmic membrane (Minamino and Namba 2008; Paul et al. 2008; Minamino et al. 2011; Lee et al. 2014; Terashima et al. 2018).
2.1 FlhA Ring Structure
FlhA forms an ion channel to conduct protons and sodium ions and plays an important role in the energy coupling mechanism along with the cytoplasmic ATPase complex (Minamino et al. 2011, 2016; Morimoto et al. 2016; Erhardt et al. 2017). FlhA consists of a hydrophobic N-terminal transmembrane domain (FlhATM) and a large C-terminal cytoplasmic domain (FlhAC) (Minamino et al. 1994). Genetic and biochemical analyses have shown that FlhATM interacts with FlhB, FliF, and FliR (Kihara et al. 2001; Barker and Samatey 2012; Hara et al. 2011; Fukumura et al. 2017). FlhAC has been visualized by electron cryotomography (ECT) and subtomogram averaging to form a ring-shaped projection in the cavity within the C ring (Fig. 1a, c) (Abrusci et al. 2013; Kawamoto et al. 2013). Similar ring-like structures formed by the C-terminal cytoplasmic domain of a FlhA homologue of the injectisome, SctV, have been identified by ECT (Fig. 1b, c) (Kawamoto et al. 2013; Hu et al. 2015, 2017; Makino et al. 2016). FlhAC and SctVC form a homo-nonamer as part of the export gate complex (Fig. 2a) (Abrusci et al. 2013; Kawamoto et al. 2013; Morimoto et al. 2014). FlhAC interacts with FliH, FliI, FliJ, flagellar type III export chaperones, and export substrates and coordinates flagellar protein export with assembly (Minamino and Macnab 2000c; Bange et al. 2010; Minamino et al. 2012a; Kinoshita et al. 2013; Furukawa et al. 2016; Kinoshita et al. 2016; Inoue et al. 2018). This suggests that the FlhAC ring structure acts as a docking platform for these proteins and plays an important role in hierarchical protein targeting and export.
Atomic models of the docking platform made of FlhAC and the cytoplasmic ATPase ring complex consisting of FliH, FliI and FliJ. a The FlhAC ring, which is involved in the interactions with FliH, FliI, FliJ, flagellar type III export chaperones and export substrates and b the FliH12FliI6FliJ ring, which plays an important role in energy transduction. Cα ribbon representation of FlhAC (PDB ID: 3A5I), the FliH2FliI complex (PDB ID: 5B0O), and FliJ (PDB ID: 3AJW) is shown. Highly conserved Gln38, Leu42, Tyr45, Tyr49, Phe72, Leu76, Ala79, and His83 residues of FliJ (indicated as Q38, L42, Y45, Y49, F72, L76, A79, and H83, respectively) are responsible for the interaction with FlhAC and the FliJ–FlhAC interaction facilitates PMF-driven protein export by the transmembrane export gate complex c crystal structure of FlhAC in complex with a FliDC–FliT fusion protein (PDB ID: 6CH2). FlhAC consists of four domains: D1, D2, D3, and D4. A highly conserved Tyr106 residue of FliT (magenta) binds to a conserved hydrophobic dimple at an interface between domains D1 and D2 of FlhAC. Well conserved Asp456, Phe459 and Thr490 residues of FlhAC (indicated as D456, F459, and T490, respectively) are responsible for the interaction with Tyr108 of FliT
Crystal structures of FlhAC and SctVC have been solved by X-ray crystallography (Bange et al. 2010; Moore and Jia 2010; Saijo-Hamano et al. 2010; Worrall et al. 2010; Abrusci et al. 2013). FlhAC consists of four domains, D1, D2, D3, and D4, and a flexible linker (FlhAL) connecting with FlhATM (Fig. 2c). The crystal structure of SctVC derived from the Shigella injectisome forms a nonameric ring structure through D1–D3 and D3–D3 interactions (Abrusci et al. 2013). Because similar subunit interactions are observed in the crystal packing of the Salmonella FlhAC structure (Saijo-Hamano et al. 2010), D1–D3 and D3–D3 interactions are likely to be responsible for the FlhAC ring formation (Kawamoto et al. 2013). In addition to these interactions, interactions of FlhAL with domains D1 and D3 of its neighboring FlhAC subunit are involved in highly cooperative FlhAC ring formation in solution (Terahara et al. 2018). FlhAC adopts two distinct, open and closed conformations (Moore and Jia 2010; Saijo-Hamano et al. 2010; Worrall et al. 2010; Abrusci et al. 2013). A large open cleft between domains D2 and D4 is observed in the open form, but the cleft is closed in the closed form. A conserved hydrophobic dimple containing Asp456, Phe459, and Thr490 resides is located at the interface between domains D1 and D2 and is directly involved in the interactions with the FlgN, FliS, and FliT chaperones in complex with their cognate substrates (Minamino et al. 2012a; Kinoshita et al. 2013). Crystal structures of FlhAC in complex with the chaperone–substrate complexes have shown that the chaperones bind to the hydrophobic dimple of the open form of FlhAC (Fig. 2c) (Xing et al. 2018). Mutations at residues involved in the interactions of FlhAL with its neighboring FlhAC subunits in the FlhAC ring structure significantly weaken the interaction of FlhAC with the chaperone–substrate complexes, thereby reducing the probability of filament assembly at the hook tip (Terahara et al. 2018). This leads to a plausible hypothesis that interactions of FlhAL with the D1 and D3 domains of its neighboring FlhAC subunits may convert the FlhAC ring structure from the closed conformation to the open one, allowing the chaperone–substrate complexes to bind to the FlhAC ring.
2.2 FlhB
The type III protein export apparatus undergoes substrate specificity switching upon completion of hook assembly, terminating hook assembly, and initiating filament assembly. FlhB is involved in the substrate specificity switching along with a secreted molecular ruler protein, FliK, which is also secreted via the type III protein export apparatus during hook assembly (Minamino 2018). FlhB consists of a hydrophobic N-terminal domain and a relatively large C-terminal cytoplasmic domain (FlhBC) (Minamino et al. 1994). Crystal structures of FlhBC and its homologue of the injectisome, SctUC, have been solved by X-ray crystallography. FlhBC and SctUC contain two distinct: CN and CC polypeptides (Zarivach et al. 2008; Meshcheryakov et al. 2013). The FlhBCN polypeptide consists of a long α helix (α1) and a β strand (β1). The FlhBCC polypeptide is composed of three α helices (α2, α3, α4), three β strands (β2, β3, β4), and a highly flexible C-terminal tail (residues 354–383, FlhBCCT). Four α helices surround a four-stranded β sheet, forming a globular domain (Meshcheryakov et al. 2013). FlhBCCT is dispensable for FlhB function, but its truncation results in autonomous substrate specificity switching of the type III protein export apparatus in the absence of FliK (Kutsukake et al. 1994). This suggests that FlhBCCT may contribute to the well-regulated substrate specificity switching. A highly conserved NPTH sequence lies on a flexible loop located between FlhBCN and FlhBCC (Minamino and Macnab 2000a; Fraser et al. 2003). FlhBC undergoes autocatalytic cleavage between Asn269 and Pro270 within the NPTH sequence to be split into FlhBCN and FlhBCC by a mechanism involving cyclization of Asn269 (Ferris et al. 2005; Zarivach et al. 2008). A conserved hydrophobic patch formed by Ala286, Pro287, Ala341, and Leu344 in FlhBCC is directly involved in interactions with the N-terminal region of the hook protein, which contains an export signal recognized by the type III protein export apparatus (Evans et al. 2013). Photo-cross-linking experiments combined with mutational analyses have shown that the C-terminal domain of FliK binds to FlhBC, thereby terminating the export of the rod and hook-type proteins and initiating the export of the filament-type proteins (Kinoshita et al. 2017).
2.3 Core Structure of the Export Gate Complex
FliP, FliQ, and FliR form a core structure of the transmembrane export gate complex (Fukumura et al. 2017). The structure of purified FliP–FliQ–FliR core complex has been determined at 4.2 Å resolution by electron cryomicroscopy (cryoEM) and single-particle image analysis. The cryoEM structure of the core complex adopts a right-handed helical assembly composed of five copies of FliP, four copies of FliQ, and one copy of FliR (Kuhlen et al. 2018) (Fig. 3). Although FliP, FliQ, and FliR are predicted to have four, two, and six transmembrane helices, respectively (Ohnishi et al. 1997), they do not adopt canonical integral membrane topologies in the core complex (Kuhlen et al. 2018). FliP and FliR form a FliP5FliR1 complex, and four FliQ subunits are associated with the FliP5FliR1 complex on its outside. FliR is a structural fusion of FliP and FliQ and so compensates for a helical rise between the first and the fifth FliP subunits to stabilize the helical structure. The assembled FliP5FliQ4FliR1 core complex has a central pore with a diameter of 1.5 nm, which seems to be the protein translocation channel (Kuhlen et al. 2018). The most distal part of the core complex is likely to interact with the most proximal end of the rod inside the basal body MS ring (Dietsche et al. 2016; Kuhlen et al. 2018). Biochemical analyses have shown that SctR, SctS, and SctT, which are FliP, FliQ, and FliR homologues of the injectisome, respectively, form a core structure in the injectisome in a way similar to the assembly of the FliP5FliQ4FliR1 complex (Wagner et al. 2010; Zilkenat et al. 2016; Dietsche et al. 2016).
FliO consists of an N-terminal periplasmic tail, a single transmembrane helix and a C-terminal cytoplasmic domain (Barker et al. 2010). FliO forms a 5 nm ring structure with three flexible clamp-like structures that bind to FliP to facilitate FliP oligomerization (Fukumura et al. 2017). Highly conserved Phe137, Phe150, and Glu178 residues of FliP, which are functionally important, contribute not only to FliO–FliP interaction but also for FliP–FliP interaction (Fukumura et al. 2017). The FliO ring complex protects FliP from proteolytic degradation and promotes stable FliP5FliR1 complex formation (Fabiani et al. 2017). Overexpression of FliP restores motility of a Salmonella fliO null mutant to the wild-type level, suggesting that the FliO ring complex acts as a structural scaffold to facilitate the helical assembly of the FliP5FliR1 complex (Barker et al. 2010; Fukumura et al. 2017; Fabiani et al. 2017).
3 Cytoplasmic ATPase Ring Complex
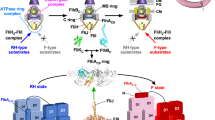
The cytoplasmic ATPase ring complex is composed of twelve copies of FliH, six copies of FliI and one copy of FliJ (Fig. 2b) (Imada et al. 2016). The ATPase ring structure has been visualized at the flagellar base by ECT (Fig. 1a, c) (Chen et al. 2011). ATP hydrolysis by the ATPase ring complex activates the transmembrane export gate complex through an interaction between FliJ and FlhA, allowing the export gate complex to utilize PMF across the cytoplasmic membrane to transport flagellar proteins (Minamino et al. 2014). The FliH12FliI6FliJ complex is structurally similar to F- and V-type rotary ATPases, suggesting that the flagellar ATPase ring complex is evolutionally related to these rotary ATPases (Imada et al. 2007; Ibuki et al. 2011; Imada et al. 2016).
FliI is a Walker-type ATPase and forms a homo-hexamer to fully exert its ATPase activity (Fig. 2b) (Fan et al. 1996; Claret et al. 2003). ATP binds to an ATP binding site located an interface between FliI subunits in a way similar to that found in F- and V-type rotary ATPases, and so FliI ring formation is required for ATP hydrolysis by the FliI ATPase (Kazetani et al. 2009). FliJ facilitates the assembly of FliI into the hexameric ring structure by binding to the center of the ring (Ibuki et al. 2011). Highly conserved, surface-exposed residues of FliJ, namely Gln38, Leu42, Tyr45, Tyr49, Phe72, Leu76, Ala79, and His83, are involved in the interaction with FlhAL (Fig. 2b) (Ibuki et al. 2013), and the interaction between FliJ and FlhAL coordinates ATP hydrolysis by the FliI6 ring with proton-coupled flagellar protein export (Minamino et al. 2014; Morimoto et al. 2016). FliH forms a homo-dimer (Minamino and Macnab 2000b) although the conformation of the two FliH monomers is different from each other (Minamino et al. 2002; Imada et al. 2016). The C-terminal domain of FliH binds to the extreme N-terminal region of FliI (Fig. 2b) (Minamino and Macnab 2000b; González-Pedrajo et al. 2002; Okabe et al. 2009; Imada et al. 2016). The FliH dimer binds to a C ring protein, FliN, and FlhA to allow the ATPase ring complex to efficiently localize to the flagellar base (González-Pedrajo et al. 2006; Minamino et al. 2009; Bai et al. 2014). Two conserved Trp7 and Trp10 residues of FliHN are directly involved in the interactions of FliH with FliN and FlhA (Minamino et al. 2009; Hara et al. 2012; Notti et al. 2015).
FliH and FliI also exist as a FliH2FliI1 complex in the cytoplasm (Minamino and Macnab 2000b; Minamino et al. 2001). Flagellar export chaperone–substrate complexes bind to the FliH2FliI1 complex through an interaction between the FliI ATPase and the chaperone (Thomas et al. 2004; Imada et al. 2010; Minamino et al. 2012b). More than six copies of FliI labeled with yellow fluorescent protein (FliI-YFP) are estimated to be associated with the basal body through the interactions of FliH with FliN and FlhA. Since FliI-YFP shows rapid exchanges between the flagellar basal body and the cytoplasmic pool, the FliH2FliI1 complex is thought to act as a dynamic carrier to bring export substrates and chaperone–substrate complexes from the cytoplasm to the FlhAC–FlhBC docking platform of the transmembrane export gate complex (Bai et al. 2014; Terashima et al. 2018).
4 Sorting Platform
The C ring is composed of FliG, FliM, and FliN. The C ring acts not only as a rotor of the flagellar motor but also as a structural switch to change the direction of flagellar motor rotation (Fig. 1c) (Berg 2003; Morimoto and Minamino 2014). The C ring has 34-fold rotational symmetry (Thomas et al. 1999). FliG consists of three domains: N-terminal (FliGN), middle (FliGM), and C-terminal (FliGC) domains (Lee et al. 2010). FliGN directly binds to the MS ring protein, FliF (Kihara et al. 2000). Crystal structures of FliGN in complex with the C-terminal portion of FliF (FliFC) have been solved by X-ray crystallography (Lynch et al. 2017; Xue et al. 2018). Two α helices of FliFC are deeply inserted into a hydrophobic groove of FliGN formed by four α helices. Helix α4 of FliGN adopts two distinct conformations. One adopts an extended conformation whereas the other is divided into helices α4a and α4b. When the FliFC–FliGN complex exists in a solution, the α4 helix adopts an extended conformation (Lynch et al. 2017). When FliG assembles into the FliG ring structure on the cytoplasmic face of the MS ring, the α4 helix induces a conformational change and so helix α4b interacts with helix α4a of its neighboring FliG subunit (Lynch et al. 2017; Xue et al. 2018). The FliGMC unit, which consists of FliGM, FliGC and a helix linker connecting these two domains, adopts a compact conformation through an intramolecular interaction between FliGM and FliGC, allowing FliG to exit as a monomer in solution (Baker et al. 2016; Kinoshita et al. 2018a). In contrast, when the FliGMC unit adopts an extended conformation, intermolecular interactions between FliGM and FliGC promote the self-assembly of FliG into the ring structure (Baker et al. 2016; Kinoshita et al. 2018b). FliM and FliN form a stable FliM1FliN3 complex through an interaction between the C-terminal domain of FliM (FliMC) and FliN (Notti et al. 2015; McDowell et al. 2016) and bind to the FliG ring through an interaction between the middle domain of FliM (FliMM) and FliGM to form the C ring wall (Paul et al. 2011; Vartanian et al. 2012). Intermolecular interactions between FliMM domains are required for the formation of the continuous wall of the C ring (Park et al. 2006). FliMC and FliN together form a spiral structure at the bottom of the C ring (McDowell et al. 2016). It has been shown that surface-exposed hydrophobic residues of FliN, Val111, Val112 and Val113, are involved in the interaction with FliH (McMurry et al. 2006; Paul et al. 2006). Overexpression of FliI partially rescues the reduced ability of flagellar protein export by the fliN null mutant (McMurry et al. 2006), suggesting that the C ring is required for efficient recruitment of the FliH2FliI1 complex to the type III protein export apparatus for efficient flagellar protein export.
It has also been shown that the sorting platform of the injectisome contributes to a strict order of protein secretion by the type III protein export apparatus (Lara-Tejero et al. 2011). However, the structure and stoichiometry of the sorting platform of the injectisome are distinct from those of the flagellar C ring structure (Fig. 1) (Kawamoto et al. 2013; Hu et al. 2015; Makino et al. 2016; Hu et al. 2017). A FliM/FliN homologue of the injectisome, SctQ, forms six pod-like structures on the cytoplasmic face of the cytoplasmic membrane ring (Fig. 1b) (Hu et al. 2015; Makino et al. 2016; Hu et al. 2017). SctK, which is not conserved in the flagellar type III protein export system, associates with the pod-like structure (Hu et al. 2017). SctL, which is a FliH homologue of the injectisome, forms a linker connecting the pod and the ATPase ring complex made of a FliI homologue, SctN (Notti et al. 2015; Hu et al. 2017). It has been shown the sorting platform is highly dynamic structure during protein secretion (Diepold et al. 2015, 2017).
5 Assembly of the Type III Protein Export Apparatus
FliF assembles into the MS ring within the cytoplasmic membrane (Kubori et al. 1992; Ueno et al. 1992). Recently, it has been reported that FliG is required for efficient MS ring formation (Li and Sourjik 2011; Morimoto et al. 2014). This suggests that FliF and FliG together form the MS–FliG ring complex. FliP and FliR form a FliP5FliR1 complex in a FliO-dependent manner (Dietsche et al. 2016; Fukumura et al. 2017; Fabiani et al. 2017). FliQ is peripherally associated around the outside of the FliP5FliR1 complex and forms a helical FliP5FliQ4FliR1 structure inside the MS ring (Kuhlen et al. 2018). FlhA and FlhB are associated with the FliP5FliQ4FliR1 core complex (Fukumura et al. 2017). FlhA also binds to the MS ring directly (Fukumura et al. 2017). Since FlhA requires FliF, FliG, FliO, FliP, FliQ, and FliR for its assembly to the flagellar basal body but not FlhB (Morimoto et al. 2014), it has been proposed that the assembly of the flagellar type III export gate complex begins with the formation of the FliP5FliR1 complex with the help of the FliO ring complex, followed by the assembly of FliQ and finally of FlhA and FlhB during MS ring formation (Fig. 4) (Wagner et al. 2010; Diepold et al. 2011; Dietsche et al. 2016; Fukumura et al. 2017).
Assembly mechanism of the type III protein export apparatus. FliP, FliQ, and FliR form a FliP5FliQ4FliR1 complex with the help of the FliO complex, followed by the assembly of FlhB and finally of FlhA during MS ring formation in the cytoplasmic membrane. Then, the FliM1FliN3 complex binds to FliG to form the C ring on the cytoplasmic face of the MS ring. Finally, the cytoplasmic ATPase ring complex made of FliH, FliI, and FliJ is formed and is associated with the C ring through interactions between FliH and FliN
The FliM1FliN3 complex forms the continuous wall of the C ring on the cytoplasmic face of the MS ring through the interactions between FliGM and FliMM (Paul et al. 2011; Vartanian et al. 2012). Finally, the cytoplasmic FliH12FliI6FliJ1 ring complex is formed at the flagellar base through the interactions of FliH with FlhA and FliN (Fig. 4). Upon completion of the type III protein export apparatus, export substrates and flagellar export chaperone–substrate complexes are efficiently recruited via the cytoplasmic FliH2FliI1 complex to the type III protein export apparatus to be transported into the central channel of the growing flagellar structure.
6 Conclusion
The type III protein export apparatus consists of a PMF-driven export gate complex made of FlhA, FlhB, FliP, FliQ, and FliR and a cytoplasmic ATPase complex consisting of FliH, FliI, and FliJ. The export apparatus utilizes ATP and PMF to efficiently couple the proton influx through the export gate complex with protein translocation into the central channel of the growing structure. Atomic structures of the C-terminal cytoplasmic domains of FlhA and FlhB, FliH, FliI, FliJ, and the FliP5FliQ4FliR1 helical assembly have been solved. However, it still remains unknown how flagellar proteins are unfolded and transported by the PMF-driven export gate complex. High-resolution structural analysis of the entire protein export apparatus by cryoEM image analysis would be essential to advance our mechanistic understanding of the type III protein export process.
References
Abrusci P, Vergara-Irigaray M, Johnson S, Beeby MD, Hendrixson DR, Roversi P, Friede ME, Deane JE, Jensen GJ, Tang CM, Lea SM (2013) Architecture of the major component of the type III secretion system export apparatus. Nat Struct Mol Biol 20:99–104
Bai F, Morimoto YV, Yoshimura SDJ, Hara N, Kami-ike N, Namba K, Minamino T (2014) Assembly dynamics and the roles of FliI ATPase of the bacterial flagellar export apparatus. Sci Rep 4:6528
Baker MA, Hynson RM, Ganuelas LA, Mohammadi NS, Liew CW, Rey AA, Duff AP, Whitten AE, Jeffries CM, Delalez NJ, Morimoto YV, Stock D, Armitage JP, Turberfield AJ, Namba K, Berry RM, Lee LK (2016) Domain-swap polymerization drives the self-assembly of the bacterial flagellar motor. Nat Struct Mol Biol 23:197–203
Bange G, Kümmerer N, Engel C, Bozkurt G, Wild K, Sinning I (2010) FlhA provides the adaptor for coordinated delivery of late flagella building blocks to the type III secretion system. Proc Natl Acad Sci USA 107:11295–11300
Barker CS, Samatey FA (2012) Cross-complementation study of the flagellar type III export apparatus membrane protein FlhB. PLoS ONE 7:e44030
Barker CS, Meshcheryakova IV, Kostyukova AS, Samatey FA (2010) FliO regulation of FliP in the formation of the Salmonella enterica flagellum. PLoS Genet 6:e1001143
Berg HC (2003) The rotary motor of bacterial flagella. Ann Rev Biochem 72:19–54
Chen S, Beeby M, Murphy GE, Leadbetter JR, Hendrixson DR, Briegel A, Li Z, Shi J, Tocheva EI, Müller A, Dobro MJ, Jensen GJ (2011) Structural diversity of bacterial flagellar motors. EMBO J 30:2972–2981
Claret L, Susannah CR, Higgins M, Hughes C (2003) Oligomerization and activation of the FliI ATPase central to bacterial flagellum assembly. Mol Microbiol 48:1349–1355
Diepold A, Wiesand U, Cornelis GR (2011) The assembly of the export apparatus (YscR, S, T, U, V) of the Yersinia type III secretion apparatus occurs independently of other structural components and involves the formation of an YscV oligomer. Mol Microbiol 82:502–514
Diepold A, Kudryashev M, Delalez NJ, Berry RM, Armitage JP (2015) Composition, formation, and regulation of the cytosolic C-ring, a dynamic component of the type III secretion injectisome. PLoS Biol 13(1):e1002039
Diepold A, Sezgin E, Huseyin M, Mortimer T, Eggeling C, Armitage JP (2017) A dynamic and adaptive network of cytosolic interactions governs protein export by the T3SS injectisome. Nat Commun 8:15940
Dietsche T, Tesfazgi Mebrhatu M, Brunner MJ, Abrusci P, Yan J, Franz-Wachtel M, Schärfe C, Zilkenat S, Grin I, Galán JE, Kohlbacher O, Lea S, Macek B, Marlovits TC, Robinson CV, Wagner S (2016) Structural and functional characterization of the bacterial type III secretion export apparatus. PLoS Pathog 12:e1006071
Erhardt M, Wheatley P, Kim EA, Hirano T, Zhang Y, Sarkar MK, Hughes KT, Blair DF (2017) Mechanism of type-III protein secretion: regulation of FlhA conformation by a functionally critical charged-residue cluster. Mol Microbiol 104:234–249
Evans LD, Poulter S, Terentjev EM, Hughes C, Fraser GM (2013) A chain mechanism for flagellum growth. Nature 504:287–290
Fabiani FD, Renault TT, Peters B, Dietsche T, Gálvez EJC, Guse A, Freier K, Charpentier E, Strowig T, Franz-Wachtel M, Macek B, Wagner S, Hensel M, Erhardt M (2017) A flagellum-specific chaperone facilitates assembly of the core type III export apparatus of the bacterial flagellum. PLoS Biol 15:e2002267
Fan F, Macnab RM (1996) Enzymatic characterization of FliI: an ATPase involved in flagellar assembly in Salmonella typhimurium. J Biol Chem 271:31981–31988
Ferris HU, Furukawa Y, Minamino T, Kroetz MB, Kihara M, Namba K, Macnab RM (2005) FlhB regulates ordered export of flagellar components via autocleavage mechanism. J Biol Chem 280:41236–41242
Fraser GM, Hirano T, Ferris HU, Devgan LL, Kihara M, Macnab RM (2003) Substrate specificity of type III flagellar protein export in Salmonella is controlled by subdomain interactions in FlhB. Mol Microbiol 48:1043–1057
Fukumura T, Makino F, Dietsche T, Kinoshita M, Kato T, Wagner S, Namba K, Imada K, Minamino T (2017) Assembly and stoichiometry of the core structure of the bacterial flagellar type III export gate complex. PLoS Biol 15:e2002281
Furukawa Y, Inoue Y, Sakaguchi A, Mori Y, Fukumura T, Miyata T, Namba K, Minamino T (2016) Structural stability of flagellin subunits affects the rate of flagellin export in the absence of FliS chaperone. Mol Microbiol 102:405–416
Galán JE, Lara-Tejero M, Marlovits TC, Wagner S (2014) Bacterial type III secretion systems: specialized nanomachines for protein delivery into target cells. Annu Rev Microbiol 68:415–438
González-Pedrajo B, Fraser GM, Minamino T, Macnab RM (2002) Molecular dissection of Salmonella FliH, a regulator of the ATPase FliI and the type III flagellar protein export pathway. Mol Microbiol 45:967–982
González-Pedrajo B, Minamino T, Kihara M, Namba K (2006) Interactions between C ring proteins and export apparatus components: a possible mechanism for facilitating type III protein export. Mol Microbiol 60:984–998
Hara N, Namba K, Minamino T (2011) Genetic characterization of conserved charged residues in the bacterial flagellar type III export protein FlhA. PLoS ONE 6:e22417
Hara N, Morimoto YV, Kawamoto A, Namba K, Minamino T (2012) Interaction of the extreme N-terminal region of FliH with FlhA is required for efficient bacterial flagellar protein export. J Bacteriol 194:5353–5360
Hu B, Morado DR, Margolin W, Rohde JR, Arizmendi O, Picking WL, Picking WD, Liu J (2015) Visualization of the type III secretion sorting platform of Shigella flexneri. Proc Natl Acad Sci USA 112:1047–1052
Hu B, Lara-Tejero M, Kong Q, Galán JE, Liu J (2017) In situ molecular architecture of the Salmonella type III secretion machine. Cell 168:1065–1074
Ibuki T, Imada K, Minamino T, Kato T, Miyata T, Namba K (2011) Common architecture between the flagellar protein export apparatus and F- and V-ATPases. Nat Struct Mol Biol 18:277–282
Ibuki T, Uchida Y, Hironaka Y, Namba K, Imada K, Minamino T (2013) Interaction between FliJ and FlhA, components of the bacterial flagellar type III export apparatus. J Bacteriol 195:466–473
Imada K, Minamino T, Tahara A, Namba K (2007) Structural similarity between the flagellar type III ATPase FliI and F1-ATPase subunits. Proc Natl Acad Sci USA 104:485–490
Imada K, Minamino T, Kinoshita M, Furukawa Y, Namba K (2010) Structural insight into the regulatory mechanisms of interactions of the flagellar type III chaperone FliT with its binding partners. Proc Natl Acad Sci USA 107:8812–8817
Imada K, Minamino T, Uchida Y, Kinoshita M, Namba K (2016) Insight into the flagella type III export revealed by the complex structure of the type III ATPase and its regulator. Proc Natl Acad Sci USA 113:3633–3638
Inoue Y, Morimoto YV, Namba K, Minamino T (2018) Novel insights into the mechanism of well-ordered assembly of bacterial flagellar proteins in Salmonella. Sci Rep 8:1787
Kawamoto A, Morimoto YV, Miyata T, Minamino T, Hughes KT, Kato T, Namba K (2013) Common and distinct structural features of Salmonella injectisome and flagellar basal body. Sci Rep 3:3369
Kazetani K, Minamino T, Miyata T, Kato T, Namba K (2009) ATP-induced FliI hexamerization facilitates bacterial flagellar protein export. Biochem Biophys Res Commun 388:323–327
Kihara M, Miller GU, Macnab RM (2000) Deletion analysis of the flagellar switch protein FliG of Salmonella. J Bacteriol 182:3022–3028
Kihara M, Minamino T, Yamaguchi S, Macnab RM (2001) Intergenic suppression between the flagellar MS ring protein FliF of Salmonella and FlhA, a membrane component of its export apparatus. J Bacteriol 183:1655–1662
Kinoshita M, Hara N, Imada K, Namba K, Minamino T (2013) Interactions of bacterial flagellar chaperone-substrate complexes with FlhA contribute to co-ordinating assembly of the flagellar filament. Mol Microbiol 90:1249–1261
Kinoshita M, Nakanishi Y, Furukawa Y, Namba K, Imada K, Minamino T (2016) Rearrangements of α-helical structures of FlgN chaperone control the binding affinity for its cognate substrates during flagellar type III export. Mol Microbiol 101:656–670
Kinoshita M, Aizawa S, Inoue Y, Namba K, Minamino T (2017) The role of intrinsically disordered C-terminal region of FliK in substrate specificity switching of the bacterial flagellar type III export. Mol Microbiol 105:572–588
Kinoshita M, Furukawa Y, Uchiyama S, Imada K, Namba K, Minamino T (2018a) Insight into adaptive remodeling of the rotor ring complex of the bacterial flagellar motor. Biochem Biophys Res Commun 496:12–17
Kinoshita M, Namba K, Minamino T. (2018) Effect of a clockwise-locked deletion in FliG on the FliG ring structure of the bacterial flagellar motor. Genes Cells 23:241–247 (2018)
Kubori T, Shimamoto N, Yamaguchi S, Namba K, Aizawa S (1992) Morphological pathway of flagellar assembly in Salmonella typhimurium. J Mol Biol 226:433–446
Kuhlen L, Abrusci P, Johnson S, Gault J, Deme J, Caesar J, Dietsche T, Mebrhatu MT, Ganief T, Macek B, Wagner S, Robinson CV, Lea SM (2018) Structure of the core of the type III secretion system export apparatus. Nat Struct Mol Biol 25:583–590
Kutsukake K, Minamino T, Yokoseki T (1994) Isolation and characterization of FliK-independent flagellation mutants from Salmonella typhimurium. J Bacteriol 176:7625–7629
Lara-Tejero M, Kato J, Wagner S, Liu X, Galán JE (2011) A sorting platform determines the order of protein secretion in bacterial type III systems. Science 331:1188–1191
Lee KL, Ginsburg MA, Crovace C, Donohoe M, Stock D (2010) Structure of the torque ring of the flagellar motor and the molecular basis for rotational switching. Nature 466:996–1000
Lee PC, Zmina SE, Stopford CM, Toska J, Rietsch A (2014) Control of type III secretion activity and substrate specificity by the cytoplasmic regulator PcrG. Proc Natl Acad Sci USA 111:2027–2036
Li H, Sourjik V (2011) Assembly and stability of flagellar motor in Escherichia coli. Mol Microbiol 80:886–899
Lynch MJ, Levenson R, Kim EA, Sircar R, Blair DF, Dahlquist FW, Crane BR (2017) Co-folding of a FliF-FliG split domain forms the basis of the MS: C ring interface within the bacterial flagellar motor. Structure 25:317–328
Macnab RM (2004) Type III flagellar protein export and flagellar assembly. Biochim Biophys Acta 1694:207–217
Makino F, Shen D, Kajimura N, Kawamoto A, Pissaridou P, Oswin H, Pain M, Murillo I, Namba K, Blocker AJ (2016) The architecture of the cytoplasmic region of type III secretion systems. Sci Rep 6:33341
McDowell MA, Marcoux J, McVicker G, Johnson S, Fong YH, Stevens R, Bowman LA, Degiacomi MT, Yan J, Wise A, Friede ME, Benesch JL, Deane JE, Tang CM, Robinson CV, Lea SM (2016) Characterisation of Shigella Spa33 and Thermotoga FliM/N reveals a new model for C-ring assembly in T3SS. Mol Microbiol 99:749–766
McMurry JL, Murphy JW, González-Pedrajo B (2006) The FliN-FliH interaction mediates localization of flagellar export ATPase FliI to the C ring complex. Biochemistry 45:11790–11798
Meshcheryakov VA, Kitao A, Matsunami H, Samatey FA (2013) Inhibition of a type III secretion system by the deletion of a short loop in one of its membrane proteins. Acta Crystallogr D Biol Crystallogr 69:812–820
Minamino T (2014) Protein export through the bacterial flagellar type III export pathway. Biochim Biophys Acta 1843:1642–1648
Minamino T (2018) Hierarchical protein export mechanism of the bacterial flagellar type III protein export apparatus. FEMS Microbiol Lett 365:fny117
Minamino T, Macnab RM (1999) Components of the Salmonella flagellar export apparatus and classification of export substrates. J Bacteriol 181:1388–1394
Minamino T, Macnab RM (2000a) Domain structure of Salmonella FlhB, a flagellar export component responsible for substrate specificity switching. J Bacteriol 182:4906–4919
Minamino T, Macnab RM (2000b) FliH, a soluble component of the type III flagellar export apparatus of Salmonella, forms a complex with FliI and inhibits its ATPase activity. Mol Microbiol 37:1494–1503
Minamino T, Macnab RM (2000c) Interactions among components of the Salmonella flagellar export apparatus and its substrates. Mol Microbiol 35:1052–1064
Minamino T, Namba K (2008) Distinct roles of the FliI ATPase and proton motive force in bacterial flagellar protein export. Nature 451:485–488
Minamino T, Iino T, Kutuskake K (1994) Molecular characterization of the Salmonella typhimurium flhB operon and its protein products. J Bacteriol 176:7630–7637
Minamino T, Tame JRH, Namba K, Macnab RM (2001) Proteolytic analysis of the FliH/FliI complex, the ATPase component of the type III flagellar export apparatus of Salmonella. J Mol Biol 312:1027–1036
Minamino T, González-Pedrajo B, Oosawa K, Namba K, Macnab RM (2002) Structural properties of FliH, an ATPase regulatory component of the Salmonella type III flagellar export apparatus. J Mol Biol 322:281–290
Minamino T, Imada K, Namba K (2008) Mechanisms of type III protein export for bacterial flagellar assembly. Mol BioSyst 4:1105–1111
Minamino T, Yoshimura SDJ, Morimoto YV, González-Pedrajo B, Kami-ike N, Namba K (2009) Roles of the extreme N-terminal region of FliH for efficient localization of the FliH-FliI complex to the bacterial flagellar type III export apparatus. Mol Microbiol 74:1471–1483
Minamino T, Morimoto YV, Hara N, Namba K (2011) An energy transduction mechanism used in bacterial type III protein export. Nat Commun 2:475
Minamino T, Kinoshita M, Hara N, Takeuchi S, Hida A, Koya S, Glenwright H, Imada K, Aldridge PD, Namba K (2012a) Interaction of a bacterial flagellar chaperone FlgN with FlhA is required for efficient export of its cognate substrates. Mol Microbiol 83:775–788
Minamino T, Kinoshita M, Imada K, Namba K (2012b) Interaction between FliI ATPase and a flagellar chaperone FliT during bacterial flagellar export. Mol Microbiol 83:168–178
Minamino T, Morimoto YV, Kinoshita M, Aldridge PD, Namba K (2014) The bacterial flagellar protein export apparatus processively transports flagellar proteins even with extremely infrequent ATP hydrolysis. Sci Rep 4:7579
Minamino T, Morimoto YV, Hara N, Aldridge PD, Namba K (2016) The Bacterial flagellar type III export gate complex is a dual fuel engine that can use both H+ and Na+ for flagellar protein export. PLoS Pathog 12:e1005495
Moore SA, Jia Y (2010) Structure of the cytoplasmic domain of the flagellar secretion apparatus component FlhA from Helicobacter pylori. J Biol Chem 285:21060–21069
Morimoto YV, Minamino T (2014) Structure and function of the bi-directional bacterial flagellar motor. Biomolecules 4:217–234
Morimoto YV, Ito M, Hiraoka KD, Che Y-S, Bai F, Kami-ike N, Namba K, Minamino T (2014) Assembly and stoichiometry of FliF and FlhA in Salmonella flagellar basal body. Mol Microbiol 91:1214–1226
Morimoto YV, Kami-ike N, Miyata T, Kawamoto A, Kato T, Namba K, Minamino T (2016) High-resolution pH imaging of living bacterial cell to detect local pH differences. mBio 7:e01911-16
Notti RQ, Bhattacharya S, Lilic M, Stebbins CE (2015) A common assembly module in injectisome and flagellar type III secretion sorting platforms. Nat Commun 6:7125
Ohnishi K, Fan F, Schoenhals GJ, Kihara M, Macnab RM (1997) The FliO, FliP, FliQ, and FliR proteins of Salmonella typhimurium: putative components for flagellar assembly. J Bacteriol 179:6092–6099
Okabe M, Minamino T, Imada K, Namba K, Kihara M (2009) Role of the N-terminal domain of FliI ATPase in bacterial flagellar protein export. FEBS Lett 583:743–748
Park SY, Lowder B, Bilwes AM, Blair DF, Crane BR (2006) Structure of FliM provides insight into assembly of the switch complex in the bacterial flagella motor. Proc Natl Acad Sci USA 103:11886–11891
Paul K, Harmon JG, Blair DF (2006) Mutational analysis of the flagellar rotor protein FliN: identification of surfaces important for flagellar assembly and switching. J Bacteriol 188:5240–5248
Paul K, Erhardt M, Hirano T, Blair DF, Hughes KT (2008) Energy source of the flagellar type III secretion. Nature 451:489–492
Paul K, Gonzalez-Bonet G, Bilwes AM, Crane BR, Blair D (2011) Architecture of the flagellar rotor. EMBO J 30:2962–2971
Saijo-Hamano Y, Imada K, Minamino T, Kihara M, Shimada M, Kitao A, Namba K (2010) Structure of the cytoplasmic domain of FlhA and implication for flagellar type III protein export. Mol Microbiol 76:260–268
Terahara N, Inoue Y, Kodera N, Morimoto YV, Uchihashi T, Imada K, Ando T, Namba K, Minamino T (2018) Insight into structural remodeling of the FlhA ring responsible for bacterial flagellar type III protein export. Sci Adv 4:eaao7054
Terashima H, Kawamoto A, Tastumi C, Namba K, Minamino T, Imada K (2018) In vitro reconstitution of functional type III protein export and insights into flagellar assembly. mBio 9:e00988-18
Thomas DR, Morgan DG, DeRosier DJ (1999) Rotational symmetry of the C ring and a mechanism for the flagellar rotary motor. Proc Natl Acad Sci USA 96:10134–10139
Thomas J, Stafford GP, Hughes C (2004) Docking of cytosolic chaperone-substrate complexes at the membrane ATPase during flagellar type III protein export. Proc Natl Acad Sci USA 101:3945–3950
Ueno T, Oosawa K, Aizawa S (1992) M ring, S ring and proximal rod of the flagellar basal body of Salmonella typhimurium are composed of subunits of a single protein, FliF. J Mol Biol 227:672–677
Vartanian AS, Paz A, Fortgang EA, Abramson J, Dahlquist FW (2012) Structure of flagellar motor proteins in complex allows for insights into motor structure and switching. J Biol Chem 287:35779–35783
Wagner S, Königsmaier L, Lara-Tejero M, Lefebre M, Marlovits TC, Galán JE (2010) Organization and coordinated assembly of the type III secretion export apparatus. Proc Natl Acad Sci USA 107:17745–17750
Wagner S, Grin I, Grin I, Malmsheimer S, Singh N, Torres-Vargas CE, Westerhausen S (2018) Bacterial type III secretion systems: a complex device for the delivery of bacterial effector proteins into eukaryotic host cells. FEMS Microbiol Lett 365:fny201
Worrall LJ, Vuckovic M, Strynadka NCJ (2010) Crystal structure of the C-terminal domain of the Salmonella type III secretion system export apparatus protein InvA. Protein Sci 19:1091–1096
Xing Q, Shi K, Portaliou A, Rossi P, Economou A, Kalodimos CG (2018) Structure of chaperone-substrate complexes docked onto the export gate in a type III secretion system. Nat Commun 9:1773
Xue C, Lam KH, Zhang H, Sun K, Lee SH, Chen X, Au SWN (2018) Crystal structure of the FliF-FliG complex from Helicobacter pylori yields insight into the assembly the motor MS–C ring in the bacterial flagellum. J Biol Chem 293:2066–2078
Zarivach R, Deng W, Vuckovic M, Felise HB, Nguyen HV, Miller SI, Finlay BB, Strynadka NC (2008) Structural analysis of the essential self-cleaving type III secretion proteins EscU and SpaS. Nature 453:124–127
Zilkenat S, Franz-Wachtel M, Stierhof YD, Galán JE, Macek B, Wagner S (2016) Determination of the stoichiometry of the complete bacterial type III secretion needle complex using a combined quantitative proteomic approach. Mol Cell Proteomics 15:1598–1609
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2019 Springer Nature Switzerland AG
About this chapter
Cite this chapter
Minamino, T., Kawamoto, A., Kinoshita, M., Namba, K. (2019). Molecular Organization and Assembly of the Export Apparatus of Flagellar Type III Secretion Systems. In: Wagner, S., Galan, J. (eds) Bacterial Type III Protein Secretion Systems. Current Topics in Microbiology and Immunology, vol 427. Springer, Cham. https://doi.org/10.1007/82_2019_170
Download citation
DOI: https://doi.org/10.1007/82_2019_170
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-030-52122-6
Online ISBN: 978-3-030-52123-3
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)