Abstract
Histophilus somni is a commensal and an opportunistic bacterial pathogen associated with multisystemic diseases in cattle and sheep. Some strains of H. somni isolated from the genital tract of cattle are biochemically and serologically similar to the pathogenic strains, but are relatively innocuous. Several virulence factors/mechanisms have been identified in H. somni, of which the phase-variable lipooligosaccharide, induction of apoptosis of host cells, intraphagocytic survival, and immunoglobulin Fc-binding proteins have been well characterized. The genomes of H. somni pneumonia strain 2336 and preputial strain 129Pt have also been sequenced, and comparative analyses of these genomes have provided novel insights into the role of horizontal gene transfer in the evolution of the respective strains. Continued analyses of the genomes of H. somni strains and comparing them to the newly sequenced genomes of other bacteria facilitated the identification of a putative integrative and conjugative element (designated ICEHso2336) encoding tetracycline resistance. Comparative genomics also showed that the uptake signal sequence (5′-AAGTGCGGT) of Haemophilus influenzae is abundant in H. somni and provided a genetic basis for the recalcitrance of some strains of this species to natural transformation. The post-genomic era for H. somni offered an opportunity for the functional characterization of genes identified by computational methods. This opportunity has been realized to a great extent by transcriptomic studies that have identified several small noncoding RNAs and new genes. These new discoveries and developments are expected to stimulate further in-depth investigations of H. somni, especially from the systems biology viewpoint.
Access provided by Autonomous University of Puebla. Download chapter PDF
Similar content being viewed by others
Keywords
- Genomic Island
- Major Outer Membrane Protein
- Conjugative Element
- Allelic Exchange
- Actinobacillus Actinomycetemcomitans
These keywords were added by machine and not by the authors. This process is experimental and the keywords may be updated as the learning algorithm improves.
1 Introduction to Genomic Analyses
The terms “genomics” and “genomic methods” describe “the molecular and bioinformatics techniques that employ all or part of the genome to answer a question about an organism or a group of organisms” (Carruthers et al. 2012). Genomics has immense applications in the quest to understand nature, and comparative microbial genomics is an indispensable tool for molecular pathogenic bacteriology. The first complete genome sequence from a free-living organism was that of Haemophilus influenzae strain Rd KW20 (Fleischmann et al. 1995), a close relative of H. somni. This pioneering work at the erstwhile Institute for Genomic Research (TIGR) popularized the concept of whole-genome random sequencing by the “shotgun” approach. Since the completion of the first bacterial genome sequence, thousands of bacterial and archaeal genomes have been sequenced and annotated using novel tools and techniques. The genomes of several species of Pasteurellaceae have also been sequenced, and whole-genome comparisons have provided new insights into the physiology and evolution of members within this very important bacterial family (Challacombe and Inzana 2008).
Numerous in vitro and in vivo studies during the pre-genomic era have shed light on the differences in virulence properties and genetic traits between H. somni pathogenic isolates from sick animals and commensal isolates from the genital tract (Corbeil et al. 1995). H. somni pneumonia strain 2336 (NCBI taxonomy ID 228400) and preputial strain 129Pt (NCBI taxonomy ID 205914) have been phenotypically well characterized in the laboratory and utilized in several comparative studies (Corbeil et al. 1997; Inzana et al. 1992, 2002). The genomes of these two strains have been completely sequenced and compared (Challacombe et al. 2007; Siddaramappa et al. 2011). This chapter will provide an overview of the pre-genomic investigations, comparative genomic analyses, and post-genomic studies of H. somni strains.
2 Comparative Genomics
Several temperate bacteriophages that infect strains of H. influenzae have been purified and described (Williams et al. 2002). However, temperate bacteriophages that infect strains of H. somni remain to be isolated and characterized. Nevertheless, prophages and their associated sequences appear to be rife in the genome of H. somni strain 2336, but less abundant in the genome of strain 129Pt (Siddaramappa et al. 2011), indicating that the natural repertoire of bacteriophages that potentially infect some strains of H. somni strains could be large. Furthermore, a large portion of strain-specific sequences occurring in strains 2336 and 129Pt appear to be due to prophages and their associated sequences (Siddaramappa et al. 2011). Although the Mu-like prophage (FluMu) found in H. influenzae strain Rd KW20 is absent in the genomes of H. somni strains, the genome of H. somni strain 2336 contains a prophage that appears to be partially related to the Mannheimia haemolytica serotype A1 lysogenic bacteriophage ϕMhaA1-PHL101.
In addition to the prophages and their associated sequences, the genomes of H. somni strains contain several genomic islands that appear to be unrelated to each other (Siddaramappa et al. 2011). A genomic island that is homologous to ICEHin1056, which is a 59,393-bp integrative and conjugative element (~39 % G+C) containing genes encoding ampicillin, chloramphenicol, and tetracycline resistance in H. influenzae type b strain 1056, has also been identified in H. somni strain 2336 (Mohd-Zain et al. 2004). The genomic island of H. somni strain 2336 was more precisely delineated upon comparison with ICEPmu1, which is an integrative and conjugative element (~42 % G+C) containing genes encoding resistance to multiple antibiotics in Pasteurella multocida strain 36950 (Michael et al. 2012). This genomic island of H. somni strain 2336 appears to be a putative integrative and conjugative element and is referred to as ICEHso2336 (~40.5 % G+C). An integrative and conjugative element (ICEMh1, ~40 % G+C) containing genes encoding resistance to multiple antibiotics and closely related to ICEPmu1 is also present in M. haemolytica strain 42548 (Eidam et al. 2015). Whereas the nucleotide identity between ICEMh1 and ICEHin1056 is only ~70 %, the nucleotide identity between ICEMh1, ICEPmu1, and ICEHso2336 is ~99 %.
Furthermore, ICEPmu1 and ICEMh1 are integrated site-specifically into tRNALeu in the chromosomes of P. multocida strain 36950 and M. haemolytica strain 42548, respectively (Eidam et al. 2015; Michael et al. 2012). A comparison of these loci as well as ICEHso2336 indicated that each element contains 11-bp (5′-GATTTTGAATC) terminal direct repeats and an 86-bp tRNALeu at the right terminus (Fig. 1a). Although ICEPmu1 is smaller in size than ICEMh1 by ~10,000 bp, it contains more antimicrobial resistance genes than the latter (Eidam et al. 2015; Michael et al. 2012). As reported previously, ICEHso2336 contains the tetracycline repressor gene tetR and the tetracycline resistance gene tetH (Michael et al. 2012; Mohd-Zain et al. 2004; Siddaramappa et al. 2011), and H. somni strain 2336 is resistant to tetracycline (MIC 8 µg/ml) (Ueno et al. 2014). A schematic map of ICEHso2336 is shown in Fig. 1b, and a comparison of the ORFs that occur among ICEPmu1, ICEMh1, and ICEHso2336 is shown in Table 1. Although these elements are closely related, they are not identical and it is evident from Table 1 that they display a mosaic structure with alternating conserved and variable regions. In particular, ICEHso2336 lacked 22 ORFs found in ICEPmu1, and 13 of these 22 ORFs are also absent in ICEMh1. In contrast, ICEPmu1 and ICEMh1 lack 12 ORFs found in ICEHso2336. Interestingly, H. somni strain 129Pt lacks an analogous ICE, but contains short stretches of homologous sequences. Not surprisingly, most of the ORFs identified in ICEPmu1 and/or ICEHso2336 have distant homologs outside of the Pasteurellaceae.
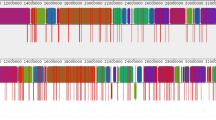
a Comparison of ICEMh1, ICEPmu1, and ICEHso2336. Each ICE contains 11-bp terminal direct repeats (DR1 and DR3) and an additional direct repeat (DR2) within the 86-bp tRNALeu gene (underlined). A 51-bp sequence occurs between the tRNALeu gene and DR3. These features are identical among the three ICEs. The sequence between DR1 and the tRNALeu gene (92110 bp in ICEMh1, 82,066 bp in ICEPmu1, and 66463 bp in ICEHso2336) contains genes that distinguish the three ICEs. Numbers above/below the maps (e.g., 2098396 and 2190664) indicate nucleotide positions within the respective chromosomes. b Schematic map of ICEHso2336. Terminal direct repeats (DR1 and DR3) shown in Fig. 1a are indicated by vertical black bars. Arrows represent ORFs found within ICEHso2336 and compared in Table 1. Blue arrows represent ORFs that have orthologs in ICEPmu1, but not in ICEMh1. Gray arrows represent ORFs that have orthologs in ICEPmu1 and ICEMh1. Black arrows represent ORFs that have no orthologs in ICEPmu1 and ICEMh1. Red arrows represent tetR and tetH (have orthologs in ICEPmu1 and ICEMh1). Gray arrows containing asterisks represent ORFs that have full-length or partial homologs in H. somni strain 129Pt
It is interesting to note that M. haemolytica strain 42548, P. multocida strain 36950, and H. somni strain 2336 were isolated from cases of naturally occurring bovine respiratory tract infections in different parts of the USA (Pennsylvania, Nebraska, and Washington, respectively) in different years (2007, 2005, and 1980s, respectively), but harbor closely related genomic elements containing antibiotic resistance determinants. Multidrug-resistant isolates of these respiratory pathogens appear to be more common among animals in bovine feedlots (Klima et al. 2014). Furthermore, horizontal transfer of ICEs that mediate antibiotic resistance from M. haemolytica and H. somni to P. multocida, and from P. multocida to Escherichia coli, has been demonstrated (Klima et al. 2014). It is possible that these ICEs have a common evolutionary origin, and indiscriminate use of antibiotics favors their preservation and dispersal in the field.
3 Comparative Transcriptomics
Computational gene prediction at best provides a “first pass” structural annotation of genomes and has many limitations, which could be overcome using experimental approaches that involve the analyses of the transcriptome. Attempts have been made to obtain a high-resolution transcriptome map of H. somni strain 2336 using the Illumina RNA-Seq technology (Kumar et al. 2012). Comparison of the transcriptome map of strain 2336 with the computationally annotated genome facilitated the identification of 94 small noncoding RNA (sRNA) of various sizes (70–695 bp, average G+C content 39.3 %). A vast majority of these sRNA (82 of 94) were reported to be novel (unidentified in previous bacterial transcriptome studies) and proposed to play roles in housekeeping and virulence, in addition to gene regulation. Sequence analyses of the 94 sRNA indicated that 31 were specific to strain 2336, 41 were specific to strains 2336 and 129Pt, 11 had homologs only in the genomes of P. multocida, H. influenzae, and H. parainfluenzae, and 11 had homologs in the genomes of other distantly related bacteria (Kumar et al. 2012). Furthermore, the start sites of five predicted genes (HSM_0031, 0525, 0789, 1019, and 1729) were corrected using the RNA-Seq data and comparison with other phylogenetically related homologs.
Genome annotation had predicted that putative proteins encoded by HSM_0603, 0748, 1385, 1666, and 1744 (hypothetical protein, α-L-fucosidase, 3-hydroxydecanoyl-ACP dehydratase, DNA damage-inducible protein, and alcohol dehydrogenase, respectively) were shorter than their homologs in other species. RNA-Seq data of strain 2336 showed the presence of full-length mRNA for these genes and confirmed that the putative proteins were truncated at the N-terminus due to either frameshift mutations (for HSM_1385 and 1744) or non-functional start codons (for the other three genes) (Kumar et al. 2012). Analyses of the RNA-Seq data indicated that 1636 of the 1980 predicted protein-coding genes were transcribed and there were 278 operons consisting of 730 genes in H. somni strain 2336 (Kumar et al. 2012).
4 Plasmids and Shuttle Vectors
Plasmid-borne resistance to multiple antibiotics is a relatively common feature among some members of the Pasteurellaceae. Isolates of H. somni resistant to tetracycline and harboring tetH, albeit lacking plasmids, have been cultured from nasal swabs of feedlot calves from Alberta, Canada (D’Amours et al. 2011). Furthermore, plasmid profiling as a means of identification and characterization of field isolates of H. somni has been reported (Appuhamy et al. 1998; Fussing and Wegener 1993). Efforts have also been made toward deciphering and describing the complete nucleotide sequences of plasmids from H. somni strains (Izadpanah et al. 2001; Siddaramappa et al. 2006). All four H. somni circular plasmids whose sequences have been deciphered/described are referred to as cryptic plasmids since they lack the genes that encode functions other than those necessary for their own replication (Izadpanah et al. 2001; Siddaramappa et al. 2006). Interestingly, the largest H. somni circular plasmid that has been completely sequenced (pHS129, 5178 bp) appears to be a dimer (Siddaramappa et al. 2006), and the natural occurrence of such plasmid dimers among bacteria is relatively rare.
The possibility of using native or non-native plasmids, after suitable modifications, as shuttle vectors that function in E. coli and H. somni has been explored. H. somni strain HS91 was transformed with plasmid pD70KanR, which is derived from M. haemolytica plasmid pD70 (Briggs and Tatum 2005). Interestingly, in vitro modification of pD70KanR using a commercially available HhaI methylase significantly improves the transformation efficiency (Briggs and Tatum 2005). Furthermore, H. somni strain 129Pt, which contains plasmid pHS129, can be transformed with pLS88, which is a broad-host-range plasmid purified from Haemophilus ducreyi (Sanders et al. 1997). In vivo modification of pLS88 using the recombination-deficient H. influenzae strain DB117 improves the transformation efficiency (Sanders et al. 1997).
H. somni strain 129Pt has also been transformed with a modified version of H. somni circular plasmid pHS649 (Siddaramappa et al. 2006). Derivatives of pLS88 that transform H. somni with a higher efficiency (e.g., pNS3K) have also been developed using kanamycin resistance as the selectable marker (Sandal et al. 2008). Therefore, it appears that pHS129 and pLS88 do not belong to the same incompatibility group, as are pHS129 and pHS649. The possibility of improving these vectors or other forms of pLS88 (such as pLSSK and pLSKS) (Wood et al. 1999) for efficient transformation of H. somni strains remains to be explored.
5 Mutagenesis
Although chemical mutagenesis is a popular technique in bacterial genetics and ethyl methanesulfonate has been used to obtain non-capsulated mutants of Actinobacillus pleuropneumoniae (Inzana et al. 1993), it has not been widely used in other members of the Pasteurellaceae. Transformation and mutagenesis of strains of H. somni using exogenous DNA molecules is difficult, at least in part due to an omnipresent restriction–modification system.
Molecular genetic analyses of H. somni were invigorated following the demonstration that in vitro or in vivo modification improves the transformation efficiency of shuttle plasmids for H. somni (Briggs and Tatum 2005; Sanders et al. 1997). Successful transformation of H. somni strain 129Pt with a putative virulence-associated gene of H. somni strain 2336, and the stable expression of the gene in the transformed strain, demonstrated the utility of H. somni preputial isolates for genetic analyses (Sanders et al. 1997). Furthermore, transformation of strain 129Pt was also used to demonstrate that lob1 is involved in lipooligosaccharide (LOS) biosynthesis in H. somni and that the 5′-(CAAT)n repeats within lob1 are involved in LOS phase variation (McQuiston et al. 2000) (see chapter on “The Many Facets of Lipooligosaccharide as a Virulence Factor for Histophilus somni”). These studies used a commercial electroporator to introduce heterologous DNA into H. somni rendered electrocompetent by growth in brain–heart infusion broth or Columbia broth and washing the bacterial pellets with 272 mM sucrose solution (Briggs and Tatum 2005; McQuiston et al. 2000; Sanders et al. 1997).
A non-replicative suicide plasmid methylated in vitro by HhaI methylase was used for mutagenesis of a H. somni strain 738 DNA locus involved in LOS biosynthesis by homologous recombination-mediated allelic exchange (Wu et al. 2000). The mutant strains had an altered LOS profile in comparison with the wild-type strain, indicating that lob2A could be involved in LOS biosynthesis (Wu et al. 2000). However, the prototype mutant strain (H. somni 738-lob2A1::Km) could not be complemented using shuttle vector pLSlob2A, reportedly due to inefficient electrotransformation (Wu et al. 2000) (see chapter on “The Many Facets of Lipooligosaccharide as a Virulence Factor for Histophilus somni”).
A combination of methylation in vivo using the H. influenzae cloning strain DB117 and in vitro using HhaI methylase has been shown to improve the transformation efficiency of plasmids for H. somni strain 8025 (Sanders et al. 2003). A fivefold increase in transformation efficiency is observed after plasmids derived from H. somni strain 8025 are reintroduced into the same strain by electroporation (Sanders et al. 2003), indicating that the restriction–modification systems among H. influenzae and H. somni strains could be different. Furthermore, homologous recombination-mediated allelic exchange was used for partial deletion of a locus encoding high molecular weight immunoglobulin-binding proteins (HMW IgBPs) in H. somni strain 8025 (Sanders et al. 2003). A significant difference (p < 0.001) in the adherence of the mutant or wild-type strain to bovine pulmonary artery endothelial cells was also reported (Sanders et al. 2003). Of interest is that both lob1 and the gene encoding for HMW IgBPs contain the H. influenzae uptake signal sequence (see Sect. 6).
A temperature-sensitive plasmid was developed to obtain in-frame, unmarked aroA deletion mutants of H. somni (Tatum and Briggs 2005). M. haemolytica native plasmid pD70 was modified by inserting the Tn903 kanamycin resistance cassette and the modified plasmid (pD70KanR) mutagenized using hydroxylamine. A single base-pair mutation from G to A at position 301 within the origin of replication renders this plasmid temperature sensitive. The aroA gene from H. somni was amplified by PCR and cloned into the temperature-sensitive plasmid pGA301oriC to create pTsHsaroC. An in-frame deletion was engineered within pTsHsaroC to create the replacement plasmid pTsHsΔaroAC (Tatum and Briggs 2005). This replacement plasmid is methylated in vitro using HhaI methylase, electroporated into H. somni strain 2336, and recovered at the permissive temperature of 30 °C for 2 h on medium containing 50 µg/ml kanamycin. The plates are then incubated at the non-permissive temperature (41 °C) for 16 h to select for single-crossover mutants containing the temperature-sensitive replacement plasmid integrated into the chromosome by homologous recombination (Tatum and Briggs 2005). Single-crossover mutants are cultured in broth without kanamycin at the permissive temperature for 16 h to facilitate a second crossover event and plasmid excision. This process is repeated twice, and bacteria from the third-pass culture are streaked onto plates without kanamycin. The plates are incubated at 37 °C for 16 h, and colonies are further replica-plated with or without kanamycin selection. After incubation at 37 °C, kanamycin-sensitive colonies are selected and the absence of the kanamycin gene is tested by PCR. Deletion of aroA is also confirmed by PCR (Tatum and Briggs 2005).
A non-replicative suicide plasmid methylated in vitro using HhaI methylase can also be used for complete deletion of the ibpA open reading frame encoding HMW IgBPs in H. somni strain 2336 by homologous recombination-mediated allelic exchange (Hoshinoo et al. 2009). The isogenic mutant strain was less cytotoxic than wild-type strain 2336 for bovine FBM-17 macrophage-like cells, murine J774.1 macrophage-like cells, and bovine primary monocyte cells (Hoshinoo et al. 2009). Although wild-type strain 2336 significantly compromised the ability of murine J774.1 macrophage-like cells and bovine primary monocyte cells to phagocytize microspheres, the isogenic mutant strain had no such effect, indicating that IbpA (specifically the Fic region; see chapter on “Histophilus somni Surface Proteins”) of H. somni may play a role in pathogenesis (Hoshinoo et al. 2009).
Homologous recombination-mediated exchange of genes encoding the major outer membrane protein (MOMP) between H. somni strains 129Pt and 2336 has been described (Ueno et al. 2014). Since plasmid-based cloning of the H. somni gene encoding MOMP proved difficult, a vector-free strategy that utilizes the direct electroporation of PCR-amplified, HhaI-methylated linear DNA into H. somni was developed (Ueno et al. 2014). Following allelic exchange, strain 129Pt stably expresses the gene encoding MOMP from strain 2336 (HSM_1447, ompH/OmpH, 1443 bp/380 aa) and strain 2336 stably expresses the gene encoding MOMP from strain 129Pt (HS_0971, ompH/OmpH, 951 bp/316 aa), and the proteins can be detected by Western and dot blots using strain-specific anti-MOMP monoclonal antibodies. Furthermore, strains 129Pt and 2336 stably express a chimeric gene encoding MOMP (due to combining parts of genes encoding MOMPs from the two strains) after allelic exchange, and the fusion proteins can be detected using strain-specific anti-MOMP monoclonal antibodies in Western and dot blots (Ueno et al. 2014). The serum susceptibilities of strain 129Pt expressing HSM_1447 and strain 129Pt expressing the fusion protein (containing portions of HSM_1447 at the C-terminus) are significantly greater than those of the wild type (Ueno et al. 2014). This is not surprising since the genomes of strains 129Pt and 2336 differ from each other, and the genes encoding the OmpH homologs have only 56 % identity.
To overcome the inherent low efficiency of transformation and recombination of non-replicative suicide plasmids used for allelic exchange in H. somni, improved methods of mutagenesis need to be developed. Mutagenesis of H. somni using a commercially available transposon (Sandal et al. 2009) represents a significant step in this direction. Electroporation of H. somni strain 2336 yields up to 100 kanamycin-resistant colonies per 20 ng of the EZ-Tn5™ <KAN-2> Tnp Transposome™ (Epicentre, Madison, WI). Of 500 transposon mutants of H. somni strain 2336 screened for biofilm formation using the crystal violet assay, 55 formed either more or less biofilm than the wild-type strain. Of the several transposon mutants confirmed to produce less biofilm than the wild-type strain by scanning electron microscopy, six contained a transposon insertion in a region of the ibpA gene that encodes a putative filamentous hemagglutinin. This indicates that filamentous hemagglutinins, which are important attachment factors in other pathogenic bacteria [such as Bordetella (Villarino Romero et al. 2014)], likely contribute to H. somni biofilm formation and possibly pathogenesis (Sandal et al. 2009). Mutagenesis of H. somni strain 2336 genes putatively encoding S-ribosylhomocysteinase (luxS), universal stress protein E (uspE), major facilitator transport protein, and a protein of unknown function has also been achieved using the EZ-Tn5™ <KAN-2> Tnp Transposome™ (Sandal et al. 2009; Shah et al. 2014). Interestingly, both luxS and uspE mutants are attenuated in an acute septicemia mouse model, whereas only the uspE mutant is deficient in biofilm formation (Shah et al. 2014).
6 Natural Transformation
The ability of bacteria to internalize chromosomal fragments and/or plasmids under natural conditions is referred to as competence. Competence is proposed to be regulated by biochemical as well as environmental cues, and the purposes for internalizing DNA within the host cell could be non-genetic (e.g., nutrition) or genetic (e.g., transformation) (Mell and Redfield 2014). Although most naturally competent bacteria are indiscriminate in DNA internalization, members of the Pasteurellaceae and the Neisseriaceae are known to prefer conspecific DNA. The preferential internalization of conspecific DNA by members of these two families appears to be facilitated by short uptake signal sequences (Mell and Redfield 2014). In H. influenzae, the uptake signal sequence is a nonamer (5′-AAGTGCGGT, or its reverse complement), and comparative genomic analyses have demonstrated the abundance of this sequence in Actinobacillus actinomycetemcomitans, P. multocida, and H. somni (Bakkali et al. 2004; Redfield et al. 2006).
Although several members of the Pasteurellaceae are believed to be competent, only H. influenzae, A. actinomycetemcomitans, and A. pleuropneumoniae have been shown to undergo natural transformation under laboratory conditions (Redfield et al. 2006). In other species, the lack of competence or transformation in the laboratory is believed to be due to the failure to mimic native conditions and/or dysfunctional genetic systems (Redfield et al. 2006). Notably, H. somni strains 2336 and 129Pt lack a comD homolog and appear to encode a shortened ComE homolog. Since it has been demonstrated that functionality of each gene within the com operon is essential for transformation of H. influenzae (Carruthers et al. 2012), it could be presumed that H. somni strains 2336 and 129Pt are non-transformable. This appears to be valid in the case of strain 129Pt since it fails to be transformed when plasmid pNS3K (Sandal et al. 2008), genomic DNA from a lob2A mutant of strain 738 (Wu et al. 2000), or genomic DNA from transposon mutants of luxS or uspE of strain 2336 is used (Shah et al. 2014). However, strain 2336 can be transformed with a low efficiency when plasmid pNS3K or genomic DNA from a lob2A mutant of strain 738 is used. Nevertheless, strain 2336 fails to transform when genomic DNA from transposon mutants of luxS or uspE is used. Moreover, H. somni strains 649, 8025, and M14-622 also fail to transform when genomic DNA from a lob2A mutant of strain 738 is used (Shah et al. 2014). Therefore, it is likely that H. somni strains differ in their competency/transformability, and this could be due to the lack of specific com genes and/or the variety of restriction–modification systems that occur in this species (Briggs and Tatum 2005; Siddaramappa et al. 2011). Differences in competence and transformation have also been observed among strains of H. influenzae lacking specific com genes (Maughan and Redfield 2009). Furthermore, transformation of H. influenzae with restriction endonuclease-digested conspecific DNA is dependent on fragment size (Beattie et al. 1982), and it has been hypothesized that restriction endonucleases released by lysed cells may cut donor DNA fragments destined for uptake and reduce recombination efficiency (Mell and Redfield 2014).
7 Conclusions
Biochemical and genetic studies in the pre-genomic era firmly establish H. somni as a potent opportunistic pathogen. Complete genome sequencing reveals the pathogenic repertoire of this species, and comparative genomic analyses facilitate the identification of chromosomal regions that resemble the pathogenicity islands of other virulent bacteria. One such pathogenicity island has now been identified as ICEHso2336 and appears to represent a classical horizontally transferred element. Transcriptome analyses indicate that ~80 % of the predicted genes of H. somni strain 2336 are readily transcribed, and ~44 % of these genes are operonic. Furthermore, electrotransformation of H. somni appears to be more efficient than natural transformation. In addition, genetic manipulation of H. somni is achievable through either suicide plasmid-based homologous recombination (targeted mutagenesis) or transposomes (random mutagenesis), and several plasmids are now available that can serve as shuttle vectors. Future investigations of H. somni are expected to be guided by the principles, technologies, and developments discussed in this chapter.
References
Appuhamy S, Low JC, Coote JG, Parton R (1998) PCR methods and plasmid profile analysis for characterisation of Histophilus ovis strains. J Med Microbiol 47(11):987–992
Bakkali M, Chen TY, Lee HC, Redfield RJ (2004) Evolutionary stability of DNA uptake signal sequences in the Pasteurellaceae. Proc Natl Acad Sci USA 101(13):4513–4518. doi:10.1073/pnas.0306366101
Beattie KL, Wakil AE, Driggers PH (1982) Action of restriction endonucleases on transforming DNA of Haemophilus influenzae. J Bacteriol 152(1):332–337
Briggs RE, Tatum FM (2005) Generation and molecular characterization of new temperature-sensitive plasmids intended for genetic engineering of Pasteurellaceae. Appl Environ Microbiol 71(11):7187–7195. doi:10.1128/AEM.71.11.7187-7195.2005
Carruthers MD, Tracy EN, Dickson AC, Ganser KB, Munson RS Jr, Bakaletz LO (2012) Biological roles of nontypeable Haemophilus influenzae type IV pilus proteins encoded by the pil and com operons. J Bacteriol 194(8):1927–1933. doi:10.1128/JB.06540-11
Challacombe JF, Inzana TJ (2008) Comparative genomics of Pasteurellaceae. In: Kuhnert P, Christensen H (eds) Pasteurellaceae: biology, genomics, and molecular aspects. Caister Academic Press, Norfolk, pp 53–77
Challacombe JF, Duncan AJ, Brettin TS, Bruce D, Chertkov O, Detter JC, Han CS, Misra M, Richardson P, Tapia R, Thayer N, Xie G, Inzana TJ (2007) Complete genome sequence of Haemophilus somnus (Histophilus somni) strain 129Pt and comparison to Haemophilus ducreyi 35000HP and Haemophilus influenzae Rd. J Bacteriol 189(5):1890–1898
Corbeil L, Gogolewski R, Stephens L, Inzana T (1995) Haemophilus somnus: antigen analysis and immune responses. In: Donachie W, Lainson F, Hodgson J (eds) Haemophilus, Actinobacillus, and Pasteurella. Plenum Press, New York, pp 63–73
Corbeil LB, Bastida-Corcuera FD, Beveridge TJ (1997) Haemophilus somnus immunoglobulin binding proteins and surface fibrils. Infect Immun 65(10):4250–4257
D’Amours GH, Ward TI, Mulvey MR, Read RR, Morck DW (2011) Genetic diversity and tetracycline resistance genes of Histophilus somni. Vet Microbiol 150(3–4):362–372. doi:10.1016/j.vetmic.2011.02.051
Eidam C, Poehlein A, Leimbach A, Michael GB, Kadlec K, Liesegang H, Daniel R, Sweeney MT, Murray RW, Watts JL, Schwarz S (2015) Analysis and comparative genomics of ICEMh1, a novel integrative and conjugative element (ICE) of Mannheimia haemolytica. J Antimicrob Chemother 70(1):93–97. doi:10.1093/jac/dku361
Fleischmann RD, Adams MD, White O, Clayton RA, Kirkness EF, Kerlavage AR, Bult CJ, Tomb JF, Dougherty BA, Merrick JM et al (1995) Whole-genome random sequencing and assembly of Haemophilus influenzae Rd. Science 269(5223):496–512
Fussing V, Wegener HC (1993) Characterization of bovine Haemophilus somnus by biotyping, plasmid profiling, REA-patterns and ribotyping. Zentralblatt fur Bakteriologie: Int J Med Microbiol 279(1):60–74
Hoshinoo K, Sasaki K, Tanaka A, Corbeil LB, Tagawa Y (2009) Virulence attributes of Histophilus somni with a deletion mutation in the ibpA gene. Microb Pathog 46(5):273–282 doi:S0882-4010(09)00033-3 [pii]
Inzana TJ, Gogolewski RP, Corbeil LB (1992) Phenotypic phase variation in Haemophilus somnus lipooligosaccharide during bovine pneumonia and after in vitro passage. Infect Immun 60(7):2943–2951
Inzana TJ, Todd J, Veit HP (1993) Safety, stability, and efficacy of noncapsulated mutants of Actinobacillus pleuropneumoniae for use in live vaccines. Infect Immun 61(5):1682–1686
Inzana TJ, Glindemann G, Cox AD, Wakarchuk W, Howard MD (2002) Incorporation of N-acetylneuraminic acid into Haemophilus somnus lipooligosaccharide (LOS): enhancement of resistance to serum and reduction of LOS antibody binding. Infect Immun 70(9):4870–4879
Izadpanah R, Dan A, Benko M, Rusvai M, Fodor L, Harrach B (2001) DNA sequence of a small, unidentified plasmid isolated from a Haemophilus somnus strain: short communication. Acta Vet Hung 49(1):11–16
Klima CL, Zaheer R, Cook SR, Booker CW, Hendrick S, Alexander TW, McAllister TA (2014) Pathogens of bovine respiratory disease in North American feedlots conferring multidrug resistance via integrative conjugative elements. J Clin Microbiol 52(2):438–448. doi:10.1128/JCM.02485-13
Kumar R, Lawrence ML, Watt J, Cooksey AM, Burgess SC, Nanduri B (2012) RNA-seq based transcriptional map of bovine respiratory disease pathogen “Histophilus somni 2336”. PLoS ONE 7(1):e29435. doi:10.1371/journal.pone.0029435
Maughan H, Redfield RJ (2009) Extensive variation in natural competence in Haemophilus influenzae. Evol: Int J Organ Evol 63(7):1852–1866. doi:10.111/j.1558-5646.2009.00658.x
McQuiston JH, McQuiston JR, Cox AD, Wu Y, Boyle SM, Inzana TJ (2000) Characterization of a DNA region containing 5′-(CAAT)(n)-3’ DNA sequences involved in lipooligosaccharide biosynthesis in Haemophilus somnus. Microb Pathog 28(5):301–312. doi:10.1006/mpat.1999.0351
Mell JC, Redfield RJ (2014) Natural competence and the evolution of DNA uptake specificity. J Bacteriol 196(8):1471–1483. doi:10.1128/JB.01293-13
Michael GB, Kadlec K, Sweeney MT, Brzuszkiewicz E, Liesegang H, Daniel R, Murray RW, Watts JL, Schwarz S (2012) ICEPmu1, an integrative conjugative element (ICE) of Pasteurella multocida: structure and transfer. J Antimicrob Chemother 67(1):91–100. doi:10.1093/jac/dkr411
Mohd-Zain Z, Turner SL, Cerdeno-Tarraga AM, Lilley AK, Inzana TJ, Duncan AJ, Harding RM, Hood DW, Peto TE, Crook DW (2004) Transferable antibiotic resistance elements in Haemophilus influenzae share a common evolutionary origin with a diverse family of syntenic genomic islands. J Bacteriol 186(23):8114–8122
Redfield RJ, Findlay WA, Bosse J, Kroll JS, Cameron AD, Nash JH (2006) Evolution of competence and DNA uptake specificity in the Pasteurellaceae. BMC Evol Biol 6:82. doi:10.1186/1471-2148-6-82
Sandal I, Seleem MN, Elswaifi SF, Sriranganathan N, Inzana TJ (2008) Construction of a high-efficiency shuttle vector for Histophilus somni. J Microbiol Methods 74(2–3):106–109. doi:10.1016/j.mimet.2008.04.002
Sandal I, Shao JQ, Annadata S, Apicella MA, Boye M, Jensen TK, Saunders GK, Inzana TJ (2009) Histophilus somni biofilm formation in cardiopulmonary tissue of the bovine host following respiratory challenge. Microb Infect/Institut Pasteur 11(2):254–263. doi:10.1016/j.micinf.2008.11.011
Sanders JD, Tagawa Y, Briggs RE, Corbeil LB (1997) Transformation of a virulence associated gene of Haemophilus somnus into a strain lacking the gene. FEMS Microbiol Lett 154(2):251–258
Sanders JD, Bastida-Corcuera FD, Arnold KF, Wunderlich AC, Corbeil LB (2003) Genetic manipulation of immunoglobulin binding proteins of Haemophilus somnus. Microb Pathog 34(3):131–139
Shah N, Bandara AB, Sandal I, Inzana TJ (2014) Natural competence in Histophilus somni strain 2336. Vet Microbiol 173(3–4):371–378. doi:10.1016/j.vetmic.2014.07.025
Siddaramappa S, Duncan AJ, Brettin T, Inzana TJ (2006) Comparative analyses of two cryptic plasmids from Haemophilus somnus (Histophilus somni). Plasmid 55(3):227–234. doi:10.1016/j.plasmid.2005.11.004
Siddaramappa S, Challacombe JF, Duncan AJ, Gillaspy AF, Carson M, Gipson J, Orvis J, Zaitshik J, Barnes G, Bruce D, Chertkov O, Detter JC, Han CS, Tapia R, Thompson LS, Dyer DW, Inzana TJ (2011) Horizontal gene transfer in Histophilus somni and its role in the evolution of pathogenic strain 2336, as determined by comparative genomic analyses. BMC Genom 12:570. doi:10.1186/1471-2164-12-570
Tatum FM, Briggs RE (2005) Construction of in-frame aroA deletion mutants of Mannheimia haemolytica, Pasteurella multocida, and Haemophilus somnus by using a new temperature-sensitive plasmid. Appl Environ Microbiol 71(11):7196–7202. doi:10.1128/AEM.71.11.7196-7202.2005
Ueno Y, Hoshinoo K, Tagawa Y (2014) Mutations in the major outer membrane protein gene from Histophilus somni by an allelic exchange method. J Microbiol Methods 106:83–92. doi:10.1016/j.mimet.2014.08.008
Villarino Romero R, Osicka R, Sebo P (2014) Filamentous hemagglutinin of Bordetella pertussis: a key adhesin with immunomodulatory properties? Future Microbiol 9(12):1339–1360. doi:10.2217/fmb.14.77
Williams BJ, Golomb M, Phillips T, Brownlee J, Olson MV, Smith AL (2002) Bacteriophage HP2 of Haemophilus influenzae. J Bacteriol 184(24):6893–6905
Wood GE, Dutro SM, Totten PA (1999) Target cell range of Haemophilus ducreyi hemolysin and its involvement in invasion of human epithelial cells. Infect Immun 67(8):3740–3749
Wu Y, McQuiston JH, Cox A, Pack TD, Inzana TJ (2000) Molecular cloning and mutagenesis of a DNA locus involved in lipooligosaccharide biosynthesis in Haemophilus somnus. Infect Immun 68(1):310–319
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2015 Springer International Publishing Switzerland
About this chapter
Cite this chapter
Siddaramappa, S. (2015). Histophilus somni Genomics and Genetics. In: Inzana, T. (eds) Histophilus somni. Current Topics in Microbiology and Immunology, vol 396. Springer, Cham. https://doi.org/10.1007/82_2015_5009
Download citation
DOI: https://doi.org/10.1007/82_2015_5009
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-319-29554-1
Online ISBN: 978-3-319-29556-5
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)