Abstract
The human microbiota consists of bacteria, archaea, viruses, and fungi that build a highly complex network of interactions between each other and the host. While there are many examples for commensal bacterial influence on host health and immune modulation, little is known about the role of commensal fungi inside the gut community. Up until now, fungal research was concentrating on opportunistic diseases caused by fungal species, leaving the possible role of fungi as part of the microbiota largely unclear. Interestingly, fungal and bacterial abundance in the gut appear to be negatively correlated and disruption of the bacterial microbiota is a prerequisite for fungal overgrowth. The mechanisms behind bacterial colonization resistance are likely diverse, including direct antagonism as well as bacterial stimulation of host defense mechanisms. In this work, we will review the current knowledge of the development of the intestinal bacterial and fungal community, the influence of the microbiota on human health and disease, and the role of the opportunistic yeast C. albicans. We will furthermore discuss the possible benefits of commensal fungal colonization. Finally, we will summarize the recent findings on bacterial–fungal interactions.
M. Joanna Niemiec and Alexander Steimle have equally contributed to this work.
Access provided by Autonomous University of Puebla. Download chapter PDF
Similar content being viewed by others
1 Development and Composition of the Intestinal Microbiota
For a long time, it was assumed that the unborn child is sterile and that the initial contact with commensal microbes occurs during birth (Stinson et al. 2017). This led to the official doctrine that any microbes found in the uterine cavity must be pathological and hazardous to the unborn child. However, in the last few years a growing body of scientific studies reported the presence of either microbial DNA or viable microbiota in the placenta, the amniotic fluid, the meconium, or the umbilical cord blood. As mother–offspring pairs share microbial signatures between placenta, amniotic fluid, and meconium, it is suggested that the early gut colonization may be initialized prenatally by a distinct trajectory of maternal microbes (Kundu et al. 2017; Collado et al. 2016).
The establishment of the human intestinal microbiota is influenced by multiple factors. One important factor is the mode of delivery, which affects diversity and colonization pattern of the infant gut microbiota. Vaginally delivered infants harbor in all body habitats bacterial communities that are in composition most similar to the vaginal communities of their mothers (Dominguez-Bello et al. 2010). The microbiota of children delivered by cesarean section is most similar to the skin communities of their mothers (Dominguez-Bello et al. 2010). While dominant bacterial taxa found in vaginally delivered infants are Lactobacilli, Prevotella, Atopobium, or Sneathia spp., typical skin taxa in samples from caesarian section delivered babies include Staphylococcus, Corynebacterium, and Propionibacterium spp. as well as Klebsiella, Veillonella, and Clostridiaceae. (Dominguez-Bello et al. 2010; Kundu et al. 2017; Rutayisire et al. 2016). There is more and more evidence that the early microbiota colonization may influence the development of diseases later in life (Goulet 2015). For instance, a low microbial diversity in infancy precedes the onset of allergic disease (West 2014), and sensitization to food in children at three months and one year was correlated with a high abundance of Enterobacteriaceae and a low abundance of Bacteroidaceae in the gut microbiota (Azad et al. 2015). Furthermore, there is growing evidence that an aberrant gut microbe composition develops as result of the delivery mode, e.g., caesarian section affects the subsequent regulation of immune responses (Knight and Girling 2003; Rutayisire et al. 2016). It is reported that the risk of developing celiac disease, asthma, obesity, or type-1-diabetes (T1D) is increased in children delivered by caesarian section as compared to vaginally born infants (Black et al. 2015; Kuhle et al. 2015; Adlercreutz et al. 2015).
In addition, intake of colostrum and breast milk possibly supplies neonates with the unique microbial pattern of the mothers’ breast milk (Martin et al. 2007). Using this route, the most frequent genera which are vertically transferred from mother to child are Lactobacillus, Staphylococcus, Enterococcus, and Bifidobacterium (Kundu et al. 2017; Makino et al. 2011; Fernandez et al. 2013).
Upon introduction of dietary supplements, solid food, and withdrawal from breast milk, the intestinal microbiota acquires greater complexity, individuality and increasingly resembles the microbiota of a typical omnivore (Kundu et al. 2017; Palmer et al. 2007). In parallel with increasing age during adolescence and puberty, the population of aerobes and facultative anaerobes decreases gradually and the number of obligate anaerobes increases (Hopkins and Macfarlane 2002). In addition, gender-specific diversifications of the gut microbiota arise during this period of life (Markle et al. 2013). Moreover, the adolescent microbiome was found to differ functionally from that of adults, expressing genes related to development and growth (Hollister et al. 2015). The adult microbiota harbors a core community of colonizers and is more stable compared to that of early life (Palmer et al. 2007; Rajilic-Stojanovic et al. 2012). However, it is still amenable to changes by environmental perturbations such as changes in nutrition (David et al. 2014), seasonal variations or temperature fluctuations (Chevalier et al. 2015), or even altitude (Zhang et al. 2016). Changes in the composition of the intestinal microbiota also account for changes in gut metabolites. Using comparative metabolomics, clear differences between gut metabolite productions can be observed between humans following omnivore, vegetarian, vegan, or a Mediterranean diet (De Filippis et al. 2016; Wu et al. 2016). However, to which extent these metabolome differences are due to diet-associated differences in the microbiota or nutrient composition of the diet itself remains to be determined.
2 Microbiota in Health and Disease
The defined composition of the intestinal microbiota is a unique feature of each individual. As demonstrated above, this characteristic composition within each individual is not static, but rather subject to frequent changes due to environmental, nutritional, or iatrogenic influences.
The impact of the intestinal microbiota on the development of a wide variety of human pathologies is the subject of intense ongoing research. In this context, developing strategies to precisely modulate the microbiota composition in order to restore gut homeostasis in a host-specific manner is thought to offer novel therapeutic approaches for the treatment of microbiota-influenced human diseases. To achieve this ambitious aim, several fundamental questions have to be answered: (1) How exactly does a certain microbiota composition influence the progress or outcome of a distinct pathology on a molecular level? Answering this question might also lead to the discovery of novel, so far neglected host-specific therapeutic targets. (2) Does the presence of certain commensal microbes enhance the risk of developing a particular disease? (3) Is the presence of such a specific disease-promoting commensal a general phenomenon or is it restricted to a certain individual host or of host group sharing particular genetic predispositions? And finally: (4) Is there a host-specific distinct ideal microbiota composition that completely prevents potential disease-driving events?
One of the most intriguing observations supporting the hypothesis of microbiota-triggered human diseases is the effect of transplanting feces from human patients into germ-free (GF) mice. For example, fecal transplantation from multiple sclerosis (MS) patients into GF mice results in a more prominent experimental autoimmune encephalomyelitis (EAE) outcome compared to using feces from healthy donor controls (Berer et al. 2017). Correspondingly, fecal transplantations from patients suffering from irritable bowel syndrome (IBS) (De Palma et al. 2017) or Parkinson’s disease (PD) (Sampson et al. 2016) induced stronger disease phenotypes in germ-free susceptible mice compared to mice transplanted with feces from healthy human donors. However, as for many of these studies, it is not completely clear if this disease-inducing dysbiotic microbiota is the cause of the pathology or just a secondary consequence emerging from other disease-promoting events in the donor host.
There are diverse interactions between gut microbes and host cells at intestinal mucosal interfaces: (1) Commensal microbes can directly interact with host cells. (2) The host, in turn, reacts to microbes by either tolerance or inflammation, thereby potentially leading to microbiota composition shifts. (3) Members of the microbiota interact with each other, and (4) environmental factors can influence this complex interplay. It is important to note that these interactions are very dynamic and interconnected. Under homeostatic conditions, the different variables are counterbalanced leading to a highly diverse microbiota that contributes to health maintenance. However, changing one of these variables can promote dysbiosis, characterized by strongly reduced microbiota diversity, possibly leading to pathology-driving effects.
In the following, we will briefly summarize the different mechanisms by which the composition of microbiota impacts human health.
2.1 Interactions Between Bacteria
Usually, a balanced or homeostatic microbiota composition helps the host to combat infections with enteropathogenic bacteria, viruses, or fungi (Ubeda et al. 2017). Therefore, a microbiota shift toward dysbiosis might facilitate the establishment of infections caused by enteropathogens. It is important to note that, on one side, microbiota alterations due to external influences facilitate pathogen invasion and repopulation. On the other side, gastrointestinal (GI) infections affect the microbiota composition.
There are several mechanisms how certain microbiota members influence the survival or colonization properties of other intestinal microbes. These intermicrobial interactions strongly affect the overall microbiota composition leading to a highly diverse and balanced microbiota and helping to prevent overgrowth of pathobiotic or pathogenic microorganisms: Commensal microbes can impact survival or repopulation of other microbiota members either directly or indirectly via modulation of the host immune system leading to feedback effects on certain microbes (Buffie and Pamer 2013).
Some bacteria are reported to secrete molecules affecting the abundance of other intestinal bacteria. For instance, the commensal symbiont Bacteroidetes thetaiotaomicron secretes a factor that represses toxin production from enterohemorrhagic Escherichia coli strains (de Sablet et al. 2009). Comparably, certain Bifidobacterium spp. produce soluble factors which promote protection against pathogenic E. coli and Citrobacter rodentium strains (Gagnon et al. 2004). The probiotic strain E. coli Nissle 1917 secretes microcins limiting the expansion of pathobiotic or pathogenic Enterobacteriacaeae strains under inflammatory conditions, thereby helping to re-establish homeostasis (Sassone-Corsi et al. 2016). Bacillus thuringiensis was reported to inhibit Clostridium difficile by secretion of a strain-specific bacteriocin (Rea et al. 2010). Notably, these bactericins and microcins seem to act in a target-specific manner, not influencing other microbiota components (Sassone-Corsi et al. 2016; Rea et al. 2011).
Another important factor is the competition for adhesion sites and nutrients. For example, Lactobacillus reuteri expresses a mucus-binding protein facilitating adherence to the host mucosa (Kankainen et al. 2009). This effect is thought to contribute to limiting the primary adhesion of pathogens like Clostridium difficile. Bacteroidetes thetaiotaomicron metabolizes plant-derived monosaccharides, thereby limiting growth of the pathogen C. rodentium (Kamada et al. 2012). However, not only competition occurs at such mucosal interfaces but also synergistic effects are observed. For example, Bacteroidetes longum upregulates carbohydrate-metabolizing enzymes of Bacteroidetes thetaiotaomicron, thereby supporting this strain’s growth (Sonnenburg et al. 2006).
Additionally, microbiota components indirectly impact repopulation of other consortium members via modulation of the host immune system. Such modulation includes, among others, the induction of antimicrobial peptide (AMP) secretion, the production of antimicrobial C-type lectins (Sonnenburg et al. 2006), induction of plasma cell-derived IgA (Ivanov et al. 2009), or modulation of T cell-mediated immunity (Gaboriau-Routhiau et al. 2009).
2.2 Intestinal Epithelial Barrier and Translocation of Microbiota Components
The intestinal lumen and its content are separated from underlying sterile tissues by the intestinal mucosal barrier. This barrier is a dynamic structure that allows for communication between microbiota components, metabolites, and the host organism; facilitates absorption of nutrients; and aids in sensing invading pathogens (Bischoff et al. 2014). It is built up by intestinal epithelial cells (IECs), a mucus layer covering IECs at the luminal side, and antimicrobial peptides (AMPs) (Antoni et al. 2014). The integrity of this barrier is of crucial importance since it prevents from uncontrolled translocation of luminal content into other body compartments, which could result in local or systemic inflammation and disease (Valentini et al. 2014).
Barrier function is regulated by both the microbiota and host immune cells. IECs as well as immune cells express pattern recognition receptors (PRRs), such as Toll-like receptors (TLRs), mediating detection of distinct bacterial components, called microbe-associated molecular patterns (MAMPs). PRR-mediated detection of MAMPs by IECs results in the activation of intracellular signaling pathways leading, for instance, to secretion of AMPs which, in turn, contribute to intestinal homeostasis (Ayabe et al. 2000; Vaishnava et al. 2011). Therefore, defects in PRR-mediated signaling in IECs increase susceptibility to intestinal infections due to reduced secretion of AMPs but also decreased mucus formation (Frantz et al. 2012; Bhinder et al. 2014). Mucus is produced by goblet cells, and the resulting layer prevents the over-activation of host cells by separating the luminal content from the intestinal epithelium (Fig. 1). An additional factor of barrier integrity is the presence of tight junction proteins which mediate impermeability of the IECs cell layer. Commensal bacteria contribute to barrier integrity through the production of certain metabolites. For example, the generation of short-chain fatty acids (SCFAs) promotes mucus secretion from goblet cells (Burger-van Paassen et al. 2009), while indole, generated from tryptophan by tryptophanase-expressing bacteria, supports tight junction protein function by activating the pregnane X receptor on IECs (Shimada et al. 2013; Venkatesh et al. 2014).
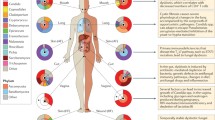
Mechanisms of microbiota-mediated influences on host pathologies. The composition of the intestinal microbiota is influenced by external factors: Intermicrobial interactions and colonization resistance effects influence the microbiota’s ability to fight enteropathogenic infections. Translocation of bacteria or bacterial components affects diseases such as type-2-diabetes (T2D) or can cause inflammatory reactions. Metabolites produced by the intestinal microbiota impact the progress of, e.g., liver disease (LD), colonic cancer (CC), or T2D. These metabolites also affect the integrity of the intestinal barrier and T cell-mediated immunity. Microbe-associated molecular patterns (MAMPs) derived from intestinal commensals or pathogens are sensed by host pattern recognition receptors (PRRs), subsequently influencing B cell- and T cell-mediated immunity, mainly via activation of intestinal dendritic cells (DCs) which are located within the lamina propria. These DCs influence the transepithelial secretion of IgA by plasma cells, which, in turn, impacts the intestinal microbiota. Microbiota-mediated influence of intestinal B cell immunity is mediated either T cell-dependent or T cell-independent. Surface expression of T cell-activating molecules and the cytokine secretion pattern of DCs directs polarization of naïve CD4+ T cells into various different T cell phenotypes: T helper 1 (Th) 1, Th2, regulatory T cells (Treg), pathological (path) or protective (prot) Th17, anti-Th17 Tregs or Th1-like Th17 cells. Transcription factors and key cytokines characterizing each subpopulation are indicated in white boxes. Path Th17 and Th1-like Th17 cells can influence the pathology of autoimmune diseases such as multiple sclerosis (MS), inflammatory bowel disease (IBD), type-1-diabetes (T1D) or rheumatoid arthritis (RA). Prot Th17 cells help to maintain the integrity of the intestinal epithelium. These adaptive immune responses can also affect the microbiota composition. Illustration by A. Steimle
The integrity of the epithelial barrier is influenced not only by microbial metabolites or MAMPs but also by host immune cells. Main mediators of this are cytokines, secreted by various leukocytes. In response to mucosal injury, host immune cells secrete IL-6 which promotes IEC proliferation, therefore, contributing to wound healing (Kuhn et al. 2014). A comparable effect is observed for IL-5 and IL-13, which induce cell proliferation via interposed macrophage activation (Seno et al. 2009). On the other hand, pro-inflammatory cytokines such as TNF or IFNγ have opposite effects, weakening barrier integrity, i.e., through the suppression of β-catenin/T cell factor signaling (Nava et al. 2010). Importantly, all these host immune cell-mediated effects can be induced by microbiota components, i.e., through MAMP sensing by PRRs (Fig. 1), representing an indirect way of microbiota-mediated influences on barrier integrity.
Dysfunction of the intestinal epithelial barrier and the resulting increased permeability have been linked to various pathologies and were observed in patients with inflammatory bowel disease (IBD), irritable bowel syndrome (IBS), various liver diseases (LD), acute pancreatitis (AP), type-1-diabetes (T1D) and type-2-diabetes (T2D), chronic kidney disease, depression, and other diseases, reviewed in (Fukui 2016). However, it is often unclear whether an increase in barrier permeability and subsequent bacterial translocation is just a secondary effect of ongoing inflammatory processes or a primary pathology-inducing effect.
However, in some cases, an association between translocated intestinal bacteria and onset of disease has been reported, i.e., for type-2-diabetes (T2D) (Sato et al. 2014) and liver disease (LD) (Gkolfakis et al. 2015). It is furthermore hypothesized that even the blood–brain barrier permeability is affected by the intestinal microbiota (Lavasani et al. 2010), therefore influencing neurodegenerative diseases and psychiatric conditions. This effect was suggested to be due to bacterial translocation under microbiota dysbiosis, finally leading to inflammatory reactions in the central nervous system (CNS) (Maes et al. 2012, 2013).
2.3 Microbial Metabolites
Among others, the host benefits from intestinal microbes by their ability to digest certain nutrients which cannot be metabolized by the host itself. Certain microbe-generated metabolites furthermore provide profound effects on the health of the host organism, such as vitamin B or K. Best studied are the effects of short-chain fatty acids (SCFAs) and long-chain fatty acids (LCFAs). SCFAs are beneficial for the host by providing nutrition for enterocytes, stimulating epithelial proliferation, and promoting epithelial barrier function. Furthermore, SCFAs promote tolerance of the intestinal immune system (de Goffau et al. 2014), while LCFAs promote the expansion of inflammation-triggering Th1 and Th17 cells (Haghikia et al. 2015). In addition, some SCFAs have a direct impact on intestinal pathobionts and thereby contribute to colonization resistance.
Microbe-derived metabolites do not only affect the intestinal tissue but also other organ systems, most prominently the liver. For example, SCFAs positively affect glucose homeostasis (Lin et al. 2012). On the other hand, not only bacterial endotoxins but also bacterial metabolites such as ethanol, ammonia, or acetaldehyde reach the liver via the gut–liver axis where they might consequently affect liver function (Gkolfakis et al. 2015). This illustrates that alterations in microbiota composition can lead to changes in metabolite production and thereby affects pathologies such as obesity, dyslipidemia and their subsequent influence on type-2-diabetes.
2.4 Interaction of Microbiota and the Immune System
The intestinal microbiota provides a considerable influence on the host immune system. This is exemplified by experiments with GF mice which showed that maturation of the immune system requires the presence of intestinal microbiota (Hamada et al. 2002; Fukata and Arditi 2013; Shi and Mu 2017).
This effect is mediated not only by the indirect mechanisms outlined above but also by direct interactions with the intestinal immune system. MAMPs play a pivotal role in activating or silencing the host’s innate as well as the adaptive immune system. Several cell types at GI mucosal interfaces express PRRs including enterocytes and phagocytes. PRRs include cytoplasmic and membrane-associated TLRs, C-type lectin receptors, NOD-like receptors, and RIG-I-like receptors. PRR activation can result in pro- or anti-inflammatory responses, depending on the MAMP being recognized. One of the most prominent MAMPs is lipopolysaccharide (LPS) of Gram-negative bacteria which is sensed by the host TLR4/MD-2 receptor complex. Interestingly, some commensal bacteria such as many Bacteroides spp. harbor lipopolysaccharide structures which fail to activate the TLR4/MD-2 receptor complex. This is thought to contribute to tolerate in the presence of beneficial commensal bacteria. However, PRR-mediated MAMP sensing is important for recognition and subsequent clearance of enteropathogens as well as for epithelial cell proliferation, maintenance of tight junctions, and AMP release as outlined above (Buffie and Pamer 2013; Sharma et al. 2010).
Importantly, PRRs are not evenly expressed on IECs but display a polarized distribution pattern. Extracellular TLRs, for instance, are usually expressed only at the basolateral plasma membranes of enterocytes (Yu and Gao 2015). This appears to be necessary in order to maintain immune balance and avoid over-activation of innate and adaptive immune responses since PAMPs are effectively sensed when luminal content crosses the epithelial barrier but are rarely recognized at the luminal side of enterocytes.
Besides enterocytes, PRRs are also expressed by phagocytes. Among them, intestinal dendritic cells (DCs) fulfill a crucial role by linking the innate with the adaptive immune system. Intestinal DCs can sense microbes either with participation of neighboring goblet cells (McDole et al. 2012), with the support of CX3CR1+ phagocytic cells (Mazzini et al. 2014), or via direct sampling of luminal antigens by DC dendrites extending through the epithelium into the lumen (Farache et al. 2013). DCs play an important role in establishing a protective adaptive immunity against pathogens. Furthermore, they crucially contribute to tolerance of commensal bacteria and food antigens (Shiokawa et al. 2017). DCs impact B cell- as well as T cell-mediated immune responses via the expression of certain surface molecules or secreted proteins. This process is tightly regulated and dependent on the sampled antigen. DCs can either display a mature phenotype which is characterized by strong expression of T cell-activating surface molecules and pro-inflammatory cytokines leading to inflammatory processes, or a tolerogenic phenotype favoring maintenance of immune homeostasis (Steimle and Frick 2016).
T cell-mediated immunity shaping is one of the main consequences resulting from activation of intestinal DCs. Naïve CD4+ T cells encountering activated DCs can develop into different T cell subsets. These subsets are characterized by the expression of certain transcription factors, surface markers, or the secretion of key cytokines which are T-bet and IFNγ for Th1 cells, GATA-3 and IL-4 for Th2 cells, or Foxp3 and CD25 for Tregs. Th17 cells play an ambivalent role. While ROR\(\upgamma\)t+IL-17+IL-10-Th17 cells are attributed to inflammation-driving properties, ROR\(\upgamma\)t+IL-17-IL-10+Th17 cells are reported to have homeostasis-promoting functions, i.e., by supporting the integrity of the intestinal barrier. Additionally, ROR\(\upgamma\)t+Th17 cells can develop into inflammation-promoting so-called ROR\(\upgamma\)t+IFN\(\upgamma\)+Th1-like Th17 cells (Fig. 1). This dichiotomic role of Th17 cells was recently excellently reviewed by Stockinger et al. (Stockinger and Omenetti 2017).
Concerning adaptive immune responses, T cell populations are not the sole immune cells being affected by the intestinal microbiota. B cell populations, i.e., intestinal plasma cells, are also influenced by luminal microbes. Intestinal plasma cells secrete IgA which is transported transepithelially into the GI lumen. Here, they coat bacteria leading to reduced bacterial interaction with host immune components. Interestingly, pathobiotic commensals are preferably coated with plasma cell-derived IgA. This process strongly contributes to homeostasis and antagonizes dysbiosis. Intestinal plasma cell-induced IgA secretion can be either T cell-mediated or T cell-independent (Fagarasan et al. 2010). IgA-producing plasma cells are influenced indirectly, via a T cell-dependent regulation loop, or directly by intestinal dendritic cells, i.e., via microbiota-dependent regulation of the expression of B cell-influencing factors such as BAFF or APRIL (Tezuka et al. 2011) (Fig. 1).
Based on these implications, it does not seem surprising that the intestinal microbiota impacts the outcome or progression of inflammatory disorders, especially autoimmune diseases. However, little is known about the definite mechanisms by which the microbiota specifically affects disease progress. Prominent contributing factors include a disturbed microbiota-mediated regulation of cathepsin S expression and/or activity, a protease of antigen-presenting cells (APCs) influencing subsequent T cell activation, and a dysregulated systemic Th17 response which is strongly affected by the local intestinal immune system (Steimle et al. 2016). Apart from Th1 and Th2 responses, characterized mainly by IFNγ and IL-4 secreting CD4+ T cells, respectively, a so-called Th17 response has gained more and more attention when it comes to microbiota-mediated immunological disorders. Th17 cells are found primarily in the gut under homeostatic conditions (Ivanov et al. 2009). However, they seem to be able to migrate to other organs or compartments under dysbiotic conditions further promoting disease progress (Krebs et al. 2016). The Th17 response plays an ambivalent role in this context since Th17 cells may express significantly different properties, depending on their particular cytokine secretion pattern, from disease-driving to protective functions. Furthermore, the outcome of experimental autoimmune encephalomyelitis (EAE), a mouse model for MS, is highly dependent on the microbiota composition, and enhanced Th17 cell activation is associated with a more severe pathology in this model (Lee et al. 2011).
Microbiota-triggered T cell immune responses are in fact shared among all microbiota-influenced autoimmune disorders (AIDs) where they contribute strongly to the extent of the respective pathology. In fact, AIDs such as MS, IBD, rheumatoid arthritis (RA), and T1D are linked to dysregulated Th17 responses in human patients (Chen et al. 2016; van Langelaar et al. 2018; Tang et al. 2017; Stadhouders et al. 2018; Fores et al. 2018; Baharlou et al. 2016), an association that is experimentally supported in mouse models for these diseases (Lee et al. 2011; Wu et al. 2010; George et al. 2016).
T cell immunity is strongly shaped by the intestinal microbiota, as demonstrated by the complete absence of Th17, Th1, and regulatory T cells (Tregs) in GF mice (Geuking et al. 2011). Tregs aggregate in the mammalian intestine helping to maintain the host’s immune homeostasis (Littman and Rudensky 2010; Ivanov et al. 2008). The importance of these cells is underlined by the observation that a depletion of Tregs results in abnormal repopulation of microbiota-directed CD4+ T cells leading to pathological inflammatory reactions (Kamada et al. 2013). AID patients often exhibit intestinal irregularities such as reduced numbers of intestinal Tregs resulting in enhanced inflammatory reactions or impaired gut barrier function (Wu and Wu 2012; Rosser and Mauri 2016). This is often traced back to a general microbiota dysbiosis in these individuals, characterized by reduced α-diversity. In fact, human AID patients provide a distinct microbiota composition compared to healthy controls. This was, for example, demonstrated in patients suffering from T1D (Giongo et al. 2011; d’Arminio Monforte et al. 2014; Davis-Richardson et al. 2014; Li and Atkinson 2015), MS (Bhargava and Mowry 2014; Miyake et al. 2015), RA (Scher et al. 2013; Maeda et al. 2016; Liu et al. 2013), IBD, or systemic lupus erythematosus (SLE) (Hevia et al. 2014). Of course, most of these studies are correlation-based studies, not elucidating if microbiota dysbiosis is cause or consequence of the disease. Usually, it is assumed that a certain microbiota composition is not the exclusive cause of such pathologies since the induction of the disease usually requires a certain predisposition, which is mostly genetic, and/or specific environmental influences.
Most of the microbiota-related research, especially concerning the influence of intestinal microbes on human health, focuses on commensal and enteropathogenic bacteria, and their direct or indirect impact on the host. Surprisingly little is however known about interactions between members of the intestinal microbe consortium and how these interactions affect microbiota composition and the host. One group of understudied organisms in this context are fungi.
3 Commensal Fungi as Part of the Microbiota
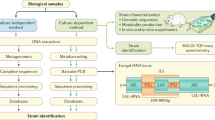
Currently, studies of the human microbiota often focus on bacteria only. Historically, this can be mostly explained by technical reasons. On the one hand, the range of microbes detected in culture-based analyses depends on the media used and thereby excludes viruses and most protozoa. While many fungi do grow on media for bacterial cultivation, they are vastly outnumbered by bacteria in the healthy gut (Qin et al. 2010), and thus difficult to assess quantitatively. Sequencing-based approaches, on the other hand, are often technically biased by the DNA extraction protocols toward isolation of bacterial genomes (Underhill and Iliev 2014), and targeted sequencing of 16S rDNA limits the range of included microbes to bacteria.
While it is true that fungi are vastly outnumbered by bacteria (Qin et al. 2010), one has to keep in mind the size and volume differences between bacteria and fungi and the resulting differences in biomass. It was estimated that the average bacterium relates to a yeast cell as a human to an elephant (Chaudhuri 2016). In addition, fungi might fill a unique niche by producing metabolites specific to fungi. And in fact, recent studies dissecting the mycobiome identified 66 genera of fungi present in human stool samples of which Saccharomyces, Candida, and Cladosporium were the most abundant genera (Hoffmann et al. 2013). Still, how the mycobiome is shaped and how it influences the human host is largely unknown.
3.1 The Birth of a Mycobiota Community
Similar to the bacterial microbiota, the human mycobiota is inherited during and after birth directly from the mother and other individuals living in close contact (Bliss et al. 2008; Nagata et al. 2012). Studies on the mycobiota of newborns revealed that the most abundant fungal species found in the intestinal tract were Candida parapsilosis, C. tropicalis, C. albicans, Saccharomyces cerevisiae, and C. orthopsilosis, corresponding to the mother’s vaginal microbiota. For vaginally born babies, C. albicans was the most dominant fungus on the skin (Juyal et al. 2013; Ward et al. 2018). After birth, the intestinal fungal population within an individual is subject to changes influenced by different internal and external factors. In contrast to the bacterial population, in which diversity reaches a maximum at adulthood and declines with old age (Yatsunenko et al. 2012), the mycobiota seems to follow an opposite trend: Diversity seems to be lowest during adulthood and higher in infants and the elderly (Strati et al. 2016) (Fig. 2). Cross-kingdom competition and colonization resistance mediated by the complex bacterial microbiota likely contribute to this, but according to our knowledge no studies have directly addressed this hypothesis. External factors shaping our entire microbiota and likely also affecting our mycobiota are, for instance, lifestyle (Evans et al. 2014; Misic et al. 2015), nutrition (David et al. 2014), and external acquisition of new microorganisms (Strati et al. 2016).
Diversity of bacteria and fungi in the gut over time. The diversity of the microbiota is changing with age. Fungi are dominant after birth when bacterial diversity is low. After bacterial species become more abundant, the fungal diversity drops. This trend is reversed with high age. Illustration by M. Kapitan
3.2 Yeasts Commonly Associated with Humans
Some Saccharomyces and Candida spp. are commonly associated with humans. Among those, some species are thought to be strictly host-associated, while others have also environmental niches or purpose during food production. One example for the latter is the baker’s yeast S. cerevisiae. This yeast is probably the fungus that had the largest impact on human life, culture, society, and science (Chambers and Pretorius 2010) and is one of the dominant fungal species found in the gastrointestinal (GI) tract (Hoffmann et al. 2013). Its environmental niche seems to be fruit, especially grapes, on which S. cerevisiae ferments sugars if the peel is damaged (Goddard 2008). This fermentation capacity has been exploited by humans for the production of bread and alcoholic beverages. Because of its natural niche on fruit and its widespread use in food production, it is not surprising that this yeast gets introduced frequently into the human GI tract. S. cerevisiae, like other environmental fungi, survives passaging through the GI tract (Strati et al. 2016). It is, however, not fully clear whether occurrence of S. cerevisiae in stool indicates transient passage or whether S. cerevisiae permanently resides in the gut. Because of its widespread and safe utilization, S. cerevisiae was even considered to be used as a probiotic (Pennacchia et al. 2008). A close relative, the yeast S. boulardii, is already used as a commercially available probiotic (Kelesidis and Pothoulakis 2012). As already demonstrated for the first time in the 1980s in animal and human experiments, its major benefits seem to be the reduction of Clostridium difficile growth (Massot et al. 1984) and positive effects on the intestinal immune system (Corthier et al. 1986).
The genus Candida comprises over 150 related species of which 15 are known to cause opportunistic infections in humans. Out of these, the most common causes of disease are C. albicans, C. glabrata, C. parapsilosis, C. tropicalis, and C. krusei (Yapar 2014). C. parapsilosis, C. tropicalis, and C. krusei account for approximately 17, 10, and 2% of fungal bloodstream infections worldwide, respectively (Pfaller et al. 2011). While these fungi can be found as part of the human microbiota, for instance, C. tropicalis in the gut (Roilides et al. 2003) and C. parapsilosis on the skin (Trofa et al. 2008), they all occur also in the environment (Carruba et al. 1991; Yang et al. 2012). The carriage rate of C. krusei in humans seems to be quite low, yet it is an important fermenter of cacao seeds (Nielsen et al. 2005) and therefore has an environmental niche intertwined with humans. C. tropicalis is widely present in soil (Yang et al. 2012), and C. parapsilosis can be found in a variety of environmental niches such as soil, water, plants, and insects (Gadanho and Sampaio 2005; Medeiros et al. 2008; Suh et al. 2008).
In contrast, C. albicans and C. glabrata, the two most commonly isolated Candida species associated with candidiasis, are found predominantly or even exclusively in association with warm-blooded hosts. C. glabrata is genetically more closely related to S. cerevisiae than to other Candida species (Kurtzman and Robnett 1998). It grows exclusively as yeast and is a colonizer of the human oral cavity and GI tract (Anderson 1917). Interestingly, colonization rates are rather low but positively correlated with age (Malani et al. 2011). Whether an environmental niche for C. glabrata exists is still debated (Gabaldon and Carrete 2016). It is associated with birds from where it could be transmitted directly or indirectly to humans or other animals, but it is not yet fully clear if these hosts are the natural niche or just a temporary reservoir for this fungus. One study found C. glabrata on spoiled oranges (Koc et al. 2007), but that might be due to contamination by bird droppings. C. albicans has been found only in association with warm-blooded animals including humans. It is present on most mucosal surfaces like the GI and urogenital tract, the mouth, but also the skin of at least 60% of healthy individuals (Calderone and Clancy 2012). In the absence of underlying risk factors, infections with C. albicans are usually superficial. Oral infections, also known as thrush, and skin infections, such as diaper’s rush, are found especially in infants, in which a stable microbiota has yet to develop (Mohamadi et al. 2014). Oral candidiasis occurs predominantly in HIV-positive individuals, but also in the elderly, where it is often associated with oral prostheses (Bianchi et al. 2016), which may influence local oral microbiota composition. 75% of otherwise healthy women experience at least one vaginal infection in their life, often following the use of antibiotics and/or oral contraceptives that influence the vaginal environment (Brown et al. 2012). Severe, life-threatening disseminated Candida infections usually only occur in immunocompromised individuals. In addition to immunosuppression, indwelling catheters (as a source of biofilm-related infections), and traumatic or surgical breaches of the intestinal barrier are common risk factors for candidiasis (Yapar 2014). It was demonstrated that the majority of C. albicans bloodstream infections (BSIs) are caused by endogenous strains from the patient’s own gut (Odds et al. 2006), illustrating the role of the gut as a source for opportunistic fungal infections. Furthermore, antibiotic treatment predisposes not only for mucosal but also systemic Candida infections (Pappas 2006). Depletion of the bacterial microbiota is frequently associated with enhanced fungal colonization which in turn correlates with the likelihood of the development of candidiasis (Azevedo et al. 2015). On the other hand, it was demonstrated that a significant proportion of C. albicans infections are polymicrobial, so Candida is accompanied by bacteria in the respective specimen (Kim et al. 2013; Pulimood et al. 2002; Bouza et al. 2013; Klotz et al. 2007; Hermann et al. 1999). This clearly indicates that bacterial–fungal interactions could play an important role in the development of invasive candidiasis. As C. albicans is by far the best-studied Candida species, we will in the following focus on this species to discuss what is known so far about the life of Candida in the gut, and its interactions with the host and bacteria in this niche.
3.3 Life of Candida Albicans in the Gut
The intestinal environment varies significantly along the length of the GI tract and across the diameter of the intestine (Zheng et al. 2015; Khutoryanskiy 2015; Fischbach and Sonnenburg 2011; Flint et al. 2012). The astonishing transcriptional and metabolic versatility of C. albicans allows the fungus to thrive within these diverse environments and also facilitates survival during local and systemic infections (Brown et al. 2014; Ene et al. 2014). Interestingly, the expression pattern of C. albicans colonizing the murine gut appears to be distinct from that observed during tissue invasion (Rosenbach et al. 2010; Thewes et al. 2007; Walker et al. 2009; Zakikhany et al. 2007), suggesting a specific adaptation to a commensal lifestyle in this host niche. This is supported by studies that analyzed the role of distinct transcription factors for C. albicans survival in different host niches. For example, C. albicans Efg1 is important for virulence during systemic candidiasis, but in the gut, EFG1-deficient C. albicans strains were found to be more efficient colonizers (Pierce et al. 2013). Contrarily, the transcription factor Efh1 appears to actively promote C. albicans commensalism but is dispensable for systemic infections (White et al. 2007). Reciprocal roles for transcription factors have also been described in the context of iron acquisition: The transcriptional repressor Sfu1, which reduces iron uptake, is required for the persistence of C. albicans in the gut. In contrast, the positive regulator Sef1, mediating enhanced iron acquisition, is essential for virulence in a mouse model of systemic infection (Chen et al. 2011).
Another important feature of C. albicans that appears to be differentially regulated in the gut, compared to other host niches, is morphogenesis. One of C. albicans’ most remarkable features is its morphological flexibility (Noble et al. 2017). The typical growth form at temperatures at or below 30 °C in the laboratory is the budding yeast, also known as “white cells.” Upon certain environmental cues (e.g., shift to 37 °C, elevated levels of CO2, contact to surfaces, serum components), C. albicans is, however, able to filament and form pseudohyphae or true hyphae (Noble et al. 2017; Sudbery 2011; Wang and Xu 2008). While yeasts proliferate faster, hyphae are capable of penetrating tissue, leading to invasion (Berman and Sudbery 2002) and also contributing to immune evasion (Torosantucci et al. 2004). Therefore, it is not surprising that filamentation is associated with virulence in systemic, but also mucosal infections (Katagiri et al. 2018). Interestingly, hyphae are rarely observed in the gut, even though this niche provides strong filamentation cues such as body temperature, high CO2, and contact. Consequently, this observation implies that even stronger hyphal repression signals occur in this niche. While it is tempting to speculate that members of the microbial gut community may contribute to filamentation repression, filamentation was not observed in GM mice colonized with only C. albicans (Böhm et al. 2017), suggesting that it is the intestinal environment, rather than the microbiota, that promotes growth in the yeast form. Furthermore, several studies found that genetically forcing C. albicans into the filamentous form reduces colonization levels (Böhm et al. 2017; Rosenbach et al. 2010; Vautier et al. 2015).
In addition to the “white cell” morphotype C. albicans can undergo transition into several other yeast phenotypes that are metabolically distinct from each other. Opaque cells, for instance, grow as elongated, spiky single cells bigger than white cells and are able to mate (Hickman et al. 2013). The white to opaque transition is regulated by a network of different transcription factors of which the two main regulators are Efg1 and Wor1, which repress each other (Morschhäuser 2010). While Wor1 promotes cells to switch to the opaque phenotype, Efg1-dominated cells are white. At 37 °C, however, opaque cells are instable and switch back to the white form (Slutsky et al. 1987), which makes mating inside the gut niche unlikely.
Recently, two other phenotypes have been discovered. The “gray cell” type is interconnected with the white–opaque transition, but is independent of the mating-type locus (Tao et al. 2014). While the role of opaque and gray cells for commensalism remains unclear, the morphotype GI-induced transition (GUT) was observed after multiple transitions of C. albicans through the mouse gut. Of note, similar to opaque cells, overexpression of Wor1 is necessary for the switch to GUT cells. However, GUT cells are elongated but smooth compared to opaque cells and thus morphologically distinct (Pande et al. 2013). GUT cells outcompete white cells in a mouse colonization model probably due to improved adaptation to the gut niche and a more efficient metabolism (Pande et al. 2013). Taking into consideration that Efg1 is involved in filament formation (Stoldt et al. 1997) and low Efg1 levels in C. albicans promotes colonization of the mouse gut (Pierce et al. 2013), it appears that these different yeast cell types have distinct roles in different host niches, regulated by Efg1 and Wor1. While Efg1 is essential for yeast-to-hypha transition and therefore virulence (Lo et al. 1997), Wor1 seems to play a major role during commensalism and therefore colonization.
Remarkably, although C. albicans grows predominantly as yeast in the murine intestinal tract, it was shown that the local population expressed several genes that were previously assumed to be exclusively involved in or associated with hyphal growth (Herwald and Kumamoto 2014; Rosenbach et al. 2010). Furthermore, no filamentation was observed in GF mice while earlier studies reported a low level of filamentation in the gut of antibiotic-treated, C. albicans-colonized mice (Böhm et al. 2017). This could indicate that a tightly regulated expression of hypha- and virulence-associated fungal attributes in the gut is supporting C. albicans during its interaction with the microbiota.
3.4 Antagonistic Interactions Between Commensal Bacteria and Candida Albicans—Limiting Fungal Colonization
The potential of the commensal microbiota to limit fungal proliferation in the gut appears obvious given the clinical observation that depletion of the commensal microbiota by prolonged use broad-spectrum antibiotics frequently results in fungal overgrowth (Ianiro et al. 2016; Eggimann et al. 2015; Mason et al. 2012b; Erb Downward et al. 2013; Pasqualotto et al. 2006). Colonization resistance by the commensal microbiota can be achieved in different ways, as outlined in Sect. 2.1. However, whether distinct members of the microbiota are responsible for colonization resistance against C. albicans and by which mechanisms this is mediated is not yet well understood.
One of the best-studied examples for antagonistic Candida–bacteria interactions are lactic acid bacteria (LAB). They are characterized by their ability to ferment glucose to lactic acid, but they also produce bactericins and anti-fungal peptides (De Vuyst and Leroy 2007) (Fig. 3). Lactobacilli are the dominant species of the vaginal microbiota where they prevent urogenital infections by other microbes (Vasquez et al. 2002), including C. albicans, by acidification of the vaginal mucosa through lactic acid (Aroutcheva et al. 2001).
Antagonistic interactions between fungi and bacteria. Top: Bacteroidetes species can stimulate intestinal cells to produce more HIF1α which leads to secretion of antimicrobial peptides (AMPs) that in turn promote colonization resistance against C. albicans in a mouse model. Lactobacilli can secrete short-chain fatty acids which are the main source of energy for enterocytes and inhibit filamentation and invasiveness of C. albicans in vitro. Right: E. faecalis can secrete a small peptide termed EntV that inhibits filamentation in nematodes. Bottom: Salmonella species and Acinetobacter (A.) baumanii directly bind to the filamentous form of C. albicans which leads to killing of the fungus in vitro. Left: E. coli secretes an anti-fungal peptide during magnesium depletion that inhibits fungal growth in vitro. Illustration by M. Kapitan
Work by Mason et al. aimed to clarify how C. albicans and lactobacilli interact in the gut and stomach after antibiotic treatment in mice. In the presence of C. albicans, lactobacilli are repressed in their capacity to recolonize the stomach after antibiotic treatment is discontinued. Instead of lactobacilli, enterococci became the dominant species. This effect was reversed when fungal cells were absent (Mason et al. 2012a). Similar results were observed in the colon, where the fungus also led to faster recovery of the Bacteroidetes population (Mason et al. 2012b). One possible explanation is that C. albicans directly antagonizes lactobacilli: Yet again, fungal promotion of enterococcal growth, which then leads to repression of other Gram-positive bacteria by secretion of bactericins, could also explain the observations (Sawa et al. 2012). Lactobacilli can potentially antagonize C. albicans by the production of SCFAs, mainly acetate, propionate, and butyrate (Pessione 2012). SCFAs not only promote intestinal barrier function, they have also been shown to inhibit C. albicans filamentation and therefore invasiveness (Noverr and Huffnagle 2004) (Fig. 3). However, in the colon lactobacilli usually comprise less than 2% of the overall microbiota and the majority of SCFAs are produced by a few other bacterial taxa and species. The most important producer for propionate, for example, is the mucus-degrading bacterium Akkermansia muciniphila (Derrien et al. 2004), while butyrate is mainly produced from resistant starch by Ruminococcus bromii and a few other species (Louis et al. 2010; Ze et al. 2012). Yet, lactobacilli-derived molecules might contribute significantly to colonization resistance in distinct intestinal niches or in distinctly composed microbiota. Also, effects mediated by other colonization resistance mechanisms appear possible.
Besides lactobacilli, other anaerobic intestinal bacteria can mediate colonization resistance. In the presence of Gram-negative bacteria, IECs and Paneth cells secrete RegIIIγ, a C-type lectin which is highly active against Gram-positive bacteria (Cash et al. 2006). Experiments in mice showed that this immune stimulation can protect the host against vancomycin-resistant Enterococcus faecium colonization (Brandl et al. 2008). Similar mechanisms have been found for fungi: Fan et al. (Fan et al. 2015) demonstrated that the presence of two anaerobic bacterial species, Bacteroides thetaiotamicron and Blautia producta, was sufficient to clear C. albicans from the mouse gut. In the presence of fungi, these bacteria stimulated intestinal cells to express more HIF1α, a regulator of the innate immune system (Nizet and Johnson 2009) promoting expression of cathelicidins, a group of antimicrobial peptides (Peyssonnaux et al. 2005). In mice co-colonized with C. albicans and either of these anaerobic bacteria, HIF1α and the cathelicidin LL-37-CRAMP were significantly upregulated. In humans, LL-37 is active against Candida (Lopez-Garcia et al. 2005). These findings demonstrate that commensal bacteria can potentially reduce colonization with fungi indirectly in the colon.
3.5 Interactions Between Pathogenic Bacteria and Candida Albicans
Dysbiosis facilitates increased colonization not only with C. albicans, but also with opportunistic bacterial pathogens (Bien et al. 2013). A broad spectrum of bacteria commonly found within the gut can cause disseminated infections in the host (Klastersky and Aoun 2004; Mootsikapun 2007). Given that mixed infections involving C. albicans and bacteria are not uncommon (Kim et al. 2013), and considering that dysbiosis favors both, growth of C. albicans and facultative pathogenic bacteria, in the gut simultaneously (Behnsen et al. 2014; Bien et al. 2013; Nerandzic et al. 2012), the type of interactions between these bacteria and the fungus might have clinical relevance. If fungal–bacterial interactions in the dysbiotic gut are antagonistic, the relative risk of infection might not be affected. In contrast, synergistic interactions might lead to an increased risk and warrant prophylactic treatment.
Acinetobacter baumanii is a Gram-negative bacterium recently emerging as a nosocomial opportunistic pathogen (Lee et al. 2007) difficult to treat due to development of resistances against a variety of antibiotics (Dijkshoorn et al. 2007). Interactions with C. albicans have been described as antagonistic. During in vitro coincubation, the bacterium can bind to C. albicans hyphae via the A. baumanii outer membrane protein A (OmpA) which results in hyphal apoptosis (Gaddy et al. 2009) (Fig. 3). Similarly, coinfection in a Caenorhabditis elegans nematode model resulted in reduced lethality mediated by inhibition of fungal filamentation (Peleg et al. 2008). However, it yet needs to be determined if interactions in the mammalian gut are likewise antagonistic.
The most commonly C. albicans-associated bacterial group are Gram-positives (Hermann et al. 1999). A variety of interactions have been found for Streptococcus (Bamford et al. 2015; Falsetta et al. 2014; Yu et al. 2018) and Staphylococcus species, especially Staphylococcus aureus (Harriott and Noverr 2010; Kong et al. 2016; Krause et al. 2015; Schlecht et al. 2015; Carlson 1983a, b). However, these interactions are mostly taking place outside of the gut. Enterococcus faecalis on the other hand shares a variety of niches with C. albicans (Hayashi et al. 2005; Horsley et al. 2013; Komiyama et al. 2016), and interactions in the gut are highly likely (Mason et al. 2012b). The high association between C. albicans and E. faecalis in clinical settings suggest synergistic relationships or at least neutral coexistence. Yet, the interactions described so far experimentally were antagonistic: In 2013, Cruz et al. (Cruz et al. 2013) observed that nematodes coinfected with E. faecalis and C. albicans survive longer compared to the respective single infections. This was partly due to inhibition of hyphae by a small peptide later identified and termed EntV (Graham et al. 2017) (Fig. 3). The peptide alone was shown to be able to protect mice from C. albicans infection in an oropharyngeal candidiasis (OPC) model by inhibiting hyphal formation and is also able to dissolve mature fungal biofilms. It is, however, unclear if EntV is expressed in the gut and whether interactions in this niche are also antagonistic.
Escherichia coli is a highly diverse bacterial species (Becker et al. 2014; Meador et al. 2014), and thus, it is not surprising that it also appears to be heterogenic in its behavior toward C. albicans. Recently, the E. coli strain MG1655 was found to secrete a fungicidal molecule that killed C. albicans during in vitro co-cultivation under magnesium-depleted conditions (Cabral et al. 2018). In contrast, enterohemorrhagic E. coli (EHEC) led to enhanced invasion and damage of enterocytes in vitro during coinfection, likely mediated by upregulation of hypha-associated genes like EFG1 and HWP1 (Yang et al. 2016) (Fig. 3). Such fungal–bacterial synergism was also observed in vivo during intraperitoneal coinfection of mice. While no mice died during single infection with either species, coinfection with both microorganisms led to drastically enhanced mortality (Klaerner et al. 1997). One strain of E. coli was furthermore shown to enhance fungal attachment to the bladder mucosa of rats and led to higher virulence during urinary tract infection (Levison and Pitsakis 1987). Of note, it was also demonstrated that supernatants of E. coli, similar to Pseudomonas aeruginosa, could inhibit biofilm formation in vitro (Holcombe et al. 2010; Bandara et al. 2013). Another in vitro study observed promotion of growth and proliferation of E. coli K12 in the presence of C. albicans. This effect was dependent on iron acquisition mechanisms from both microbial species. Deletion of either the fungal genes SEF1 or SFU1, transcription factors involved in iron uptake (Chen et al. 2011), or the bacterial ferric transporters FepD or FebG abolished growth promotion (Chenault and Earhart 1992). Interestingly, this growth promotion also occurred with E. coli mutants lacking their own siderophores. The authors suggested that a fungal siderophore-like molecule might support E. coli in taking up environmental iron. Even though the mechanism is poorly understood, it might be potentially important in the gut under iron limitation occurring in certain microniches. C. albicans could help stabilize the microbiota in its vicinity by yet unknown secreted factors depending on the environmental setting and interacting partner.
Metal acquisition is also important during Salmonella infection. Salmonella enterica serovar Typhimurium (S. typhimurium) colonization leads to severe inflammatory diarrhea (Hohmann 2001). In response to Salmonella infection, the host restricts iron as well as zinc and manganese, triggered by induction of the cytokine IL-22 (Godinez et al. 2008). IL-22 leads to secretion of different antimicrobial peptides such as RegIIIβ and RegIIIγ (Stelter et al. 2011), two C-type lectins able to kill Gram-negative and Gram-positive commensal bacteria but not S. typhimurium (Cash et al. 2006; Stelter et al. 2011). Two other downstream targets of IL-22 are lipocalin-2, which binds to the bacterial siderophore enterochelin preventing iron acquisition (Raffatellu et al. 2009), and calprotectin, a zinc and manganese chelator (Corbin et al. 2008; Sohnle et al. 1996). This leads to repression of commensal bacteria like E. coli and overgrowth of S. typhimurium (Behnsen et al. 2014). IL-22 has, however, no effect on C. albicans (Kagami et al. 2010). Direct antagonism between C. albicans and S. typhimurium has been also described in a nematode model where bacteria attached to hyphae and actively killed fungal cells over a type III secretion system (Kim and Mylonakis 2011) (Fig. 3).
In contrast to potentially harmful interplays with certain bacteria, C. albicans protects the host by antagonistic interactions with pathogens like Clostridium difficile (Markey et al. 2018). C. difficile is a Gram-positive obligate anaerobic spore-forming bacterium which can cause severe diarrhea and colitis, often with a lethal outcome (Burke and Lamont 2014). A major risk factor for infection is antibiotic treatment, which allows C. difficile to overcome colonization resistance provided by the commensal microbiota (Pepin et al. 2005). One last resort against reoccurring C. difficile infection is fecal microbiota transplantation (van Nood et al. 2013). It was previously shown that C. albicans can allow the obligate anaerobe C. difficile to grow under aerobic conditions during co-cultivation. This effect was independent of fungal biofilm formation which has been known to promote anaerobic growth (Fox et al. 2014); on the contrary, the bacterium secreted a molecule p-cresol which inhibited hyphal formation and led to conversion of hyphae back to yeast cells (van Leeuwen et al. 2016). This implies that C. albicans might provide a niche for the bacterium to thrive in. However, a recent in vivo study demonstrated that pre-colonization of mice with C. albicans protects the animals against a lethal challenge with C. difficile (Markey et al. 2018). This protective effect was not due to higher colonization resistance in the presence of the fungus, even though microbiota composition was significantly altered for some bacterial species in C. albicans-colonized mice, e.g., Akkermansia and Bifidobacterium species. Clostridium-derived toxin concentration was also unaltered in both groups. However, fungal colonization resulted in higher levels of pro-inflammatory IL-17A which attracts neutrophils (Nakagawa et al. 2016; Rubino et al. 2012) and promotes epithelial integrity via tight junctions (Lee et al. 2015) (Fig. 4).
Interactions between C. albicans and Clostridium difficile. Left: The obligate anaerobic bacterium C. difficile is able to grow in the presence of C. albicans in vitro under normoxic conditions. This effect is independent of fungal biofilm formation. In fact the bacterium secretes p-cresol, a small molecule that prevents fungal filamentation and therefore biofilm formation. Right: Intestinal colonization with C. albicans protects mice against C. difficile-induced killing. The fungus stimulates intestinal cells to secrete more IL-17A which boosts enterocyte regeneration, tight junctions’ expression and attracts protective immune cells. Illustration by M. Kapitan
While the exact mechanism by which C. albicans protects the host from pathogenic bacteria remains to be identified, these findings demonstrate that intestinal commensal fungi might protect the host against microbial threats under certain circumstances. Indeed Jiang et al. (Jiang et al. 2017) could show that mono-colonization by either C. albicans or S. cerevisiae after antibiotic treatment could protect mice from chemically induced colitis. Mice carrying commensal C. albicans did not show signs of colonic shortening or inflammation and in this regard behaved like mice with intact microbiota. Strikingly, intestinal mono-colonization with both fungi mediated protection against mortality upon pulmonary Influenza A virus infection. Responsible for this protection were mannans present in the fungal cell wall (Ruiz-Herrera et al. 2006) which led to the accumulation of protective immune cells. This work demonstrates that while antibiotic treatment favors fungal growth, this might not necessarily be detrimental, but that under distinct circumstances, fungi can also counteract infections by other microbes.
4 Conclusion
Historically, the view on C. albicans and other commensal fungi was dominated by their ability to cause opportunistic infections and their role during food digestion. Comparatively little is, however, known on the role of fungi as part of the microbiota in the gut. While it is obvious that bacteria mediate colonization resistance against C. albicans, its interactions with distinct bacterial species and strains appear to range from antagonistic to synergistic and likely depend on the environment in which these interactions take place. Under discrete circumstances, C. albicans, which occurs in the gut microbiota of the majority of humans and appears to be adapted to the human host, may even have beneficial effects on the host. With the growing understanding about the role that the microbiota has in different aspect of human life and well-being, it becomes increasingly clear that we also need a better understanding of fungal functions inside the bacterial community. For this, interactions between fungal and bacterial species need to be investigated in more detail. Identifying mechanisms of interactions and the environmental factors that affect these interactions will provide a basis for personalized risk assessment in patients with dysbiosis. And even though fungi are mainly considered relevant as pathogens, they may provide benefits to the host that yet remain to be discovered.
References
Adlercreutz EH, Wingren CJ, Vincente RP, Merlo J, Agardh D (2015) Perinatal risk factors increase the risk of being affected by both type 1 diabetes and coeliac disease. Acta Paediatr 104(2):178–184. https://doi.org/10.1111/apa.12836
Anderson HW (1917) Yeast-like fungi of the human intestinal tract. J Infect Dis 21(4):341–354. https://doi.org/10.1093/infdis/21.4.341
Antoni L, Nuding S, Wehkamp J, Stange EF (2014) Intestinal barrier in inflammatory bowel disease. World J Gastroenterol 20(5):1165–1179. https://doi.org/10.3748/wjg.v20.i5.1165
Aroutcheva A, Gariti D, Simon M, Shott S, Faro J, Simoes JA, Gurguis A, Faro S (2001) Defense factors of vaginal lactobacilli. Am J Obstet Gynecol 185(2):375–379. https://doi.org/10.1067/mob.2001.115867
Ayabe T, Satchell DP, Wilson CL, Parks WC, Selsted ME, Ouellette AJ (2000) Secretion of microbicidal alpha-defensins by intestinal Paneth cells in response to bacteria. Nat Immunol 1(2):113–118. https://doi.org/10.1038/77783
Azad MB, Konya T, Guttman DS, Field CJ, Sears MR, HayGlass KT, Mandhane PJ, Turvey SE, Subbarao P, Becker AB, Scott JA, Kozyrskyj AL, Investigators CS (2015) Infant gut microbiota and food sensitization: associations in the first year of life. Clin Exp Allergy 45(3):632–643. https://doi.org/10.1111/cea.12487
Azevedo MM, Teixeira-Santos R, Silva AP, Cruz L, Ricardo E, Pina-Vaz C, Rodrigues AG (2015) The effect of antibacterial and non-antibacterial compounds alone or associated with antifugals upon fungi. Front Microbiol 6:669. https://doi.org/10.3389/fmicb.2015.00669
Baharlou R, Ahmadi-Vasmehjani A, Davami MH, Faraji F, Atashzar MR, Karimipour F, Sadeghi A, Asadi MA, Khoubyari M (2016) Elevated Levels of T-helper 17-associated cytokines in diabetes type I patients: indicators for following the course of disease. Immunol Invest 45(7):641–651. https://doi.org/10.1080/08820139.2016.1197243
Bamford CV, Nobbs AH, Barbour ME, Lamont RJ, Jenkinson HF (2015) Functional regions of Candida albicans hyphal cell wall protein Als3 that determine interaction with the oral bacterium Streptococcus gordonii. Microbiology 161(Pt 1):18–29. https://doi.org/10.1099/mic.0.083378-0
Bandara HM, Cheung BP, Watt RM, Jin LJ, Samaranayake LP (2013) Secretory products of Escherichia coli biofilm modulate Candida biofilm formation and hyphal development. J Investig Clin Dent 4(3):186–199. https://doi.org/10.1111/jicd.12048
Becker HM, Apladas A, Scharl M, Fried M, Rogler G (2014) Probiotic Escherichia coli Nissle 1917 and commensal E. coli K12 differentially affect the inflammasome in intestinal epithelial cells. Digestion 89(2):110–118. https://doi.org/10.1159/000357521
Behnsen J, Jellbauer S, Wong CP, Edwards RA, George MD, Ouyang W, Raffatellu M (2014) The cytokine IL-22 promotes pathogen colonization by suppressing related commensal bacteria. Immunity 40(2):262–273. https://doi.org/10.1016/j.immuni.2014.01.003
Berer K, Gerdes LA, Cekanaviciute E, Jia X, Xiao L, Xia Z, Liu C, Klotz L, Stauffer U, Baranzini SE, Kumpfel T, Hohlfeld R, Krishnamoorthy G, Wekerle H (2017) Gut microbiota from multiple sclerosis patients enables spontaneous autoimmune encephalomyelitis in mice. Proc Natl Acad Sci U S A 114(40):10719–10724. https://doi.org/10.1073/pnas.1711233114
Berman J, Sudbery PE (2002) Candida albicans: a molecular revolution built on lessons from budding yeast. Nat Rev Genet 3(12):918–930
Bhargava P, Mowry EM (2014) Gut microbiome and multiple sclerosis. Curr Neurol Neurosci Rep 14(10):492. https://doi.org/10.1007/s11910-014-0492-2
Bhinder G, Stahl M, Sham HP, Crowley SM, Morampudi V, Dalwadi U, Ma C, Jacobson K, Vallance BA (2014) Intestinal epithelium-specific MyD88 signaling impacts host susceptibility to infectious colitis by promoting protective goblet cell and antimicrobial responses. Infect Immun 82(9):3753–3763. https://doi.org/10.1128/IAI.02045-14
Bianchi CM, Bianchi HA, Tadano T, Paula CR, Hoffmann-Santos HD, Leite DP Jr, Hahn RC (2016) Factors related to oral candidiasis in elderly users and non-users of removable dental prostheses. Rev Inst Med Trop Sao Paulo 58:17. https://doi.org/10.1590/s1678-9946201658017
Bien J, Palagani V, Bozko P (2013) The intestinal microbiota dysbiosis and Clostridium difficile infection: is there a relationship with inflammatory bowel disease? Therap Adv Gastroenterol 6(1):53–68. https://doi.org/10.1177/1756283x12454590
Bischoff SC, Barbara G, Buurman W, Ockhuizen T, Schulzke JD, Serino M, Tilg H, Watson A, Wells JM (2014) Intestinal permeability—a new target for disease prevention and therapy. BMC Gastroenterol 14:189. https://doi.org/10.1186/s12876-014-0189-7
Black M, Bhattacharya S, Philip S, Norman JE, McLernon DJ (2015) Planned cesarean delivery at term and adverse outcomes in childhood health. JAMA 314(21):2271–2279. https://doi.org/10.1001/jama.2015.16176
Bliss JM, Basavegowda KP, Watson WJ, Sheikh AU, Ryan RM (2008) Vertical and horizontal transmission of Candida albicans in very low birth weight infants using DNA fingerprinting techniques. J Pediatr 27(3):231–235. https://doi.org/10.1097/INF.0b013e31815bb69d
Böhm L, Torsin S, Tint SH, Eckstein MT, Ludwig T, Perez JC (2017) The yeast form of the fungus Candida albicans promotes persistence in the gut of gnotobiotic mice. PLoS Pathog 13(10):e1006699. https://doi.org/10.1371/journal.ppat.1006699
Bouza E, Burillo A, Munoz P, Guinea J, Marin M, Rodriguez-Creixems M (2013) Mixed bloodstream infections involving bacteria and Candida spp. J Antimicrob Chemother 68(8):1881–1888. https://doi.org/10.1093/jac/dkt099
Brandl K, Plitas G, Mihu CN, Ubeda C, Jia T, Fleisher M, Schnabl B, DeMatteo RP, Pamer EG (2008) Vancomycin-resistant enterococci exploit antibiotic-induced innate immune deficits. Nature 455(7214):804–807. https://doi.org/10.1038/nature07250
Brown AJ, Brown GD, Netea MG, Gow NA (2014) Metabolism impacts upon Candida immunogenicity and pathogenicity at multiple levels. Trends Microbiol 22(11):614–622. https://doi.org/10.1016/j.tim.2014.07.001
Brown GD, Denning DW, Gow NA, Levitz SM, Netea MG, White TC (2012) Hidden killers: human fungal infections. Sci Transl Med 4 (165):165rv113. https://doi.org/10.1126/scitranslmed.3004404
Buffie CG, Pamer EG (2013) Microbiota-mediated colonization resistance against intestinal pathogens. Nat Rev Immunol 13(11):790–801. https://doi.org/10.1038/nri3535
Burger-van Paassen N, Vincent A, Puiman PJ, van der Sluis M, Bouma J, Boehm G, van Goudoever JB, van Seuningen I, Renes IB (2009) The regulation of intestinal mucin MUC2 expression by short-chain fatty acids: implications for epithelial protection. Biochem J 420(2):211–219. https://doi.org/10.1042/BJ20082222
Burke KE, Lamont JT (2014) Clostridium difficile infection: a worldwide disease. Gut Liver 8(1):1–6. https://doi.org/10.5009/gnl.2014.8.1.1
Cabral DJ, Penumutchu S, Norris C, Morones-Ramirez JR, Belenky P (2018) Microbial competition between Escherichia coli and Candida albicans reveals a soluble fungicidal factor. Microb Cell 5(5):249–255. https://doi.org/10.15698/mic2018.05.631
Calderone RA, Clancy CJ (2012) Candida and Candidiasis, Second Edition. MBio. https://doi.org/10.1128/9781555817176
Carlson E (1983a) Effect of strain of Staphylococcus aureus on synergism with Candida albicans resulting in mouse mortality and morbidity. Infect Immun 42(1):285–292
Carlson E (1983b) Enhancement by Candida albicans of Staphylococcus aureus, Serratia marcescens, and Streptococcus faecalis in the establishment of infection in mice. Infect Immun 39(1):193–197
Carruba G, Pontieri E, De Bernardis F, Martino P, Cassone A (1991) DNA fingerprinting and electrophoretic karyotype of environmental and clinical isolates of Candida parapsilosis. J Clin Microbiol 29(5):916–922
Cash HL, Whitham CV, Behrendt CL, Hooper LV (2006) Symbiotic bacteria direct expression of an intestinal bactericidal lectin. Science 313(5790):1126–1130. https://doi.org/10.1126/science.1127119
Chambers PJ, Pretorius IS (2010) Fermenting knowledge: the history of winemaking, science and yeast research. EMBO Rep 11(12):914–920. https://doi.org/10.1038/embor.2010.179
Chaudhuri A (2016) Cell biology by the numbers. Yale J Biol Med 89(3):425–426
Chen C, Pande K, French SD, Tuch BB, Noble SM (2011) An iron homeostasis regulatory circuit with reciprocal roles in Candida albicans commensalism and pathogenesis. Cell Host Microbe 10(2):118–135. https://doi.org/10.1016/j.chom.2011.07.005
Chen J, Wright K, Davis JM, Jeraldo P, Marietta EV, Murray J, Nelson H, Matteson EL, Taneja V (2016) An expansion of rare lineage intestinal microbes characterizes rheumatoid arthritis. Genome Med 8(1):43. https://doi.org/10.1186/s13073-016-0299-7
Chenault SS, Earhart CF (1992) Identification of hydrophobic proteins FepD and FepG of the Escherichia coli ferrienterobactin permease. J Gen Microbiol 138(10):2167–2171. https://doi.org/10.1099/00221287-138-10-2167
Chevalier C, Stojanovic O, Colin DJ, Suarez-Zamorano N, Tarallo V, Veyrat-Durebex C, Rigo D, Fabbiano S, Stevanovic A, Hagemann S, Montet X, Seimbille Y, Zamboni N, Hapfelmeier S, Trajkovski M (2015) Gut microbiota orchestrates energy homeostasis during cold. Cell 163(6):1360–1374. https://doi.org/10.1016/j.cell.2015.11.004
Collado MC, Rautava S, Aakko J, Isolauri E, Salminen S (2016) Human gut colonisation may be initiated in utero by distinct microbial communities in the placenta and amniotic fluid. Sci Rep 6:23129. https://doi.org/10.1038/srep23129
Corbin BD, Seeley EH, Raab A, Feldmann J, Miller MR, Torres VJ, Anderson KL, Dattilo BM, Dunman PM, Gerads R, Caprioli RM, Nacken W, Chazin WJ, Skaar EP (2008) Metal chelation and inhibition of bacterial growth in tissue abscesses. Science 319(5865):962–965. https://doi.org/10.1126/science.1152449
Corthier G, Dubos F, Ducluzeau R (1986) Prevention of Clostridium difficile induced mortality in gnotobiotic mice by Saccharomyces boulardii. Can J Microbiol 32(11):894–896
Cruz MR, Graham CE, Gagliano BC, Lorenz MC, Garsin DA (2013) Enterococcus faecalis Inhibits Hyphal Morphogenesis and Virulence of Candida albicans. Infect Immun 81(1):189–200. https://doi.org/10.1128/IAI.00914-12
d’Arminio Monforte A, Gianotti N, Cozzi-Lepri A, Pinnetti C, Andreoni M, di Perri G, Galli M, Poli A, Costantini A, Orofino G, Maggiolo F, Mazzarello G, Celesia BM, Luciani F, Lazzarin A, Sighinolfi L, Rizzardini G, Bonfanti P, Perno CF, Antinori A, Cohort IF (2014) Durability of lopinavir/ritonavir monotherapy in individuals with viral load </=50 copies/ml in an observational setting. Antivir Ther 19(3):319–324. https://doi.org/10.3851/IMP2687
David LA, Maurice CF, Carmody RN, Gootenberg DB, Button JE, Wolfe BE, Ling AV, Devlin AS, Varma Y, Fischbach MA, Biddinger SB, Dutton RJ, Turnbaugh PJ (2014) Diet rapidly and reproducibly alters the human gut microbiome. Nature 505(7484):559–563. https://doi.org/10.1038/nature12820
Davis-Richardson AG, Ardissone AN, Dias R, Simell V, Leonard MT, Kemppainen KM, Drew JC, Schatz D, Atkinson MA, Kolaczkowski B, Ilonen J, Knip M, Toppari J, Nurminen N, Hyoty H, Veijola R, Simell T, Mykkanen J, Simell O, Triplett EW (2014) Bacteroides dorei dominates gut microbiome prior to autoimmunity in Finnish children at high risk for type 1 diabetes. Front Microbiol 5:678. https://doi.org/10.3389/fmicb.2014.00678
De Filippis F, Pellegrini N, Vannini L, Jeffery IB, La Storia A, Laghi L, Serrazanetti DI, Di Cagno R, Ferrocino I, Lazzi C, Turroni S, Cocolin L, Brigidi P, Neviani E, Gobbetti M, O’Toole PW, Ercolini D (2016) High-level adherence to a Mediterranean diet beneficially impacts the gut microbiota and associated metabolome. Gut 65(11):1812–1821. https://doi.org/10.1136/gutjnl-2015-309957
de Goffau MC, Fuentes S, van den Bogert B, Honkanen H, de Vos WM, Welling GW, Hyoty H, Harmsen HJ (2014) Aberrant gut microbiota composition at the onset of type 1 diabetes in young children. Diabetologia 57(8):1569–1577. https://doi.org/10.1007/s00125-014-3274-0
De Palma G, Lynch MD, Lu J, Dang VT, Deng Y, Jury J, Umeh G, Miranda PM, Pigrau Pastor M, Sidani S, Pinto-Sanchez MI, Philip V, McLean PG, Hagelsieb MG, Surette MG, Bergonzelli GE, Verdu EF, Britz-McKibbin P, Neufeld JD, Collins SM, Bercik P (2017) Transplantation of fecal microbiota from patients with irritable bowel syndrome alters gut function and behavior in recipient mice. Sci Transl Med 9 (379). https://doi.org/10.1126/scitranslmed.aaf6397
de Sablet T, Chassard C, Bernalier-Donadille A, Vareille M, Gobert AP, Martin C (2009) Human microbiota-secreted factors inhibit shiga toxin synthesis by enterohemorrhagic Escherichia coli O157:H7. Infect Immun 77(2):783–790. https://doi.org/10.1128/IAI.01048-08
De Vuyst L, Leroy F (2007) Bacteriocins from lactic acid bacteria: production, purification, and food applications. J Mol Microbiol Biotechnol 13(4):194–199. https://doi.org/10.1159/000104752
Derrien M, Vaughan EE, Plugge CM, de Vos WM (2004) Akkermansia muciniphila gen. nov., sp. nov., a human intestinal mucin-degrading bacterium. Int J Syst Evol Microbiol 54 (Pt 5):1469–1476. https://doi.org/10.1099/ijs.0.02873-0
Dijkshoorn L, Nemec A, Seifert H (2007) An increasing threat in hospitals: multidrug-resistant Acinetobacter baumannii. Nat Rev Microbiol 5(12):939–951. https://doi.org/10.1038/nrmicro1789
Dominguez-Bello MG, Costello EK, Contreras M, Magris M, Hidalgo G, Fierer N, Knight R (2010) Delivery mode shapes the acquisition and structure of the initial microbiota across multiple body habitats in newborns. Proc Natl Acad Sci U S A 107(26):11971–11975. https://doi.org/10.1073/pnas.1002601107
Eggimann P, Que YA, Revelly JP, Pagani JL (2015) Preventing invasive Candida infections. Where could we do better? J Hosp Infect 89(4):302–308. https://doi.org/10.1016/j.jhin.2014.11.006
Ene IV, Brunke S, Brown AJ, Hube B (2014) Metabolism in fungal pathogenesis. Cold Spring Harb Perspect Med 4(12):a019695. https://doi.org/10.1101/cshperspect.a019695
Erb Downward JR, Falkowski NR, Mason KL, Muraglia R, Huffnagle GB (2013) Modulation of post-antibiotic bacterial community reassembly and host response by Candida albicans. Sci Rep 3:2191. https://doi.org/10.1038/srep02191
Evans CC, LePard KJ, Kwak JW, Stancukas MC, Laskowski S, Dougherty J, Moulton L, Glawe A, Wang Y, Leone V, Antonopoulos DA, Smith D, Chang EB, Ciancio MJ (2014) Exercise prevents weight gain and alters the gut microbiota in a mouse model of high fat diet-induced obesity. PLoS ONE 9(3):e92193. https://doi.org/10.1371/journal.pone.0092193
Fagarasan S, Kawamoto S, Kanagawa O, Suzuki K (2010) Adaptive immune regulation in the gut: T cell-dependent and T cell-independent IgA synthesis. Annu Rev Immunol 28:243–273. https://doi.org/10.1146/annurev-immunol-030409-101314
Falsetta ML, Klein MI, Colonne PM, Scott-Anne K, Gregoire S, Pai CH, Gonzalez-Begne M, Watson G, Krysan DJ, Bowen WH, Koo H (2014) Symbiotic relationship between Streptococcus mutans and Candida albicans synergizes virulence of plaque biofilms in vivo. Infect Immun 82(5):1968–1981. https://doi.org/10.1128/iai.00087-14
Fan D, Coughlin LA, Neubauer MM, Kim J, Kim MS, Zhan X, Simms-Waldrip TR, Xie Y, Hooper LV, Koh AY (2015) Activation of HIF-1alpha and LL-37 by commensal bacteria inhibits Candida albicans colonization. Nat Med 21(7):808–814. https://doi.org/10.1038/nm.3871
Farache J, Koren I, Milo I, Gurevich I, Kim KW, Zigmond E, Furtado GC, Lira SA, Shakhar G (2013) Luminal bacteria recruit CD103+ dendritic cells into the intestinal epithelium to sample bacterial antigens for presentation. Immunity 38(3):581–595. https://doi.org/10.1016/j.immuni.2013.01.009
Fernandez L, Langa S, Martin V, Jimenez E, Martin R, Rodriguez JM (2013) The microbiota of human milk in healthy women. Cell Mol Biol (Noisy-le-grand) 59 (1):31–42
Fischbach MA, Sonnenburg JL (2011) Eating for two: how metabolism establishes interspecies interactions in the gut. Cell Host Microbe 10(4):336–347. https://doi.org/10.1016/j.chom.2011.10.002
Flint HJ, Scott KP, Duncan SH, Louis P, Forano E (2012) Microbial degradation of complex carbohydrates in the gut. Gut Microbes 3(4):289–306. https://doi.org/10.4161/gmic.19897
Fores JP, Crisostomo LG, Orii NM, Santos AS, Fukui RT, Matioli SR, de Moraes Vasconcelos D, Silva M (2018) Th17 pathway in recent-onset autoimmune diabetes. Cell Immunol 324:8–13. https://doi.org/10.1016/j.cellimm.2017.11.005
Fox EP, Cowley ES, Nobile CJ, Hartooni N, Newman DK, Johnson AD (2014) Anaerobic bacteria grow within Candida albicans biofilms and induce biofilm formation in suspension cultures. Curr Biol 24(20):2411–2416. https://doi.org/10.1016/j.cub.2014.08.057
Frantz AL, Rogier EW, Weber CR, Shen L, Cohen DA, Fenton LA, Bruno ME, Kaetzel CS (2012) Targeted deletion of MyD88 in intestinal epithelial cells results in compromised antibacterial immunity associated with downregulation of polymeric immunoglobulin receptor, mucin-2, and antibacterial peptides. Mucosal Immunol 5(5):501–512. https://doi.org/10.1038/mi.2012.23
Fukata M, Arditi M (2013) The role of pattern recognition receptors in intestinal inflammation. Mucosal Immunol 6(3):451–463. https://doi.org/10.1038/mi.2013.13
Fukui H (2016) Increased intestinal permeability and decreased barrier function: does it really influence the risk of inflammation? Inflamm Intest Dis 1(3):135–145
Gabaldon T, Carrete L (2016) The birth of a deadly yeast: tracing the evolutionary emergence of virulence traits in Candida glabrata. FEMS Yeast Res 16 (2):fov110. https://doi.org/10.1093/femsyr/fov110
Gaboriau-Routhiau V, Rakotobe S, Lecuyer E, Mulder I, Lan A, Bridonneau C, Rochet V, Pisi A, De Paepe M, Brandi G, Eberl G, Snel J, Kelly D, Cerf-Bensussan N (2009) The key role of segmented filamentous bacteria in the coordinated maturation of gut helper T cell responses. Immunity 31(4):677–689. https://doi.org/10.1016/j.immuni.2009.08.020
Gadanho M, Sampaio JP (2005) Occurrence and diversity of yeasts in the mid-atlantic ridge hydrothermal fields near the Azores Archipelago. Microb Ecol 50(3):408–417. https://doi.org/10.1007/s00248-005-0195-y
Gaddy JA, Tomaras AP, Actis LA (2009) The Acinetobacter baumannii 19606 OmpA protein plays a role in biofilm formation on abiotic surfaces and in the interaction of this pathogen with eukaryotic cells. Infect Immun 77(8):3150–3160. https://doi.org/10.1128/iai.00096-09
Gagnon M, Kheadr EE, Le Blay G, Fliss I (2004) In vitro inhibition of Escherichia coli O157:H7 by bifidobacterial strains of human origin. Int J Food Microbiol 92(1):69–78. https://doi.org/10.1016/j.ijfoodmicro.2003.07.010
George JA, Bashir G, Qureshi MM, Mohamed YA, Azzi J, Al-Ramadi BK, Fernandez-Cabezudo MJ (2016) Cholinergic stimulation prevents the development of autoimmune diabetes: evidence for the modulation of Th17 effector cells via an IFNgamma-dependent mechanism. Front Immunol 7:419. https://doi.org/10.3389/fimmu.2016.00419
Geuking MB, Cahenzli J, Lawson MA, Ng DC, Slack E, Hapfelmeier S, McCoy KD, Macpherson AJ (2011) Intestinal bacterial colonization induces mutualistic regulatory T cell responses. Immunity 34(5):794–806. https://doi.org/10.1016/j.immuni.2011.03.021
Giongo A, Gano KA, Crabb DB, Mukherjee N, Novelo LL, Casella G, Drew JC, Ilonen J, Knip M, Hyoty H, Veijola R, Simell T, Simell O, Neu J, Wasserfall CH, Schatz D, Atkinson MA, Triplett EW (2011) Toward defining the autoimmune microbiome for type 1 diabetes. ISME J 5(1):82–91. https://doi.org/10.1038/ismej.2010.92
Gkolfakis P, Dimitriadis G, Triantafyllou K (2015) Gut microbiota and non-alcoholic fatty liver disease. Hepatobiliary Pancreat Dis Int 14(6):572–581
Goddard MR (2008) Quantifying the complexities of Saccharomyces cerevisiae’s ecosystem engineering via fermentation. Ecology 89(8):2077–2082
Godinez I, Haneda T, Raffatellu M, George MD, Paixao TA, Rolan HG, Santos RL, Dandekar S, Tsolis RM, Baumler AJ (2008) T cells help to amplify inflammatory responses induced by Salmonella enterica serotype Typhimurium in the intestinal mucosa. Infect Immun 76(5):2008–2017. https://doi.org/10.1128/iai.01691-07
Goulet O (2015) Potential role of the intestinal microbiota in programming health and disease. Nutr Rev 73(Suppl 1):32–40. https://doi.org/10.1093/nutrit/nuv039
Graham CE, Cruz MR, Garsin DA, Lorenz MC (2017) Enterococcus faecalis bacteriocin EntV inhibits hyphal morphogenesis, biofilm formation, and virulence of Candida albicans. Proc Natl Acad Sci U S A 114(17):4507–4512. https://doi.org/10.1073/pnas.1620432114
Haghikia A, Jorg S, Duscha A, Berg J, Manzel A, Waschbisch A, Hammer A, Lee DH, May C, Wilck N, Balogh A, Ostermann AI, Schebb NH, Akkad DA, Grohme DA, Kleinewietfeld M, Kempa S, Thone J, Demir S, Muller DN, Gold R, Linker RA (2015) Dietary fatty acids directly impact central nervous system autoimmunity via the small intestine. Immunity 43(4):817–829. https://doi.org/10.1016/j.immuni.2015.09.007
Hamada H, Hiroi T, Nishiyama Y, Takahashi H, Masunaga Y, Hachimura S, Kaminogawa S, Takahashi-Iwanaga H, Iwanaga T, Kiyono H, Yamamoto H, Ishikawa H (2002) Identification of multiple isolated lymphoid follicles on the antimesenteric wall of the mouse small intestine. J Immunol 168(1):57–64
Harriott MM, Noverr MC (2010) Ability of Candida albicans mutants to induce Staphylococcus aureus vancomycin resistance during polymicrobial biofilm formation. Antimicrob Agents Chemother 54(9):3746–3755. https://doi.org/10.1128/AAC.00573-10
Hayashi H, Takahashi R, Nishi T, Sakamoto M, Benno Y (2005) Molecular analysis of jejunal, ileal, caecal and recto-sigmoidal human colonic microbiota using 16S rRNA gene libraries and terminal restriction fragment length polymorphism. J Med Microbiol 54(Pt 11):1093–1101. https://doi.org/10.1099/jmm.0.45935-0
Hermann C, Hermann J, Munzel U, Ruchel R (1999) Bacterial flora accompanying Candida yeasts in clinical specimens. Mycoses 42(11–12):619–627
Herwald SE, Kumamoto CA (2014) Candida albicans niche specialization: features that distinguish biofilm cells from commensal cells. Curr Fungal Infect Rep 8(2):179–184. https://doi.org/10.1007/s12281-014-0178-x
Hevia A, Milani C, Lopez P, Cuervo A, Arboleya S, Duranti S, Turroni F, Gonzalez S, Suarez A, Gueimonde M, Ventura M, Sanchez B, Margolles A (2014) Intestinal dysbiosis associated with systemic lupus erythematosus. MBio 5(5):e01548–01514. https://doi.org/10.1128/mBio.01548-14
Hickman MA, Zeng G, Forche A, Hirakawa MP, Abbey D, Harrison BD, Wang YM, Su CH, Bennett RJ, Wang Y, Berman J (2013) The ‘obligate diploid’ Candida albicans forms mating-competent haploids. Nature 494(7435):55–59. https://doi.org/10.1038/nature11865
Hoffmann C, Dollive S, Grunberg S, Chen J, Li H, Wu GD, Lewis JD, Bushman FD (2013) Archaea and fungi of the human gut microbiome: correlations with diet and bacterial residents. PLoS ONE 8(6):e66019. https://doi.org/10.1371/journal.pone.0066019
Hohmann EL (2001) Nontyphoidal salmonellosis. Clin Infect Dis 32(2):263–269. https://doi.org/10.1086/318457
Holcombe LJ, McAlester G, Munro CA, Enjalbert B, Brown AJP, Gow NAR, Ding C, Butler G, O’Gara F, Morrissey JP (2010) Pseudomonas aeruginosa secreted factors impair biofilm development in Candida albicans. Microbiology 156(Pt 5):1476–1486. https://doi.org/10.1099/mic.0.037549-0
Hollister EB, Riehle K, Luna RA, Weidler EM, Rubio-Gonzales M, Mistretta TA, Raza S, Doddapaneni HV, Metcalf GA, Muzny DM, Gibbs RA, Petrosino JF, Shulman RJ, Versalovic J (2015) Structure and function of the healthy pre-adolescent pediatric gut microbiome. Microbiome 3:36. https://doi.org/10.1186/s40168-015-0101-x
Hopkins MJ, Macfarlane GT (2002) Changes in predominant bacterial populations in human faeces with age and with Clostridium difficile infection. J Med Microbiol 51(5):448–454. https://doi.org/10.1099/0022-1317-51-5-448
Horsley H, Malone-Lee J, Holland D, Tuz M, Hibbert A, Kelsey M, Kupelian A, Rohn JL (2013) Enterococcus faecalis subverts and invades the host urothelium in patients with chronic urinary tract infection. PLoS ONE 8(12):e83637. https://doi.org/10.1371/journal.pone.0083637
Ianiro G, Tilg H, Gasbarrini A (2016) Antibiotics as deep modulators of gut microbiota: between good and evil. Gut 65(11):1906–1915. https://doi.org/10.1136/gutjnl-2016-312297
Ivanov II, Atarashi K, Manel N, Brodie EL, Shima T, Karaoz U, Wei D, Goldfarb KC, Santee CA, Lynch SV, Tanoue T, Imaoka A, Itoh K, Takeda K, Umesaki Y, Honda K, Littman DR (2009) Induction of intestinal Th17 cells by segmented filamentous bacteria. Cell 139(3):485–498. https://doi.org/10.1016/j.cell.2009.09.033
Ivanov II, Frutos Rde L, Manel N, Yoshinaga K, Rifkin DB, Sartor RB, Finlay BB, Littman DR (2008) Specific microbiota direct the differentiation of IL-17-producing T-helper cells in the mucosa of the small intestine. Cell Host Microbe 4(4):337–349. https://doi.org/10.1016/j.chom.2008.09.009
Jiang TT, Shao TY, Ang WXG, Kinder JM, Turner LH, Pham G, Whitt J, Alenghat T, Way SS (2017) Commensal fungi recapitulate the protective benefits of intestinal bacteria. Cell Host Microbe 22 (6):809–816.e804. https://doi.org/10.1016/j.chom.2017.10.013
Juyal D, Sharma M, Pal S, Rathaur VK, Sharma N (2013) Emergence of non-albicans Candida species in neonatal candidemia. N Am J Med Sci 5(9):541–545. https://doi.org/10.4103/1947-2714.118919
Kagami S, Rizzo HL, Kurtz SE, Miller LS, Blauvelt A (2010) IL-23 and IL-17A, but not IL-12 and IL-22, are required for optimal skin host defense against Candida albicans. J Immunol 185(9):5453–5462. https://doi.org/10.4049/jimmunol.1001153
Kamada N, Kim YG, Sham HP, Vallance BA, Puente JL, Martens EC, Nunez G (2012) Regulated virulence controls the ability of a pathogen to compete with the gut microbiota. Science 336(6086):1325–1329. https://doi.org/10.1126/science.1222195
Kamada N, Seo SU, Chen GY, Nunez G (2013) Role of the gut microbiota in immunity and inflammatory disease. Nat Rev Immunol 13(5):321–335. https://doi.org/10.1038/nri3430
Kankainen M, Paulin L, Tynkkynen S, von Ossowski I, Reunanen J, Partanen P, Satokari R, Vesterlund S, Hendrickx AP, Lebeer S, De Keersmaecker SC, Vanderleyden J, Hamalainen T, Laukkanen S, Salovuori N, Ritari J, Alatalo E, Korpela R, Mattila-Sandholm T, Lassig A, Hatakka K, Kinnunen KT, Karjalainen H, Saxelin M, Laakso K, Surakka A, Palva A, Salusjarvi T, Auvinen P, de Vos WM (2009) Comparative genomic analysis of Lactobacillus rhamnosus GG reveals pili containing a human- mucus binding protein. Proc Natl Acad Sci U S A 106(40):17193–17198. https://doi.org/10.1073/pnas.0908876106
Katagiri H, Fukui K, Nakamura K, Tanaka A (2018) Systemic hematogenous dissemination of mouse oral candidiasis is induced by oral mucositis. Odontology. https://doi.org/10.1007/s10266-018-0366-1
Kelesidis T, Pothoulakis C (2012) Efficacy and safety of the probiotic Saccharomyces boulardii for the prevention and therapy of gastrointestinal disorders. Therap Adv Gastroenterol 5(2):111–125. https://doi.org/10.1177/1756283x11428502
Khutoryanskiy VV (2015) Supramolecular materials: Longer and safer gastric residence. Nat Mater 14(10):963–964. https://doi.org/10.1038/nmat4432
Kim SH, Yoon YK, Kim MJ, Sohn JW (2013) Risk factors for and clinical implications of mixed Candida/bacterial bloodstream infections. Clin Microbiol Infect 19(1):62–68. https://doi.org/10.1111/j.1469-0691.2012.03906.x
Kim Y, Mylonakis E (2011) Killing of Candida albicans filaments by Salmonella enterica serovar Typhimurium is mediated by sopB effectors, parts of a type III secretion system. Eukaryot Cell 10(6):782–790. https://doi.org/10.1128/EC.00014-11
Klaerner HG, Uknis ME, Acton RD, Dahlberg PS, Carlone-Jambor C, Dunn DL (1997) Candida albicans and Escherichia coli are synergistic pathogens during experimental microbial peritonitis. J Surg Res 70(2):161–165. https://doi.org/10.1006/jsre.1997.5110
Klastersky J, Aoun M (2004) Opportunistic infections in patients with cancer. Ann Oncol 15 Suppl 4:iv329–335. https://doi.org/10.1093/annonc/mdh947
Klotz SA, Chasin BS, Powell B, Gaur NK, Lipke PN (2007) Polymicrobial bloodstream infections involving Candida species: analysis of patients and review of the literature. Diagn Microbiol Infect Dis 59(4):401–406. https://doi.org/10.1016/j.diagmicrobio.2007.07.001
Knight DJ, Girling KJ (2003) Gut flora in health and disease. Lancet 361 (9371):1831
Koc AN, Silici S, Mutlu-Sariguzel F, Sagdic O (2007) Antifungal activity of propolis in four different fruit juices. Food Technol Biotechnol 45(1):57–61
Komiyama EY, Lepesqueur LS, Yassuda CG, Samaranayake LP, Parahitiyawa NB, Balducci I, Koga-Ito CY (2016) Enterococcus species in the oral cavity: prevalence, virulence factors and antimicrobial susceptibility. PLoS ONE 11(9):e0163001. https://doi.org/10.1371/journal.pone.0163001
Kong EF, Tsui C, Kucharikova S, Andes D, Van Dijck P, Jabra-Rizk MA (2016) Commensal protection of Staphylococcus aureus against antimicrobials by Candida albicans Biofilm Matrix. MBio 7(5). https://doi.org/10.1128/mbio.01365-16
Krause J, Geginat G, Tammer I (2015) Prostaglandin E2 from Candida albicans stimulates the growth of Staphylococcus aureus in mixed biofilms. PLoS ONE 10(8):e0135404. https://doi.org/10.1371/journal.pone.0135404
Krebs CF, Paust HJ, Krohn S, Koyro T, Brix SR, Riedel JH, Bartsch P, Wiech T, Meyer-Schwesinger C, Huang J, Fischer N, Busch P, Mittrucker HW, Steinhoff U, Stockinger B, Perez LG, Wenzel UO, Janneck M, Steinmetz OM, Gagliani N, Stahl RAK, Huber S, Turner JE, Panzer U (2016) Autoimmune renal disease is exacerbated by S1P-receptor-1-dependent intestinal Th17 cell migration to the kidney. Immunity 45(5):1078–1092. https://doi.org/10.1016/j.immuni.2016.10.020
Kuhle S, Tong OS, Woolcott CG (2015) Association between caesarean section and childhood obesity: a systematic review and meta-analysis. Obes Rev 16(4):295–303. https://doi.org/10.1111/obr.12267
Kuhn KA, Manieri NA, Liu TC, Stappenbeck TS (2014) IL-6 stimulates intestinal epithelial proliferation and repair after injury. PLoS ONE 9(12):e114195. https://doi.org/10.1371/journal.pone.0114195
Kundu P, Blacher E, Elinav E, Pettersson S (2017) Our gut microbiome: the evolving inner self. Cell 171(7):1481–1493. https://doi.org/10.1016/j.cell.2017.11.024
Kurtzman CP, Robnett CJ (1998) Identification and phylogeny of ascomycetous yeasts from analysis of nuclear large subunit (26S) ribosomal DNA partial sequences. Antonie Van Leeuwenhoek 73(4):331–371
Lavasani S, Dzhambazov B, Nouri M, Fak F, Buske S, Molin G, Thorlacius H, Alenfall J, Jeppsson B, Westrom B (2010) A novel probiotic mixture exerts a therapeutic effect on experimental autoimmune encephalomyelitis mediated by IL-10 producing regulatory T cells. PLoS ONE 5(2):e9009. https://doi.org/10.1371/journal.pone.0009009
Lee JS, Tato CM, Joyce-Shaikh B, Gulen MF, Cayatte C, Chen Y, Blumenschein WM, Judo M, Ayanoglu G, McClanahan TK, Li X, Cua DJ (2015) Interleukin-23-independent IL-17 production regulates intestinal epithelial permeability. Immunity 43(4):727–738. https://doi.org/10.1016/j.immuni.2015.09.003
Lee NY, Lee HC, Ko NY, Chang CM, Shih HI, Wu CJ, Ko WC (2007) Clinical and economic impact of multidrug resistance in nosocomial Acinetobacter baumannii bacteremia. Infect Control Hosp Epidemiol 28(6):713–719. https://doi.org/10.1086/517954
Lee YK, Menezes JS, Umesaki Y, Mazmanian SK (2011) Proinflammatory T-cell responses to gut microbiota promote experimental autoimmune encephalomyelitis. Proc Natl Acad Sci U S A 108(Suppl 1):4615–4622. https://doi.org/10.1073/pnas.1000082107
Levison ME, Pitsakis PG (1987) Susceptibility to experimental Candida albicans urinary tract infection in the rat. J Infect Dis 155(5):841–846
Li X, Atkinson MA (2015) The role for gut permeability in the pathogenesis of type 1 diabetes—a solid or leaky concept? Pediatr Diabetes 16(7):485–492. https://doi.org/10.1111/pedi.12305
Lin HV, Frassetto A, Kowalik EJ Jr, Nawrocki AR, Lu MM, Kosinski JR, Hubert JA, Szeto D, Yao X, Forrest G, Marsh DJ (2012) Butyrate and propionate protect against diet-induced obesity and regulate gut hormones via free fatty acid receptor 3-independent mechanisms. PLoS ONE 7(4):e35240. https://doi.org/10.1371/journal.pone.0035240
Littman DR, Rudensky AY (2010) Th17 and regulatory T cells in mediating and restraining inflammation. Cell 140(6):845–858. https://doi.org/10.1016/j.cell.2010.02.021
Liu X, Zou Q, Zeng B, Fang Y, Wei H (2013) Analysis of fecal Lactobacillus community structure in patients with early rheumatoid arthritis. Curr Microbiol 67(2):170–176. https://doi.org/10.1007/s00284-013-0338-1
Lo HJ, Kohler JR, DiDomenico B, Loebenberg D, Cacciapuoti A, Fink GR (1997) Nonfilamentous C. albicans mutants are avirulent. Cell 90(5):939–949
Lopez-Garcia B, Lee PH, Yamasaki K, Gallo RL (2005) Anti-fungal activity of cathelicidins and their potential role in Candida albicans skin infection. J Invest Dermatol 125(1):108–115. https://doi.org/10.1111/j.0022-202X.2005.23713.x
Louis P, Young P, Holtrop G, Flint HJ (2010) Diversity of human colonic butyrate-producing bacteria revealed by analysis of the butyryl-CoA:acetate CoA-transferase gene. Environ Microbiol 12(2):304–314. https://doi.org/10.1111/j.1462-2920.2009.02066.x
Maeda Y, Kurakawa T, Umemoto E, Motooka D, Ito Y, Gotoh K, Hirota K, Matsushita M, Furuta Y, Narazaki M, Sakaguchi N, Kayama H, Nakamura S, Iida T, Saeki Y, Kumanogoh A, Sakaguchi S, Takeda K (2016) Dysbiosis contributes to arthritis development via activation of autoreactive t cells in the intestine. Arthritis Rheumatol 68(11):2646–2661. https://doi.org/10.1002/art.39783
Maes M, Kubera M, Leunis JC, Berk M (2012) Increased IgA and IgM responses against gut commensals in chronic depression: further evidence for increased bacterial translocation or leaky gut. J Affect Disord 141(1):55–62. https://doi.org/10.1016/j.jad.2012.02.023
Maes M, Kubera M, Leunis JC, Berk M, Geffard M, Bosmans E (2013) In depression, bacterial translocation may drive inflammatory responses, oxidative and nitrosative stress (O&NS), and autoimmune responses directed against O&NS-damaged neoepitopes. Acta Psychiatr Scand 127(5):344–354. https://doi.org/10.1111/j.1600-0447.2012.01908.x
Makino H, Kushiro A, Ishikawa E, Muylaert D, Kubota H, Sakai T, Oishi K, Martin R, Ben Amor K, Oozeer R, Knol J, Tanaka R (2011) Transmission of intestinal Bifidobacterium longum subsp. longum strains from mother to infant, determined by multilocus sequencing typing and amplified fragment length polymorphism. Appl Environ Microbiol 77(19):6788–6793. https://doi.org/10.1128/AEM.05346-11
Malani AN, Psarros G, Malani PN, Kauffman CA (2011) Is age a risk factor for Candida glabrata colonisation? Mycoses 54(6):531–537. https://doi.org/10.1111/j.1439-0507.2010.01941.x
Markey L, Shaban L, Green ER, Lemon KP, Mecsas J, Kumamoto CA (2018) Pre-colonization with the commensal fungus Candida albicans reduces murine susceptibility to Clostridium difficile infection. Gut Microbes:0. https://doi.org/10.1080/19490976.2018.1465158
Markle JG, Frank DN, Mortin-Toth S, Robertson CE, Feazel LM, Rolle-Kampczyk U, von Bergen M, McCoy KD, Macpherson AJ, Danska JS (2013) Sex differences in the gut microbiome drive hormone-dependent regulation of autoimmunity. Science 339(6123):1084–1088. https://doi.org/10.1126/science.1233521
Martin R, Heilig HG, Zoetendal EG, Jimenez E, Fernandez L, Smidt H, Rodriguez JM (2007) Cultivation-independent assessment of the bacterial diversity of breast milk among healthy women. Res Microbiol 158(1):31–37. https://doi.org/10.1016/j.resmic.2006.11.004
Mason KL, Erb Downward JR, Falkowski NR, Young VB, Kao JY, Huffnagle GB (2012a) Interplay between the Gastric Bacterial Microbiota and Candida albicans during Postantibiotic Recolonization and Gastritis. Infect Immun 80(1):150–158. https://doi.org/10.1128/IAI.05162-11
Mason KL, Erb Downward JR, Mason KD, Falkowski NR, Eaton KA, Kao JY, Young VB, Huffnagle GB (2012b) Candida albicans and bacterial microbiota interactions in the cecum during recolonization following broad-spectrum antibiotic therapy. Infect Immun 80(10):3371–3380. https://doi.org/10.1128/iai.00449-12
Massot J, Sanchez O, Couchy R, Astoin J, Parodi AL (1984) Bacterio-pharmacological activity of Saccharomyces boulardii in clindamycin-induced colitis in the hamster. Arzneimittelforschung 34(7):794–797
Mazzini E, Massimiliano L, Penna G, Rescigno M (2014) Oral tolerance can be established via gap junction transfer of fed antigens from CX3CR1(+) macrophages to CD103(+) dendritic cells. Immunity 40(2):248–261. https://doi.org/10.1016/j.immuni.2013.12.012
McDole JR, Wheeler LW, McDonald KG, Wang B, Konjufca V, Knoop KA, Newberry RD, Miller MJ (2012) Goblet cells deliver luminal antigen to CD103+ dendritic cells in the small intestine. Nature 483(7389):345–349. https://doi.org/10.1038/nature10863
Meador JP, Caldwell ME, Cohen PS, Conway T (2014) Escherichia coli pathotypes occupy distinct niches in the mouse intestine. Infect Immun 82(5):1931–1938. https://doi.org/10.1128/iai.01435-13
Medeiros AO, Kohler LM, Hamdan JS, Missagia BS, Barbosa FA, Rosa CA (2008) Diversity and antifungal susceptibility of yeasts from tropical freshwater environments in Southeastern Brazil. Water Res 42(14):3921–3929. https://doi.org/10.1016/j.watres.2008.05.026
Misic AM, Davis MF, Tyldsley AS, Hodkinson BP, Tolomeo P, Hu B, Nachamkin I, Lautenbach E, Morris DO, Grice EA (2015) The shared microbiota of humans and companion animals as evaluated from Staphylococcus carriage sites. Microbiome 3:2. https://doi.org/10.1186/s40168-014-0052-7
Miyake S, Kim S, Suda W, Oshima K, Nakamura M, Matsuoka T, Chihara N, Tomita A, Sato W, Kim SW, Morita H, Hattori M, Yamamura T (2015) Dysbiosis in the gut microbiota of patients with multiple sclerosis, with a striking depletion of species belonging to clostridia XIVa and IV clusters. PLoS ONE 10(9):e0137429. https://doi.org/10.1371/journal.pone.0137429
Mohamadi J, Motaghi M, Panahi J, Havasian MR, Delpisheh A, Azizian M, Pakzad I (2014) Anti-fungal resistance in candida isolated from oral and diaper rash candidiasis in neonates. Bioinformation 10(11):667–670. https://doi.org/10.6026/97320630010667
Mootsikapun P (2007) Bacteremia in adult patients with acquired immunodeficiency syndrome in the northeast of Thailand. Int J Infect Dis 11(3):226–231. https://doi.org/10.1016/j.ijid.2006.02.010
Morschhäuser J (2010) Regulation of white-opaque switching in Candida albicans. Med Microbiol Immunol 199(3):165–172. https://doi.org/10.1007/s00430-010-0147-0
Nagata R, Nagano H, Ogishima D, Nakamura Y, Hiruma M, Sugita T (2012) Transmission of the major skin microbiota, Malassezia, from mother to neonate. Pediatr In 54 (3):350–355. https://doi.org/10.1111/j.1442-200x.2012.03563.x
Nakagawa T, Mori N, Kajiwara C, Kimura S, Akasaka Y, Ishii Y, Saji T, Tateda K (2016) Endogenous IL-17 as a factor determining the severity of Clostridium difficile infection in mice. J Med Microbiol 65(8):821–827. https://doi.org/10.1099/jmm.0.000273
Nava P, Koch S, Laukoetter MG, Lee WY, Kolegraff K, Capaldo CT, Beeman N, Addis C, Gerner-Smidt K, Neumaier I, Skerra A, Li L, Parkos CA, Nusrat A (2010) Interferon-gamma regulates intestinal epithelial homeostasis through converging beta-catenin signaling pathways. Immunity 32(3):392–402. https://doi.org/10.1016/j.immuni.2010.03.001
Nerandzic MM, Mullane K, Miller MA, Babakhani F, Donskey CJ (2012) Reduced acquisition and overgrowth of vancomycin-resistant enterococci and Candida species in patients treated with fidaxomicin versus vancomycin for Clostridium difficile infection. Clin Infect Dis 55(Suppl 2):121–126. https://doi.org/10.1093/cid/cis440
Nielsen DS, Honholt S, Tano-Debrah K, Jespersen L (2005) Yeast populations associated with Ghanaian cocoa fermentations analysed using denaturing gradient gel electrophoresis (DGGE). Yeast 22(4):271–284. https://doi.org/10.1002/yea.1207
Nizet V, Johnson RS (2009) Interdependence of hypoxic and innate immune responses. Nat Rev Immunol 9(9):609–617. https://doi.org/10.1038/nri2607
Noble SM, Gianetti BA, Witchley JN (2017) Candida albicans cell-type switching and functional plasticity in the mammalian host. Nat Rev Microbiol 15(2):96–108. https://doi.org/10.1038/nrmicro.2016.157
Noverr MC, Huffnagle GB (2004) Regulation of Candida albicans morphogenesis by fatty acid metabolites. Infect Immun 72(11):6206–6210
Odds FC, Davidson AD, Jacobsen MD, Tavanti A, Whyte JA, Kibbler CC, Ellis DH, Maiden MC, Shaw DJ, Gow NA (2006) Candida albicans strain maintenance, replacement, and microvariation demonstrated by multilocus sequence typing. J Clin Microbiol 44(10):3647–3658
Palmer C, Bik EM, DiGiulio DB, Relman DA, Brown PO (2007) Development of the human infant intestinal microbiota. PLoS Biol 5(7):e177. https://doi.org/10.1371/journal.pbio.0050177
Pande K, Chen C, Noble SM (2013) Passage through the mammalian gut triggers a phenotypic switch that promotes Candida albicans commensalism. Nat Genet 45(9):1088–1091. https://doi.org/10.1038/ng.2710
Pappas PG (2006) Invasive candidiasis. Infect Dis Clin North Am 20(3):485–506. https://doi.org/10.1016/j.idc.2006.07.004
Pasqualotto AC, Nedel WL, Machado TS, Severo LC (2006) Risk factors and outcome for nosocomial breakthrough candidaemia. J Infect 52(3):216–222. https://doi.org/10.1016/j.jinf.2005.04.020
Peleg AY, Tampakakis E, Fuchs BB, Eliopoulos GM, Moellering RC Jr, Mylonakis E (2008) Prokaryote-eukaryote interactions identified by using Caenorhabditis elegans. Proc Natl Acad Sci U S A 105(38):14585–14590
Pennacchia C, Blaiotta G, Pepe O, Villani F (2008) Isolation of Saccharomyces cerevisiae strains from different food matrices and their preliminary selection for a potential use as probiotics. J Appl Microbiol 105(6):1919–1928. https://doi.org/10.1111/j.1365-2672.2008.03968.x
Pepin J, Saheb N, Coulombe MA, Alary ME, Corriveau MP, Authier S, Leblanc M, Rivard G, Bettez M, Primeau V, Nguyen M, Jacob CE, Lanthier L (2005) Emergence of fluoroquinolones as the predominant risk factor for Clostridium difficile-associated diarrhea: a cohort study during an epidemic in Quebec. Clin Infect Dis 41(9):1254–1260. https://doi.org/10.1086/496986
Pessione E (2012) Lactic acid bacteria contribution to gut microbiota complexity: lights and shadows. Front Cell Infect Microbiol 2:86. https://doi.org/10.3389/fcimb.2012.00086
Peyssonnaux C, Datta V, Cramer T, Doedens A, Theodorakis EA, Gallo RL, Hurtado-Ziola N, Nizet V, Johnson RS (2005) HIF-1alpha expression regulates the bactericidal capacity of phagocytes. J Clin Invest 115(7):1806–1815. https://doi.org/10.1172/jci23865
Pfaller MA, Moet GJ, Messer SA, Jones RN, Castanheira M (2011) Geographic variations in species distribution and echinocandin and azole antifungal resistance rates among Candida bloodstream infection isolates: report from the SENTRY antimicrobial surveillance program (2008–2009). J Clin Microbiol 49(1):396–399. https://doi.org/10.1128/jcm.01398-10
Pierce JV, Dignard D, Whiteway M, Kumamoto CA (2013) Normal adaptation of Candida albicans to the murine gastrointestinal tract requires Efg1p-dependent regulation of metabolic and host defense genes. Eukaryot Cell 12(1):37–49. https://doi.org/10.1128/EC.00236-12
Pulimood S, Ganesan L, Alangaden G, Chandrasekar P (2002) Polymicrobial candidemia. Diagn Microbiol Infect Dis 44(4):353–357
Qin J, Li R, Raes J, Arumugam M, Burgdorf KS, Manichanh C, Nielsen T, Pons N, Levenez F, Yamada T, Mende DR, Li J, Xu J, Li S, Li D, Cao J, Wang B, Liang H, Zheng H, Xie Y, Tap J, Lepage P, Bertalan M, Batto JM, Hansen T, Le Paslier D, Linneberg A, Nielsen HB, Pelletier E, Renault P, Sicheritz-Ponten T, Turner K, Zhu H, Yu C, Li S, Jian M, Zhou Y, Li Y, Zhang X, Li S, Qin N, Yang H, Wang J, Brunak S, Dore J, Guarner F, Kristiansen K, Pedersen O, Parkhill J, Weissenbach J, Bork P, Ehrlich SD, Wang J (2010) A human gut microbial gene catalogue established by metagenomic sequencing. Nature 464(7285):59–65. https://doi.org/10.1038/nature08821
Raffatellu M, George MD, Akiyama Y, Hornsby MJ, Nuccio SP, Paixao TA, Butler BP, Chu H, Santos RL, Berger T, Mak TW, Tsolis RM, Bevins CL, Solnick JV, Dandekar S, Baumler AJ (2009) Lipocalin-2 resistance confers an advantage to Salmonella enterica serotype Typhimurium for growth and survival in the inflamed intestine. Cell Host Microbe 5(5):476–486. https://doi.org/10.1016/j.chom.2009.03.011
Rajilic-Stojanovic M, Heilig HG, Tims S, Zoetendal EG, de Vos WM (2012) Long-term monitoring of the human intestinal microbiota composition. Environ Microbiol. https://doi.org/10.1111/1462-2920.12023
Rea MC, Dobson A, O’Sullivan O, Crispie F, Fouhy F, Cotter PD, Shanahan F, Kiely B, Hill C, Ross RP (2011) Effect of broad- and narrow-spectrum antimicrobials on Clostridium difficile and microbial diversity in a model of the distal colon. Proc Natl Acad Sci U S A 108(Suppl 1):4639–4644. https://doi.org/10.1073/pnas.1001224107
Rea MC, Sit CS, Clayton E, O’Connor PM, Whittal RM, Zheng J, Vederas JC, Ross RP, Hill C (2010) Thuricin CD, a posttranslationally modified bacteriocin with a narrow spectrum of activity against Clostridium difficile. Proc Natl Acad Sci U S A 107(20):9352–9357. https://doi.org/10.1073/pnas.0913554107
Roilides E, Farmaki E, Evdoridou J, Francesconi A, Kasai M, Filioti J, Tsivitanidou M, Sofianou D, Kremenopoulos G, Walsh TJ (2003) Candida tropicalis in a neonatal intensive care unit: epidemiologic and molecular analysis of an outbreak of infection with an uncommon neonatal pathogen. J Clin Microbiol 41(2):735–741
Rosenbach A, Dignard D, Pierce JV, Whiteway M, Kumamoto CA (2010) Adaptations of Candida albicans for growth in the mammalian intestinal tract. Eukaryot Cell 9(7):1075–1086. https://doi.org/10.1128/EC.00034-10
Rosser EC, Mauri C (2016) A clinical update on the significance of the gut microbiota in systemic autoimmunity. J Autoimmun 74:85–93. https://doi.org/10.1016/j.jaut.2016.06.009
Rubino SJ, Geddes K, Girardin SE (2012) Innate IL-17 and IL-22 responses to enteric bacterial pathogens. Trends Immunol 33(3):112–118. https://doi.org/10.1016/j.it.2012.01.003
Ruiz-Herrera J, Victoria Elorza M, Valentín E, Sentandreu R (2006) Molecular organization of the cell wall of Candida albicans and its relation to pathogenicity. FEMS Yeast Res 6(1):14–29. https://doi.org/10.1111/j.1567-1364.2005.00017.x
Rutayisire E, Huang K, Liu Y, Tao F (2016) The mode of delivery affects the diversity and colonization pattern of the gut microbiota during the first year of infants’ life: a systematic review. BMC Gastroenterol 16(1):86. https://doi.org/10.1186/s12876-016-0498-0
Sampson TR, Debelius JW, Thron T, Janssen S, Shastri GG, Ilhan ZE, Challis C, Schretter CE, Rocha S, Gradinaru V, Chesselet MF, Keshavarzian A, Shannon KM, Krajmalnik-Brown R, Wittung-Stafshede P, Knight R, Mazmanian SK (2016) Gut microbiota regulate motor deficits and neuroinflammation in a model of Parkinson’s disease. Cell 167 (6):1469–1480 e1412. https://doi.org/10.1016/j.cell.2016.11.018
Sassone-Corsi M, Nuccio SP, Liu H, Hernandez D, Vu CT, Takahashi AA, Edwards RA, Raffatellu M (2016) Microcins mediate competition among Enterobacteriaceae in the inflamed gut. Nature 540(7632):280–283. https://doi.org/10.1038/nature20557
Sato J, Kanazawa A, Ikeda F, Yoshihara T, Goto H, Abe H, Komiya K, Kawaguchi M, Shimizu T, Ogihara T, Tamura Y, Sakurai Y, Yamamoto R, Mita T, Fujitani Y, Fukuda H, Nomoto K, Takahashi T, Asahara T, Hirose T, Nagata S, Yamashiro Y, Watada H (2014) Gut dysbiosis and detection of “live gut bacteria” in blood of Japanese patients with type 2 diabetes. Diabetes Care 37(8):2343–2350. https://doi.org/10.2337/dc13-2817
Sawa N, Wilaipun P, Kinoshita S, Zendo T, Leelawatcharamas V, Nakayama J, Sonomoto K (2012) Isolation and characterization of enterocin W, a novel two-peptide lantibiotic produced by Enterococcus faecalis NKR-4-1. Appl Environ Microbiol 78(3):900–903. https://doi.org/10.1128/aem.06497-11
Scher JU, Sczesnak A, Longman RS, Segata N, Ubeda C, Bielski C, Rostron T, Cerundolo V, Pamer EG, Abramson SB, Huttenhower C, Littman DR (2013) Expansion of intestinal Prevotella copri correlates with enhanced susceptibility to arthritis. Elife 2:e01202. https://doi.org/10.7554/eLife.01202
Schlecht LM, Peters BM, Krom BP, Freiberg JA, Hansch GM, Filler SG, Jabra-Rizk MA, Shirtliff ME (2015) Systemic Staphylococcus aureus infection mediated by Candida albicans hyphal invasion of mucosal tissue. Microbiology 161(Pt 1):168–181. https://doi.org/10.1099/mic.0.083485-0
Seno H, Miyoshi H, Brown SL, Geske MJ, Colonna M, Stappenbeck TS (2009) Efficient colonic mucosal wound repair requires Trem2 signaling. Proc Natl Acad Sci U S A 106(1):256–261. https://doi.org/10.1073/pnas.0803343106
Sharma R, Young C, Neu J (2010) Molecular modulation of intestinal epithelial barrier: contribution of microbiota. J Biomed Biotechnol 2010:305879. https://doi.org/10.1155/2010/305879
Shi Y, Mu L (2017) An expanding stage for commensal microbes in host immune regulation. Cell Mol Immunol 14(4):339–348. https://doi.org/10.1038/cmi.2016.64
Shimada Y, Kinoshita M, Harada K, Mizutani M, Masahata K, Kayama H, Takeda K (2013) Commensal bacteria-dependent indole production enhances epithelial barrier function in the colon. PLoS ONE 8(11):e80604. https://doi.org/10.1371/journal.pone.0080604
Shiokawa A, Kotaki R, Takano T, Nakajima-Adachi H, Hachimura S (2017) Mesenteric lymph node CD11b(−) CD103(+) PD-L1(High) dendritic cells highly induce regulatory T cells. Immunology 152(1):52–64. https://doi.org/10.1111/imm.12747
Slutsky B, Staebell M, Anderson J, Risen L, Pfaller M, Soll DR (1987) “White-opaque transition”: a second high-frequency switching system in Candida albicans. J Bacteriol 169(1):189–197
Sohnle PG, Hahn BL, Santhanagopalan V (1996) Inhibition of Candida albicans growth by calprotectin in the absence of direct contact with the organisms. J Infect Dis 174(6):1369–1372
Sonnenburg JL, Chen CT, Gordon JI (2006) Genomic and metabolic studies of the impact of probiotics on a model gut symbiont and host. PLoS Biol 4(12):e413. https://doi.org/10.1371/journal.pbio.0040413
Stadhouders R, Lubberts E, Hendriks RW (2018) A cellular and molecular view of T helper 17 cell plasticity in autoimmunity. J Autoimmun 87:1–15. https://doi.org/10.1016/j.jaut.2017.12.007
Steimle A, Frick JS (2016) Molecular mechanisms of induction of tolerant and tolerogenic intestinal dendritic cells in mice. J Immunol Res 2016:1958650. https://doi.org/10.1155/2016/1958650
Steimle A, Gronbach K, Beifuss B, Schafer A, Harmening R, Bender A, Maerz JK, Lange A, Michaelis L, Maurer A, Menz S, McCoy K, Autenrieth IB, Kalbacher H, Frick JS (2016) Symbiotic gut commensal bacteria act as host cathepsin S activity regulators. J Autoimmun 75:82–95. https://doi.org/10.1016/j.jaut.2016.07.009
Stelter C, Kappeli R, Konig C, Krah A, Hardt WD, Stecher B, Bumann D (2011) Salmonella-induced mucosal lectin RegIIIbeta kills competing gut microbiota. PLoS ONE 6(6):e20749. https://doi.org/10.1371/journal.pone.0020749
Stinson LF, Payne MS, Keelan JA (2017) Planting the seed: Origins, composition, and postnatal health significance of the fetal gastrointestinal microbiota. Crit Rev Microbiol 43(3):352–369. https://doi.org/10.1080/1040841X.2016.1211088
Stockinger B, Omenetti S (2017) The dichotomous nature of T helper 17 cells. Nat Rev Immunol 17(9):535–544. https://doi.org/10.1038/nri.2017.50
Stoldt VR, Sonneborn A, Leuker CE, Ernst JF (1997) Efg1p, an essential regulator of morphogenesis of the human pathogen Candida albicans, is a member of a conserved class of bHLH proteins regulating morphogenetic processes in fungi. EMBO J 16(8):1982–1991
Strati F, Di Paola M, Stefanini I, Albanese D, Rizzetto L, Lionetti P, Calabro A, Jousson O, Donati C, Cavalieri D, De Filippo C (2016) Age and gender affect the composition of fungal population of the human gastrointestinal tract. Front Microbiol 7:1227. https://doi.org/10.3389/fmicb.2016.01227
Sudbery PE (2011) Growth of Candida albicans hyphae. Nat Rev Microbiol 9(10):737–748. https://doi.org/10.1038/nrmicro2636
Suh SO, Nguyen NH, Blackwell M (2008) Yeasts isolated from plant-associated beetles and other insects: seven novel Candida species near Candida albicans. FEMS Yeast Res 8(1):88–102. https://doi.org/10.1111/j.1567-1364.2007.00320.x
Tang MS, Bowcutt R, Leung JM, Wolff MJ, Gundra UM, Hudesman D, Malter LB, Poles MA, Chen LA, Pei Z, Neto AG, Abidi WM, Ullman T, Mayer L, Bonneau RA, Cho I, Loke P (2017) Integrated analysis of biopsies from inflammatory bowel disease patients identifies SAA1 as a link between mucosal microbes with TH17 and TH22 cells. Inflamm Bowel Dis 23(9):1544–1554. https://doi.org/10.1097/MIB.0000000000001208
Tao L, Du H, Guan G, Dai Y, Nobile CJ, Liang W, Cao C, Zhang Q, Zhong J, Huang G (2014) Discovery of a “white-gray-opaque” tristable phenotypic switching system in Candida albicans: roles of non-genetic diversity in host adaptation. PLoS Biol 12(4):e1001830. https://doi.org/10.1371/journal.pbio.1001830
Tezuka H, Abe Y, Asano J, Sato T, Liu J, Iwata M, Ohteki T (2011) Prominent role for plasmacytoid dendritic cells in mucosal T cell-independent IgA induction. Immunity 34(2):247–257. https://doi.org/10.1016/j.immuni.2011.02.002
Thewes S, Kretschmar M, Park H, Schaller M, Filler SG, Hube B (2007) In vivo and ex vivo comparative transcriptional profiling of invasive and non-invasive Candida albicans isolates identifies genes associated with tissue invasion. Mol Microbiol 63(6):1606–1628
Torosantucci A, Romagnoli G, Chiani P, Stringaro A, Crateri P, Mariotti S, Teloni R, Arancia G, Cassone A, Nisini R (2004) Candida albicans yeast and germ tube forms interfere differently with human monocyte differentiation into dendritic cells: a novel dimorphism-dependent mechanism to escape the host’s immune response. Infect Immun 72(2):833–843
Trofa D, Gacser A, Nosanchuk JD (2008) Candida parapsilosis, an emerging fungal pathogen. Clin Microbiol Rev 21(4):606–625. https://doi.org/10.1128/CMR.00013-08
Ubeda C, Djukovic A, Isaac S (2017) Roles of the intestinal microbiota in pathogen protection. Clin Transl Immunology 6(2):e128. https://doi.org/10.1038/cti.2017.2
Underhill DM, Iliev ID (2014) The mycobiota: interactions between commensal fungi and the host immune system. Nat Rev Immunol 14(6):405–416. https://doi.org/10.1038/nri3684
Vaishnava S, Yamamoto M, Severson KM, Ruhn KA, Yu X, Koren O, Ley R, Wakeland EK, Hooper LV (2011) The antibacterial lectin RegIIIgamma promotes the spatial segregation of microbiota and host in the intestine. Science 334(6053):255–258. https://doi.org/10.1126/science.1209791
Valentini L, Ramminger S, Haas V, Postrach E, Werich M, Fischer A, Koller M, Swidsinski A, Bereswill S, Lochs H, Schulzke JD (2014) Small intestinal permeability in older adults. Physiol Rep 2(4):e00281. https://doi.org/10.14814/phy2.281
van Langelaar J, van der Vuurst de Vries RM, Janssen M, Wierenga-Wolf AF, Spilt IM, Siepman TA, Dankers W, Verjans G, de Vries HE, Lubberts E, Hintzen RQ, van Luijn MM (2018) T helper 17.1 cells associate with multiple sclerosis disease activity: perspectives for early intervention. Brain 141 (5):1334–1349. https://doi.org/10.1093/brain/awy069
van Leeuwen PT, van der Peet JM, Bikker FJ, Hoogenkamp MA, Oliveira Paiva AM, Kostidis S, Mayboroda OA, Smits WK, Krom BP (2016) Interspecies Interactions between Clostridium difficile and Candida albicans. mSphere 1(6). https://doi.org/10.1128/msphere.00187-16
van Nood E, Vrieze A, Nieuwdorp M, Fuentes S, Zoetendal EG, de Vos WM, Visser CE, Kuijper EJ, Bartelsman JF, Tijssen JG, Speelman P, Dijkgraaf MG, Keller JJ (2013) Duodenal infusion of donor feces for recurrent Clostridium difficile. N Engl J Med 368(5):407–415. https://doi.org/10.1056/NEJMoa1205037
Vasquez A, Jakobsson T, Ahrne S, Forsum U, Molin G (2002) Vaginal Lactobacillus flora of healthy Swedish women. J Clin Microbiol 40(8):2746–2749
Vautier S, Drummond RA, Chen K, Murray GI, Kadosh D, Brown AJ, Gow NA, MacCallum DM, Kolls JK, Brown GD (2015) Candida albicans colonization and dissemination from the murine gastrointestinal tract: the influence of morphology and Th17 immunity. Cell Microbiol 17(4):445–450. https://doi.org/10.1111/cmi.12388
Venkatesh M, Mukherjee S, Wang H, Li H, Sun K, Benechet AP, Qiu Z, Maher L, Redinbo MR, Phillips RS, Fleet JC, Kortagere S, Mukherjee P, Fasano A, Le Ven J, Nicholson JK, Dumas ME, Khanna KM, Mani S (2014) Symbiotic bacterial metabolites regulate gastrointestinal barrier function via the xenobiotic sensor PXR and Toll-like receptor 4. Immunity 41(2):296–310. https://doi.org/10.1016/j.immuni.2014.06.014
Walker LA, Maccallum DM, Bertram G, Gow NA, Odds FC, Brown AJ (2009) Genome-wide analysis of Candida albicans gene expression patterns during infection of the mammalian kidney. Fungal Genet Biol 46(2):210–219
Wang Y, Xu XL (2008) Bacterial peptidoglycan-derived molecules activate Candida albicans hyphal growth. Commun Integr Biol 1(2):137–139
Ward TL, Dominguez-Bello MG, Heisel T, Al-Ghalith G, Knights D, Gale CA (2018) Development of the human mycobiome over the first month of life and across body sites. mSystems 3(3). https://doi.org/10.1128/msystems.00140-17
West CE (2014) Gut microbiota and allergic disease: new findings. Curr Opin Clin Nutr Metab Care 17(3):261–266. https://doi.org/10.1097/MCO.0000000000000044
White SJ, Rosenbach A, Lephart P, Nguyen D, Benjamin A, Tzipori S, Whiteway M, Mecsas J, Kumamoto CA (2007) Self-regulation of Candida albicans population size during GI colonization. PLoS Pathog 3(12):e184
Wu GD, Compher C, Chen EZ, Smith SA, Shah RD, Bittinger K, Chehoud C, Albenberg LG, Nessel L, Gilroy E, Star J, Weljie AM, Flint HJ, Metz DC, Bennett MJ, Li H, Bushman FD, Lewis JD (2016) Comparative metabolomics in vegans and omnivores reveal constraints on diet-dependent gut microbiota metabolite production. Gut 65(1):63–72. https://doi.org/10.1136/gutjnl-2014-308209
Wu HJ, Ivanov II, Darce J, Hattori K, Shima T, Umesaki Y, Littman DR, Benoist C, Mathis D (2010) Gut-residing segmented filamentous bacteria drive autoimmune arthritis via T helper 17 cells. Immunity 32(6):815–827. https://doi.org/10.1016/j.immuni.2010.06.001
Wu HJ, Wu E (2012) The role of gut microbiota in immune homeostasis and autoimmunity. Gut Microbes 3(1):4–14. https://doi.org/10.4161/gmic.19320
Yang W, Zhou Y, Wu C, Tang J (2016) Enterohemorrhagic Escherichia coli promotes the invasion and tissue damage of enterocytes infected with Candida albicans in vitro. Sci Rep 6:37485. https://doi.org/10.1038/srep37485
Yang YL, Lin CC, Chang TP, Lauderdale TL, Chen HT, Lee CF, Hsieh CW, Chen PC, Lo HJ (2012) Comparison of human and soil Candida tropicalis isolates with reduced susceptibility to fluconazole. PLoS ONE 7(4):e34609. https://doi.org/10.1371/journal.pone.0034609
Yapar N (2014) Epidemiology and risk factors for invasive candidiasis. Ther Clin Risk Manag 10:95–105. https://doi.org/10.2147/TCRM.S40160
Yatsunenko T, Rey FE, Manary MJ, Trehan I, Dominguez-Bello MG, Contreras M, Magris M, Hidalgo G, Baldassano RN, Anokhin AP, Heath AC, Warner B, Reeder J, Kuczynski J, Caporaso JG, Lozupone CA, Lauber C, Clemente JC, Knights D, Knight R, Gordon JI (2012) Human gut microbiome viewed across age and geography. Nature 486(7402):222–227. https://doi.org/10.1038/nature11053
Yu S, Gao N (2015) Compartmentalizing intestinal epithelial cell toll-like receptors for immune surveillance. Cell Mol Life Sci 72(17):3343–3353. https://doi.org/10.1007/s00018-015-1931-1
Yu XY, Fu F, Kong WN, Xuan QK, Wen DH, Chen XQ, He YM, He LH, Guo J, Zhou AP, Xi YH, Ni LJ, Yao YF, Wu WJ (2018) Streptococcus agalactiae Inhibits Candida albicans hyphal development and diminishes host vaginal mucosal TH17 response. Front Microbiol 9:198. https://doi.org/10.3389/fmicb.2018.00198
Zakikhany K, Naglik JR, Schmidt-Westhausen A, Holland G, Schaller M, Hube B (2007) In vivo transcript profiling of Candida albicans identifies a gene essential for interepithelial dissemination. Cell Microbiol 9(12):2938–2954
Ze X, Duncan SH, Louis P, Flint HJ (2012) Ruminococcus bromii is a keystone species for the degradation of resistant starch in the human colon. ISME J 6(8):1535–1543. https://doi.org/10.1038/ismej.2012.4
Zhang Z, Xu D, Wang L, Hao J, Wang J, Zhou X, Wang W, Qiu Q, Huang X, Zhou J, Long R, Zhao F, Shi P (2016) Convergent evolution of rumen microbiomes in high-altitude mammals. Curr Biol 26(14):1873–1879. https://doi.org/10.1016/j.cub.2016.05.012
Zheng L, Kelly CJ, Colgan SP (2015) Physiologic hypoxia and oxygen homeostasis in the healthy intestine. A review in the theme: cellular responses to hypoxia. Am J Physiol Cell Physiol 309(6):350–360. https://doi.org/10.1152/ajpcell.00191.2015
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2018 Springer Nature Switzerland AG
About this chapter
Cite this chapter
Kapitan, M., Niemiec, M.J., Steimle, A., Frick, J.S., Jacobsen, I.D. (2018). Fungi as Part of the Microbiota and Interactions with Intestinal Bacteria. In: Rodrigues, M. (eds) Fungal Physiology and Immunopathogenesis . Current Topics in Microbiology and Immunology, vol 422. Springer, Cham. https://doi.org/10.1007/82_2018_117
Download citation
DOI: https://doi.org/10.1007/82_2018_117
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-030-30236-8
Online ISBN: 978-3-030-30237-5
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)