Abstract
Post-traumatic stress disorder (PTSD) is a psychiatric disorder that can develop following exposure to or witnessing of a (potentially) threatening event. A critical issue is to pinpoint the (neuro)biological mechanisms underlying the susceptibility to stress-related disorder such as PTSD, which develops in the minority of ~15% of individuals exposed to trauma. Over the last few years, a first wave of epigenetic studies has been performed in an attempt to identify the molecular underpinnings of the long-lasting behavioral and mental effects of trauma exposure. The potential roles of non-coding RNAs (ncRNAs) such as microRNAs (miRNAs) in moderating or mediating the impact of severe stress and trauma are increasingly gaining attention. To date, most studies focusing on the roles of miRNAs in PTSD have, however, been completed in animals, using cross-sectional study designs and focusing almost exclusively on subjects with susceptible phenotypes. Therefore, there is a strong need for new research comprising translational and cross-species approaches that use longitudinal designs for studying trajectories of change contrasting susceptible and resilient subjects. The present review offers a comprehensive overview of available studies of miRNAs in PTSD and discusses the current challenges, pitfalls, and future perspectives of this field.
Clara Snijders and Laurence de Nijs contributed equally to this work.
Access provided by CONRICYT-eBooks. Download chapter PDF
Similar content being viewed by others
Keywords
1 Introduction
Over the last few decades, epigenetic mechanisms have been proposed to be key mediators of the lasting behavioral and molecular effects of traumatic stress exposure (Schmidt et al. 2011). While a first wave of epigenetic studies in this area focused mostly on DNA methylation, epigenetic studies in more recent years have expanded this approach by analyzing the expression of non-coding RNA (ncRNA) species and their impact on gene expression. These RNA molecules of different sizes and forms include non-coding stretches of 20–25 nucleotides named microRNAs (miRNAs). These are increasingly being investigated for their pathophysiological connection to psychiatric disorders including post-traumatic stress disorder (PTSD). More recently, studies have started to focus on the potential use of miRNAs as biomarkers of PTSD.
The present review provides an overview of the current status of the literature on miRNAs in relation to exposure to traumatic stress and its impact on mental health in humans and other mammals. To do so, we briefly describe PTSD-related neurobiological alterations along with the basic concepts of epigenetic mechanisms. Next, an overview of the current scientific evidence on miRNAs in relation to PTSD in humans and PTSD-related symptoms in animals is provided. Finally, current challenges, pitfalls, and future perspectives in studying the potential role of miRNAs in PTSD are discussed.
2 Post-traumatic Stress Disorder
As we know, PTSD is a psychiatric disorder that is triggered by a (potentially) life-threatening traumatic event, i.e., an event capable of producing intense feelings of fear, helplessness, and horror (American Psychiatric Association 2013). Characteristic symptoms include re-experiencing of the traumatic event through intrusive imagery or recurrent nightmares, constant avoidance of reminders of the event, negative mood, and hyperarousal reflected by insomnia and/or hypervigilance. Although these symptoms are often of limited intensity and duration, in a small, susceptible minority of the population they persist longer than 1 month following trauma exposure and create significant distress. Long-term persistence of symptoms is characteristic of PTSD, while the ability to withstand trauma without developing any stress symptoms or rapid recovery from an acute stress reaction without progression to PTSD is referred to as resiliency.
Over the past few decades, PTSD has repeatedly been associated with several neurobiological alterations including decreased hippocampal volume (Smith 2005; Karl et al. 2006; Shin et al. 2006), hyperactivity of the amygdala and hypoactivity of the dorsal and rostral anterior cingulate (AC) cortices and ventromedial prefrontal cortex (vmPFC) (Shin et al. 2006; Etkin and Wager 2007; El Khoury-Malhame et al. 2011). In an attempt to further elucidate the (neuro)biological processes underlying the observed differential susceptibility to traumatic stress, a large number of studies have focused on alterations in the hypothalamus-pituitary-adrenal (HPA) axis. Since the HPA axis is a core component of the mammalian stress response, its (dys)function has been extensively studied in the context of PTSD. In healthy individuals, stressful events trigger neurons of the hypothalamic paraventricular nucleus (PVN) to secrete corticotropin-releasing hormone (CRH) and vasopressin, which causes the release of adrenocorticotropin (ACTH) from the anterior pituitary and finally glucocorticoids from the adrenal cortex (Chrousos and Gold 1992). The activity of the HPA axis is modulated via several brain regions; for example, CRH neurons in the PVN are inhibited by the hippocampus and PFC and stimulated by areas such as the amygdala (Sherin and Nemeroff 2011). Finally, in order to regulate their own synthesis, glucocorticoids inhibit excessive synthesis and release of CRH and ACTH by controlling hippocampal and PVN neurons, and downregulating CRH1 receptors and corticotrope function in the anterior pituitary, thereby creating a negative feedback mechanism (Sherin and Charles 2011).
Several studies have found that subjects with PTSD show increased levels of CRH in cerebrospinal fluid (CSF) (Baker et al. 1999), as well as a blunted ACTH response to CRH (Yehuda 2006), a disturbed negative feedback loop (Geracioti et al. 2008), and increased sensitivity of glucocorticoid receptors (GRs) and chronically lowered cortisol levels (Yehuda 2001; Yehuda et al. 2000). Although dysregulation of the HPA axis is well-documented in the context of stress-related disorders and PTSD has repeatedly been associated with reduced cortisol levels, variability in response between individuals remains. The current hypothesis is that cortisol levels depend upon gender and the type of trauma exposure among other factors (Meewisse et al. 2007; Young and Breslau 2004; Lemieux and Coe 1995). To further unravel the molecular regulation of biological mechanisms underlying the onset and course of PTSD, more recent research has also focused on the involvement of epigenetic mechanisms.
3 Epigenetics: The Role of miRNAs
The term epigenetics refers to a variety of heritable but reversible processes involved in the regulation of gene expression under influence of environmental factors without the original genetic code being altered (Peschansky and Wahlestedt 2014). These epigenetic modifications are numerous and include (hydroxy)methylation of DNA cytosine residues, post-translational modifications (PTMs) of histone proteins and ncRNAs (Kouzarides 2007; Venkatesh and Workman 2015). ncRNAs refer to a class of small RNA molecules that are transcribed from genomic DNA without being translated into proteins (Peschansky and Wahlestedt 2014). Instead, these RNAs are directly involved in cellular function and gene regulation. Next to ribosomal and transfer RNAs, ncRNAs include the most commonly studied small interfering RNAs (Zamore 2002), circular RNAs (Memczak et al. 2013), piwi-interacting RNAs (Aravin et al. 2007), and miRNAs.
3.1 Biogenesis and Mode of Action of miRNAs
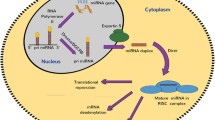
miRNAs are small (~22 nt in length) ncRNA molecules found in most eukaryotes (Fabian and Sonenberg 2012). Hundreds of different miRNAs are expressed within an organism and are involved in post-transcriptional regulation of gene expression (Pritchard et al. 2012). miRNAs are commonly classified as “intergenic” or “intronic.” Intergenic miRNA are transcribed from genomic DNA by RNA polymerase II and/or III (Borchert et al. 2006) and intronic miRNA are processed from intronic regions of heterogeneous nuclear RNA (hnRNA) (Ramalingam et al. 2014). In both cases, a primary miRNA (pri-miRNA) is formed and further cleaved and stabilized by the protein complex microprocessor that includes the ribonuclease III Drosha and its co-factor, DiGeorge syndrome critical region 8 (DGCR8) (Borchert et al. 2006). This process takes place within the nucleus and results in a precursor miRNA (pre-miRNA) of 70–100 nt in length forming a hairpin structure (Issler and Chen 2015; Lee et al. 2003). Following transport to the cytoplasm by the nuclear transport factor Exportin-5, a complex including the RNase III Dicer further processes the pre-miRNA to yield a miRNA duplex containing the final mature miRNA strand and a so-called passenger strand (Fig. 1) (Davis-Dusenbery and Hata 2010).
miRNA biogenesis and cellular locations. miRNAs are transcribed into pri-miRNA by RNA polymerase II and/or III before being further processed by Drosha and DGCR8 to form a cleaved pre-miRNA. After transportation to the cytoplasm by Exportin-5, this pre-miRNA is further digested by a complex including the RNase III Dicer. The mature miRNA is then involved in translational repression and/or mRNA degradation through interaction with the RISC. In the extracellular space, miRNAs are protected from degradation by RNases through binding to RNA-binding proteins (e.g., Ago 1 or 2) or (high-density) lipoproteins, or packaging into exosomes or microvesicles
Binding of the 5′ end of the mature miRNA (i.e., the “seed” sequence) to an almost complementary 6–8 nt seed match sequence in the 3′ UTR of mRNA induces mRNA degradation or translational inhibition (Pritchard et al. 2012; Davis-Dusenbery and Hata 2010). Thus, miRNAs hold the potential to post-transcriptionally regulate gene expression. Specifically, the mature miRNA triggers the activation of the RNA-induced silencing complex (RISC), a large protein complex containing an Argonaute protein (Ago2) needed for gene silencing, and the mature single-stranded miRNA that leads the complex towards the appropriate mRNA target (Fig. 1) (Fabian and Sonenberg 2012). It was commonly believed that, at this point, only the functional guide strand of the double-stranded miRNA product was incorporated into the RISC and the passenger strand was being degraded (Issler and Chen 2015). However, increasing evidence shows that the passenger strand also has biological functions and target mRNAs (Yang et al. 2013). In either case, depending on the type of Ago protein, the target will be cleaved directly or additional proteins may be needed to achieve silencing. However, exactly how this complex interacts with mRNA strands and which additional proteins are recruited remains unclear.
Currently, it is believed that miRNAs regulate 30–60% of human protein-coding genes (Friedman et al. 2009; Lewis et al. 2005). Several studies have investigated genetic variations such as single nucleotide polymorphisms (SNPs) in the 3′ UTRs of mRNAs (Hanin et al. 2014; Jin and Lee 2013). Since base-pair matching between miRNAs and mRNAs relies on imprecise complementarity, one single miRNA can target many different mRNAs. Therefore, genetic variations in one miRNA target can cause a wide variety of molecular and behavioral effects due to the potential of one miRNA to bind multiple targets.
3.2 miRNAs in the Nervous System
miRNAs are widely expressed within the central nervous system (CNS) and are suggested to be crucially involved in its development (Smith et al. 2010). Studies have demonstrated that several miRNAs are implicated in the proliferation and differentiation of neural stem cells (NSCs) (Bian et al. 2013), dendritic development (Magill et al. 2010), axon outgrowth and branching (Dajas-Bailador et al. 2012), and synaptic plasticity (Aksoy-Aksel et al. 2014; Hu and Li 2017). Given their central involvement in neural development and function, CNS miRNA dysregulations have been identified in several neuropsychiatric and neurodegenerative disorders such as major depressive disorder (MDD) (Smalheiser et al. 2012; Bai et al. 2012), Alzheimer’s disease (AD) (Absalon et al. 2013; Hu et al. 2013), and Parkinson’s disease (PD) (Wang et al. 2008; Doxakis 2010). Identifying exactly which and how miRNAs within the CNS interact to exert their regulatory effects will be crucial for our understanding of their precise involvement in these and other neurological disorders.
3.3 Circulating miRNAs
While most miRNAs are found inside the cells, a significant number of miRNAs have been observed in extracellular compartments such as biofluids, including blood plasma, serum, saliva, urine, tears, and CSF (Park et al. 2009; Taylor and Gercel-Taylor 2013; Hanke et al. 2010; Weber et al. 2010). These extracellular miRNAs are relatively stable since they are commonly bound to proteins such as Ago1 or 2 and (mostly high density) lipoproteins or packed into vesicles and thus protected from degradation by RNases (Fig. 1) (Taylor and Gercel-Taylor 2013; Turchinovich et al. 2013; Camussi et al. 2011; Valadi et al. 2007; Mitchell et al. 2008; Vickers et al. 2011; Wagner et al. 2013).
Packaging of miRNAs is the most common mechanism used to protect circulating miRNAs. miRNAs can be packaged into apoptotic bodies, shedding vesicles called microvesicles, or exosomes resulting from multivesicular bodies (MVBs) fusing with the plasma membrane (Taylor and Gercel-Taylor 2013; Turchinovich et al. 2013). miRNAs encapsulated within MVBs are believed to arise from the disassembled RISC and are packed along with several RISC-associated components (Gibbings et al. 2009). Once secreted, exosomes translocate easily across cell membranes, thus allowing miRNAs to be taken up by other cells where they hold the potential to actively alter gene expression, among other functions (Wang et al. 2010). Although packaged miRNAs are thought to be specifically involved in RNA-mediated cell-to-cell communication, Ago-bound miRNAs appear to be non-specific residues of cellular activity or cell death (Turchinovich et al. 2013). Indeed, Ago-miRNA complexes have not been found to be actively released or taken up by recipient cells, unlike exosomal miRNAs (Turchinovich et al. 2013). Although several theories have been postulated with regard to extracellular miRNA origin, stability and precise function in recipient cells, many questions remain to be answered. However, circulating miRNAs have several properties that make them interesting relevant candidates to be investigated as biomarkers; they are stable in various biofluids, their sequences are conserved among different species, the expression of some miRNAs is specific to tissues or biological stages, and the level of miRNAs can be easily assessed by various methods, such as small-RNA sequencing, microarrays and quantitative polymerase chain reaction (PCR) (Etheridge et al. 2011). As such, circulating miRNAs in biofluids may reflect miRNA expression and/or dysfunction in the brain.
3.4 Mechanism of miRNA Regulation
In the past few years it has become clear that miRNA expression is regulated by DNA methylation and histone modifications and vice versa (Satrom et al. 2007). Several proteins of the methyl-CpG-binding domain (MBDs) family, i.e. proteins binding to methylated DNA cytosine residues, directly influence miRNA expression (Liu et al. 2010; Chen et al. 2012). Moreover, disturbed methylation patterns arising in promoter regions of miRNA genes have been linked to several human diseases, including neurodegenerative disorders (reviewed in (Van den Hove et al. 2014) for AD). Similarly, histone modifiers have not only been shown to interact with DNA methyltransferases (Dnmts), enzymes involved in maintaining or establishing de novo DNA methylation patterns (Rose and Klose 2014; Raabe and Spengler 2013), but are also suggested to affect miRNA expression levels (Scott et al. 2006). Interestingly, miRNAs themselves have been shown to target histone modifier molecules involved in histone PTM and Dnmt1, 3a, and 3b (Fabbri et al. 2007) through a process termed RNA-directed DNA methylation (Sato et al. 2011). For instance, Dicer-null mouse embryonic stem cells have been shown to express significantly lower levels of Dnmt1, Dnmt3a, and Dnmt3b, further resulting in altered DNA methylation patterns (Sinkkonen et al. 2008). The presence of such epigenetic feedback loops highlights the complex interaction between miRNAs and other epigenetic mechanisms (Schouten et al. 2013).
4 miRNAs, Stress and PTSD
The results of studies examining miRNAs in the context of stress and PTSD in humans or PTSD-related symptoms in animals are described below and summarized in Tables 1 and 2, respectively.
4.1 Evidence from Animal Studies
4.1.1 miRNAs and Fear Conditioning
Patients with PTSD are known to show enhanced fear conditioning and to benefit from exposure-based therapy (Blechert et al. 2007). This therapy is very similar to the fear extinction training used in animals (Norberg et al. 2008). Therefore, the first study to indirectly examine the role of miRNAs in PTSD focused on their involvement in fear extinction (Lin et al. 2011). In this study, the level of miR-128b was increased in the infra-limbic PFC (ILPFC) of mice following fear extinction training, implicating its involvement in fear conditioning (Lin et al. 2011). Previously, proteins involved in miRNA biogenesis had already been shown to play a role in memory formation. Indeed, the deletion of Dicer1 in the forebrain of mice caused a decrease in several miRNAs and enhanced learning and memory strength (Konopka et al. 2010). Several recent animal studies have confirmed that specific miRNAs in several brain regions are involved in fear memory consolidation (Dias et al. 2014; Griggs et al. 2013), contextual fear memory (Vetere et al. 2014), state-dependent fear (Jovasevic et al. 2015), and memory acquisition of trace fear conditioning (Wang et al. 2013).
4.1.2 Circulating miRNAs as Biomarkers of PTSD
Over the past few years, fluctuations of miRNA levels in body fluids have been shown to correlate with psychiatric disorders, including MDD (Bocchio-Chiavetto et al. 2013), schizophrenia (Lai et al. 2011), and bipolar disorder (Rong et al., n.d.). These studies suggest potential for the use of circulating miRNAs as diagnostic biomarkers of mental disorders. The first study investigating circulating miRNAs as biomarkers of PTSD-related symptoms found that the expression of nine miRNAs was increased both in the amygdala and serum of rats exposed to 3 days of immobilization and tail shock sessions (Balakathiresan et al. 2014). One of the increased stress-responsive miRNAs, miR-19b, was also found to be involved in the regulation of fear-associated genes. A third lead for miR-19b involvement comes from a study using mice undergoing chronic social defeat stress (CSDS) that reported significant increases in the basolateral amygdala (BLA) following CSDS as compared to non-stressed controls (Volk et al. 2014). Finally, miR-19b was also found associated with Ago2 and to target the amygdalar Adrenergic Receptor Beta 1 (Adrb1).
More recently, the potential of miRNAs to be used as biomarkers of both vulnerability and resilience to stress was examined. In one study, circulating miRNA profiles were examined 3 days before and 24 h following CSDS in rats (Chen et al. 2015). Prior to the stressful event, four miRNAs (miR-4-2-5p, miR-27a-3p, miR-30e-5p, miR-362-3p) were significantly decreased only in those rats that later became vulnerable to stress. Following stress exposure, four different miRNAs (miR-139-5p, miR-28-3p, miR-326-3p, miR-99b-5p) were decreased in resilient animals. These results show that different miRNAs potentially confer vulnerability to future stress or promote sustained resilience. Taken together, these studies show promise for using miRNAs as biomarkers of vulnerability and resiliency to stress.
4.1.3 miRNAs in Transgenerational Inheritance of Early Stress
Several animal studies have shown that ncRNAs are abundantly present in sperm and may be involved in non-Mendelian inheritance of behavioral phenotypes (Rassoulzadegan et al. 2006; Liu et al. 2012). Therefore, to assess the potential role of miRNAs in the transgenerational inheritance of parental stress, Gapp et al. (2014) examined sperm samples of a mouse model of unpredictable maternal separation with unpredictable maternal stress (MSUS). Several miRNAs (among other ncRNAs) were upregulated in F1 MSUS sperm (but not F2 sperm) as compared to the sperm of non-stressed control mice. Several miRNA levels were further altered in serum, hippocampus and hypothalamus of F1 MSUS mice, and in serum and hippocampus of F2 MSUS mice. Interestingly, following injection of RNAs purified from MSUS male sperm into wild-type fertilized mouse oocytes, similar behavioral, metabolic, and molecular effects were obtained as compared to direct exposure to MSUS. Additionally, the offspring of these mice showed depressive-like behaviors. These and other results (Rodgers et al. 2013) provide support for the involvement of RNAs, including miRNAs, in the transgenerational transmission of behavioral phenotypes.
4.1.4 miRNAs Targets the FK506 Binding Protein 5 (FKBP5) Gene
The only stress-related gene that has been suggested to be regulated by miRNAs is FKBP5. Genetic variations in FKBP5 have been extensively studied in the context of gene x environment (GxE) interactions and the influence of early life adversity with regard to PTSD (Binder et al. 2004, 2008; Mehta et al. 2011). The immunophilin FKBP5 is a HSP90 co-chaperone that strongly controls glucocorticoid receptor (GR) sensitivity and signaling by binding to GRs in the cytosol thereby decreasing GR ligand affinity and nuclear translocation (Zannas et al. 2015). Several studies have shown that homozygous genotypes for SNPs in FKBP5 interact with early life (but not adult) adversity, increasing the risk for later development of PTSD (Binder et al. 2008; Zimmermann et al. 2011). Epigenetic mechanisms have repeatedly been found to contribute to the regulation of FKBP5 expression (Klengel et al. 2013; Yehuda et al. 2016). Moreover, FKBP51, one of the proteins encoded by FKBP5, presents an interesting target for the treatment of stress-related disorders. Increased levels of FKBP51 have been suggested to increase the risk of MDD and PTSD and the deletion of FKBP5 has been shown to prevent age-related depression-like phenotypes (Sabbagh et al. 2014). However, pharmacologically targeting FKBP51 has proven to be challenging due to the strong sequence similarity between this and other FKBP proteins (Schmidt et al. 2012). Recently, two independent studies have shown that miR-15a and miR-511 affect FKBP51 levels by targeting FKBP5 mRNAs (Zheng et al. 2016; Volk et al. 2016). In the first study, FKBP51 levels were found to be decreased and miR-15a levels significantly increased in the amygdala of mice subjected to CSDS as compared to non-stressed controls (Volk et al. 2016). This same pattern was found in peripheral blood of healthy humans following dexamethasone treatment and in individuals exposed to early life trauma (Volk et al. 2016). In the second study, FKBP5 mRNA and protein levels were found to be decreased by miR-511, which was further shown to be involved in neuronal differentiation (Zheng et al. 2016). These findings indicate that both miRNAs are interesting potential candidates for the treatment of stress-related disorders and set the foundations for further studies to examine the exact roles of both miRNAs in FKBP5 regulation.
4.2 Evidence from Clinical Studies
Apart from Volk et al. (2016) examining miR-15a profiles in blood samples from patients exposed to childhood trauma and healthy individuals administered with dexamethasone, most human studies researching the link between miRNAs and PTSD so far have focused on miRNAs in relation to immunological dysregulations.
Immune dysfunctions are well documented in PTSD and have been reviewed recently (Neigh and Ali 2016; Michopoulos et al. 2017). PTSD has repeatedly been linked to an excessive inflammatory state, possibly resulting from insufficient counter regulation of PTSD-induced immune activation due to cortisol hyposecretion (Gill et al. 2009; Daskalakis et al. 2016). The first study examining peripheral blood mononuclear cells (PBMCs) of combat veterans diagnosed with PTSD found that alterations in specific miRNAs correlated with immunological changes (Zhou et al. 2014). Specifically, miR-125a and miR-181c were significantly decreased in PTSD patients as compared to healthy controls. Further analyses revealed that miR-125a targeted IFN-γ and downregulated the production of the pro-inflammatory cytokine IFN-γ. Therefore, the observed increase in IFN-γ in PBMCs of PTSD patients appears to be, at least in part, epigenetically regulated. Intriguingly, miR-27a-3p, which was downregulated in the circulation of rats vulnerable to future stress (Chen et al. 2015), and miR-19b (Balakathiresan et al. 2014; Volk et al. 2014) and miR-223 (Balakathiresan et al. 2014), which were increased in the serum and the amygdala of stressed rodents, were also dysregulated in the present cohort of combat veterans with PTSD (Zhou et al. 2014). However, it is worth mentioning that, while two independent animal studies found miR-19b levels to be increased in several tissues following stress exposure (Balakathiresan et al. 2014; Volk et al. 2014), one study reported increased levels of miR-223 (Balakathiresan et al. 2014), the same miRNAs were significantly decreased in PBMCs of the human cohort (Zhou et al. 2014).
Following this initial study linking miRNAs and immune dysfunctions in PTSD, two recent studies by the same research group provide further evidence for the epigenetic regulation of inflammation in PTSD (Bam et al. 2016a, b). In addition to IFN-γ, the pro-inflammatory cytokine IL-12 was increased in the same cohort of combat veterans, and miR-193a-5p, which was suggested to target IL-12B, was downregulated (Bam et al. 2016a). These results further suggest that pro-inflammatory gene expression is regulated by miRNAs.
Recently, one study found 8 miRNAs to be differentially expressed (4 upregulated and 4 downregulated) in peripheral blood samples of returning combat veterans as compared to controls (Martin et al. 2017). Pathway analyses revealed that these miRNAs target genes involved in Wnt signaling and axon guidance. However, being limited by a small sample size, this study encourages larger studies to further unravel the involvement of miRNAs in PTSD vulnerability.
5 Current Challenges, Pitfalls, and Future Perspectives
As reflected by the present overview, most studies to date that examine the role of miRNAs in PTSD have used (almost exclusively male) animals. Human studies of this subject are now beginning to emerge and have so far only examined peripheral blood samples. Moreover, most studies have included animals or humans, rarely both, and have focused on susceptible phenotypes only, i.e., those animals and individuals suffering the consequences of trauma exposure. To the best of our knowledge, only one study has examined the potential of miRNAs as biomarkers of both vulnerability and resiliency (Chen et al. 2015). Furthermore, a major limitation of current epigenetic research is the lack of longitudinal studies that would enable identification of dynamic epigenetic changes over time (Fig. 2). For these reasons, future research is critically needed to overcome a few pressing issues.
Current research and future perspectives with regard to miRNA analyses in relation to PTSD. Green and red silhouettes represent mental health or illness, respectively. The lightning bolt represents a stressful event (e.g., CSDS for rodents, combat trauma in humans) and the blood drop represents PBMCs as well as serum and plasma analyses. The Eppendorf tube represents a CSF sample
First, given the tissue specificity of epigenetic alterations and the evident inability to study the brains of living human beings, there is a strong need for researchers to incorporate human postmortem brain analyses in their study design. This approach could not only yield additional information with regard to location and quantity of miRNAs but also shed light on the extent to which blood-based miRNA results are informative for the CNS. In this context, it is becoming clear that focusing on exosome-associated biomarkers provides interesting insights into the brain. Exosomes are secreted membrane vesicles, derived from intracellular endosomes that are generated by the endocytic pathway. The exosomal process traffics damaged or excess proteins and miRNAs as cargo from the cytosol of neurons to the extracellular space where the exosomes can be transported from the CNS to the peripheral circulation. Since exosomes are capable of crossing the blood-brain barrier, when secreted from neural cells, they can be accessed through the bloodstream and further isolated and enriched for neural origin using neural-specific membrane markers (Kapogiannis et al. 2015; Goetzl et al. 2015). Recent studies have shown that Aβ42 levels in blood exosomes, presumably derived from neurons, were abnormally higher in subjects with mild cognitive impairment (MCI), MCI that progressed to dementia, and AD (Winston et al. 2016). In another study, blood exosomal levels of Aβ42 and tau phosphorylated at Thr181 and Ser396 predicted development of AD 10 years before clinical onset (Fiandaca et al. 2015). Exosomal plasma Aβ42 also correlated with CSF levels of phosphorylated tau (Winston et al. 2016). Therefore CNS-derived blood-based exosomes are extremely interesting biomarker candidates. Following on from this suggestion, one could imagine the use of CSF to reflect the neural environment more directly. Although more invasive, the collection and analyses of CSF-associated exosomes, which are currently understudied, could provide additional and valuable insights into the brain’s pathological processes. Similarly, examining several body fluids jointly, including plasma, serum, PBMCs, sperm and CSF, could further deepen our understanding of miRNA distribution and overlap. Finally, the use of longitudinal designs could yield valuable information regarding dynamic changes over time and how these changes potentially relate to differential susceptibility to traumatic stress.
It is worth noting that guidelines such as the prospective-specimen-collection, retrospective-blinded-evaluation (PRoBE) design (Pepe et al. 2008) or the Strengthening the Reporting of Observational studies in Epidemiology for Molecular Epidemiology (STROBE-ME) (Gallo et al. 2011) offer valuable overviews to help researchers in the design, execution, and reporting of biomarker studies. With respect to analyzing miRNAs in particular, Nair et al. (2014) recently provided a comprehensive overview of helpful study requirements for researchers involved in studying miRNAs in human diseases. Importantly, both human and animal studies have shown that differences in genetic backgrounds between subjects can have a considerable effect on the resolution of biomarker studies (Ahanda et al. 2014; Zhao et al. 2010). Therefore, it is critical for future research to take variations in genetic backgrounds into account and correct for additional factors such as current or previous smoking habits, alcohol abuse or medication use of patients. Finally, when comparing the obtained results, one should keep in mind the heterogeneity of miRNA expression in different tissues. Indeed, PBMC or whole blood-derived miRNA profiles will most likely differ from those obtained through serum or plasma.
Taken together, current preclinical and preliminary clinical evidence show great potential for the use of miRNAs as biomarkers of PTSD, which would enable us to detect at-risk individuals and provide specific preventive strategies and early interventions on an individual basis. This approach is especially relevant because currently no true treatment exists for PTSD. Therefore, the presented findings build an emerging foundation for future research to further examine the exact roles of miRNAs in PTSD using appropriate study designs.
References
Absalon S, Kochanek DM, Raghavan V, Krichevsky AM (2013) MiR-26b, upregulated in Alzheimer’s disease, activates cell cycle entry, tau-phosphorylation, and apoptosis in postmitotic neurons. J Neurosci 33(37):14645–14659. https://doi.org/10.1523/jneurosci.1327-13.2013
Ahanda M-LE, Zerjal T, Dhorne-Pollet S, Rau A, Cooksey A, Giuffra E (2014) Impact of the genetic background on the composition of the chicken plasma MiRNome in response to a stress. PLoS One 9(12):e114598. https://doi.org/10.1371/journal.pone.0114598
Aksoy-Aksel A, Zampa F, Schratt G (2014) MicroRNAs and synaptic plasticity—a mutual relationship. Philos Trans R Soc Lond B Biol Sci 369(1652):20130515. https://doi.org/10.1098/rstb.2013.0515
American Psychiatric Association (2013) Diagnostic and statistical manual of mental disorders, 5th edn. American Psychiatric Association, Washington, DC
Aravin AA, Hannon GJ, Brennecke J (2007) The Piwi-piRNA pathway provides an adaptive defense in the transposon arms race. Science 318(5851):761–764. https://doi.org/10.1126/science.1146484
Bai M, Zhu X, Zhang Y, Zhang S, Zhang L, Xue L, Yi J, Yao S, Zhang X (2012) Abnormal hippocampal BDNF and miR-16 expression is associated with depression-like behaviors induced by stress during early life. PLoS One 7(10):e46921. https://doi.org/10.1371/journal.pone.0046921
Baker DG, West SA, Nicholson WE, Ekhator NN, Kasckow JW, Hill KK, Bruce AB, Orth DN, Geracioti TD Jr (1999) Serial CSF corticotropin-releasing hormone levels and adrenocortical activity in combat veterans with posttraumatic stress disorder. Am J Psychiatry 156(4):585–588. https://doi.org/10.1176/ajp.156.4.585
Balakathiresan NS, Chandran R, Bhomia M, Jia M, Li H, Maheshwari RK (2014) Serum and amygdala microRNA signatures of posttraumatic stress: fear correlation and biomarker potential. J Psychiatr Res 57:65–73. https://doi.org/10.1016/j.jpsychires.2014.05.020
Bam M, Yang X, Zhou J, Ginsberg JP, Leyden Q, Nagarkatti PS, Nagarkatti M (2016a) Evidence for epigenetic regulation of pro-inflammatory cytokines, interleukin-12 and interferon gamma, in peripheral blood mononuclear cells from PTSD patients. J Neuroimmune Pharmacol 11(1):168–181. https://doi.org/10.1007/s11481-015-9643-8
Bam M, Yang X, Zumbrun EE, Zhong Y, Zhou J, Ginsberg JP, Leyden Q, Zhang J, Nagarkatti PS, Nagarkatti M (2016b) Dysregulated immune system networks in war veterans with PTSD is an outcome of altered miRNA expression and DNA methylation. Sci Rep 6:31209. https://doi.org/10.1038/srep31209
Bian S, TL X, Sun T (2013) Tuning the cell fate of neurons and glia by microRNAs. Curr Opin Neurobiol 23(6):928–934. https://doi.org/10.1016/j.conb.2013.08.002
Binder EB, Salyakina D, Lichtner P, Wochnik GM, Ising M, Putz B, Papiol S, Seaman S, Lucae S, Kohli MA, Nickel T, Kunzel HE, Fuchs B, Majer M, Pfennig A, Kern N, Brunner J, Modell S, Baghai T, Deiml T, Zill P, Bondy B, Rupprecht R, Messer T, Kohnlein O, Dabitz H, Bruckl T, Muller N, Pfister H, Lieb R, Mueller JC, Lohmussaar E, Strom TM, Bettecken T, Meitinger T, Uhr M, Rein T, Holsboer F, Muller-Myhsok B (2004) Polymorphisms in FKBP5 are associated with increased recurrence of depressive episodes and rapid response to antidepressant treatment. Nat Genet 36(12):1319–1325. https://doi.org/10.1038/ng1479
Binder EB, Bradley RG, Liu W, Epstein MP, Deveau TC, Mercer KB, Tang Y, Gillespie CF, Heim CM, Nemeroff CB, Schwartz AC, Cubells JF, Ressler KJ (2008) Association of FKBP5 polymorphisms and childhood abuse with risk of posttraumatic stress disorder symptoms in adults. JAMA 299(11):1291–1305. https://doi.org/10.1001/jama.299.11.1291
Blechert J, Michael T, Vriends N, Margraf J, Wilhelm FH (2007) Fear conditioning in posttraumatic stress disorder: evidence for delayed extinction of autonomic, experiential, and behavioural responses. Behav Res Ther 45(9):2019–2033. https://doi.org/10.1016/j.brat.2007.02.012
Bocchio-Chiavetto L, Maffioletti E, Bettinsoli P, Giovannini C, Bignotti S, Tardito D, Corrada D, Milanesi L, Gennarelli M (2013) Blood microRNA changes in depressed patients during antidepressant treatment. Eur Neuropsychopharmacol 23(7):602–611. https://doi.org/10.1016/j.euroneuro.2012.06.013
Borchert GM, Lanier W, Davidson BL (2006) RNA polymerase III transcribes human microRNAs. Nat Struct Mol Biol 13(12):1097–1101. http://www.nature.com/nsmb/journal/v13/n12/suppinfo/nsmb1167_S1.html
Camussi G, Deregibus M-C, Bruno S, Grange C, Fonsato V, Tetta C (2011) Exosome/microvesicle-mediated epigenetic reprogramming of cells. Am J Cancer Res 1(1):98–110
Chen Y-J, Luo J, Yang G-Y, Yang K, Wen S-Q, Zou S-Q (2012) Mutual regulation between microRNA-373 and methyl-CpG-binding domain protein 2 in hilar cholangiocarcinoma. World J Gastroenterol 18(29):3849–3861. https://doi.org/10.3748/wjg.v18.i29.3849
Chen RJ, Kelly G, Sengupta A, Heydendael W, Nicholas B, Beltrami S, Luz S, Peixoto L, Abel T, Bhatnagar S (2015) MicroRNAs as biomarkers of resilience or vulnerability to stress. Neuroscience 305:36–48. https://doi.org/10.1016/j.neuroscience.2015.07.045
Cho J-H, Lee I, Hammamieh R, Wang K, Baxter D, Scherler K, Etheridge A, Kulchenko A, Gautam A, Muhie S, Chakraborty N, Galas DJ, Jett M, Hood L (2014) Molecular evidence of stress-induced acute heart injury in a mouse model simulating posttraumatic stress disorder. Proc Natl Acad Sci 111(8):3188–3193
Chrousos GP, Gold PW (1992) The concepts of stress and stress system disorders. Overview of physical and behavioral homeostasis. JAMA 267(9):1244–1252
Dajas-Bailador F, Bonev B, Garcez P, Stanley P, Guillemot F, Papalopulu N (2012) microRNA-9 regulates axon extension and branching by targeting Map 1b in mouse cortical neurons. Nat Neurosci 15:697. https://doi.org/10.1038/nn.3082
Daskalakis NP, Cohen H, Nievergelt CM, Baker DG, Buxbaum JD, Russo SJ, Yehuda R (2016) New translational perspectives for blood-based biomarkers of PTSD: from glucocorticoid to immune mediators of stress susceptibility. Exp Neurol 284(Pt B):133–140. https://doi.org/10.1016/j.expneurol.2016.07.024
Davis-Dusenbery BN, Hata A (2010) Mechanisms of control of microRNA biogenesis. J Biochem 148(4):381–392. https://doi.org/10.1093/jb/mvq096
Dias BG, Goodman JV, Ahluwalia R, Easton AE, Andero R, Ressler KJ (2014) Amygdala-dependent fear memory consolidation via miR-34a and Notch signaling. Neuron 83(4):906–918. https://doi.org/10.1016/j.neuron.2014.07.019
Doxakis E (2010) Post-transcriptional regulation of alpha-synuclein expression by mir-7 and mir-153. J Biol Chem 285(17):12726–12734. https://doi.org/10.1074/jbc.M109.086827
El Khoury-Malhame M, Reynaud E, Soriano A, Michael K, Salgado-Pineda P, Zendjidjian X, Gellato C, Eric F, Lefebvre M-N, Rouby F, Samuelian J-C, Anton J-L, Blin O, Khalfa S (2011) Amygdala activity correlates with attentional bias in PTSD. Neuropsychologia 49(7):1969–1973. https://doi.org/10.1016/j.neuropsychologia.2011.03.025
Etheridge A, Lee I, Hood L, Galas D, Wang K (2011) Extracellular microRNA: a new source of biomarkers. Mutat Res 717(1–2):85–90. https://doi.org/10.1016/j.mrfmmm.2011.03.004
Etkin A, Wager TD (2007) Functional neuroimaging of anxiety: a meta-analysis of emotional processing in PTSD, social anxiety disorder, and specific phobia. Am J Psychiatry 164(10):1476–1488. https://doi.org/10.1176/appi.ajp.2007.07030504
Fabbri M, Garzon R, Cimmino A, Liu Z, Zanesi N, Callegari E, Liu S, Alder H, Costinean S, Fernandez-Cymering C, Volinia S, Guler G, Morrison CD, Chan KK, Marcucci G, Calin GA, Huebner K, Croce CM (2007) MicroRNA-29 family reverts aberrant methylation in lung cancer by targeting DNA methyltransferases 3A and 3B. Proc Natl Acad Sci U S A 104(40):15805–15810. https://doi.org/10.1073/pnas.0707628104
Fabian MR, Sonenberg N (2012) The mechanics of miRNA-mediated gene silencing: a look under the hood of miRISC. Nat Struct Mol Biol 19(6):586–593. https://doi.org/10.1038/nsmb.2296
Fiandaca MS, Kapogiannis D, Mapstone M, Boxer A, Eitan E, Schwartz JB, Abner EL, Petersen RC, Federoff HJ, Miller BL, Goetzl EJ (2015) Identification of preclinical Alzheimer’s disease by a profile of pathogenic proteins in neurally derived blood exosomes: a case-control study. Alzheimers Dement 11(6):600–607.e601. https://doi.org/10.1016/j.jalz.2014.06.008
Friedman RC, Farh KK, Burge CB, Bartel DP (2009) Most mammalian mRNAs are conserved targets of microRNAs. Genome Res 19(1):92–105. https://doi.org/10.1101/gr.082701.108
Gallo V, Egger M, McCormack V, Farmer PB, Ioannidis JPA, Kirsch-Volders M, Matullo G, Phillips DH, Schoket B, Stromberg U, Vermeulen R, Wild C, Porta M, Vineis P (2011) Strengthening the reporting of observational studies in epidemiology-molecular epidemiology STROBE-ME: an extension of the STROBE statement. J Clin Epidemiol 64(12):1350–1363. https://doi.org/10.1016/j.jclinepi.2011.07.010
Gapp K, Jawaid A, Sarkies P, Bohacek J, Pelczar P, Prados J, Farinelli L, Miska E, Mansuy IM (2014) Implication of sperm RNAs in transgenerational inheritance of the effects of early trauma in mice. Nat Neurosci 17(5):667–669. https://doi.org/10.1038/nn.3695
Geracioti TD Jr, Baker DG, Kasckow JW, Strawn JR, Jeffrey Mulchahey J, Dashevsky BA, Horn PS, Ekhator NN (2008) Effects of trauma-related audiovisual stimulation on cerebrospinal fluid norepinephrine and corticotropin-releasing hormone concentrations in post-traumatic stress disorder. Psychoneuroendocrinology 33(4):416–424. https://doi.org/10.1016/j.psyneuen.2007.12.012
Gibbings DJ, Ciaudo C, Erhardt M, Voinnet O (2009) Multivesicular bodies associate with components of miRNA effector complexes and modulate miRNA activity. Nat Cell Biol 11(9):1143–1149. https://doi.org/10.1038/ncb1929
Gill JM, Saligan L, Woods S, Page G (2009) PTSD is associated with an excess of inflammatory immune activities. Perspect Psychiatr Care 45(4):262–277. https://doi.org/10.1111/j.1744-6163.2009.00229.x
Goetzl EJ, Boxer A, Schwartz JB, Abner EL, Petersen RC, Miller BL, Kapogiannis D (2015) Altered lysosomal proteins in neural-derived plasma exosomes in preclinical Alzheimer disease. Neurology 85(1):40–47. https://doi.org/10.1212/wnl.0000000000001702
Griggs EM, Young EJ, Rumbaugh G, Miller CA (2013) MicroRNA-182 regulates amygdala-dependent memory formation. J Neurosci 33(4):1734
Hanin G, Shenhar-Tsarfaty S, Yayon N, Hoe YY, Bennett ER, Sklan EH, Rao DC, Rankinen T, Bouchard C, Geifman-Shochat S, Shifman S, Greenberg DS, Soreq H (2014) Competing targets of microRNA-608 affect anxiety and hypertension. Hum Mol Genet 23(17):4569–4580
Hanke M, Hoefig K, Merz H, Feller AC, Kausch I, Jocham D, Warnecke JM, Sczakiel G (2010) A robust methodology to study urine microRNA as tumor marker: microRNA-126 and microRNA-182 are related to urinary bladder cancer. Urol Oncol 28(6):655–661. https://doi.org/10.1016/j.urolonc.2009.01.027
Hu Z, Li Z (2017) miRNAs in synapse development and synaptic plasticity. Curr Opin Neurobiol 45:24–31. https://doi.org/10.1016/j.conb.2017.02.014
Hu YK, Wang X, Li L, YH D, Ye HT, Li CY (2013) MicroRNA-98 induces an Alzheimer’s disease-like disturbance by targeting insulin-like growth factor 1. Neurosci Bull 29(6):745–751. https://doi.org/10.1007/s12264-013-1348-5
Issler O, Chen A (2015) Determining the role of microRNAs in psychiatric disorders. Nat Rev Neurosci 16(4):201–212. https://doi.org/10.1038/nrn3879
Jin Y, Lee CG (2013) Single nucleotide polymorphisms associated with microRNA regulation. Biomol Ther 3(2):287–302. https://doi.org/10.3390/biom3020287
Jovasevic V, Corcoran KA, Leaderbrand K, Yamawaki N, Guedea AL, Chen HJ, Shepherd GM, Radulovic J (2015) GABAergic mechanisms regulated by miR-33 encode state-dependent fear. Nat Neurosci 18(9):1265–1271. https://doi.org/10.1038/nn.4084
Kapogiannis D, Boxer A, Schwartz JB, Abner EL, Biragyn A, Masharani U, Frassetto L, Petersen RC, Miller BL, Goetzl EJ (2015) Dysfunctionally phosphorylated type 1 insulin receptor substrate in neural-derived blood exosomes of preclinical Alzheimer’s disease. FASEB J 29(2):589–596. https://doi.org/10.1096/fj.14-262048
Karl A, Schaefer M, Malta LS, Dörfel D, Rohleder N, Werner A (2006) A meta-analysis of structural brain abnormalities in PTSD. Neurosci Biobehav Rev 30(7):1004–1031. https://doi.org/10.1016/j.neubiorev.2006.03.004
Klengel T, Mehta D, Anacker C, Rex-Haffner M, Pruessner JC, Pariante CM, Pace TW, Mercer KB, Mayberg HS, Bradley B, Nemeroff CB, Holsboer F, Heim CM, Ressler KJ, Rein T, Binder EB (2013) Allele-specific FKBP5 DNA demethylation mediates gene-childhood trauma interactions. Nat Neurosci 16(1):33–41. https://doi.org/10.1038/nn.3275
Konopka W, Kiryk A, Novak M, Herwerth M, Parkitna JR, Wawrzyniak M, Kowarsch A, Michaluk P, Dzwonek J, Arnsperger T, Wilczynski G, Merkenschlager M, Theis FJ, Kohr G, Kaczmarek L, Schutz G (2010) MicroRNA loss enhances learning and memory in mice. J Neurosci 30(44):14835–14842. https://doi.org/10.1523/JNEUROSCI.3030-10.2010
Kouzarides T (2007) Chromatin modifications and their function. Cell 128(4):693–705. https://doi.org/10.1016/j.cell.2007.02.005
Lai CY, SL Y, Hsieh MH, Chen CH, Chen HY, Wen CC, Huang YH, Hsiao PC, Hsiao CK, Liu CM, Yang PC, Hwu HG, Chen WJ (2011) MicroRNA expression aberration as potential peripheral blood biomarkers for schizophrenia. PLoS One 6(6):e21635. https://doi.org/10.1371/journal.pone.0021635
Lee Y, Ahn C, Han J, Choi H, Kim J, Yim J, Lee J, Provost P, Radmark O, Kim S, Kim VN (2003) The nuclear RNase III Drosha initiates microRNA processing. Nature 425(6956):415–419. https://doi.org/10.1038/nature01957
Lemieux AM, Coe CL (1995) Abuse-related posttraumatic stress disorder: evidence for chronic neuroendocrine activation in women. Psychosom Med 57(2):105–115
Lewis BP, Burge CB, Bartel DP (2005) Conserved seed pairing, often flanked by adenosines, indicates that thousands of human genes are microRNA targets. Cell 120(1):15–20. https://doi.org/10.1016/j.cell.2004.12.035
Li C, Liu Y, Liu D, Jiang H, Pan F (2016) Dynamic alterations of miR-34c expression in the hypothalamus of male rats after early adolescent traumatic stress. Neural Plast 2016:5249893. https://doi.org/10.1155/2016/5249893
Lin Q, Wei W, Coelho CM, Li X, Baker-Andresen D, Dudley K, Ratnu VS, Boskovic Z, Kobor MS, Sun YE, Bredy TW (2011) The brain-specific microRNA miR-128b regulates the formation of fear-extinction memory. Nat Neurosci 14(9):1115–1117. https://doi.org/10.1038/nn.2891
Liu C, Teng Z-Q, Santistevan NJ, Szulwach KE, Guo W, Jin P, Zhao X (2010) Epigenetic regulation of miR-184 by MBD1 governs neural stem cell proliferation and differentiation. Cell Stem Cell 6(5):433–444. https://doi.org/10.1016/j.stem.2010.02.017
Liu W-M, Pang RTK, Chiu PCN, Wong BPC, Lao K, Lee K-F, Yeung WSB (2012) Sperm-borne microRNA-34c is required for the first cleavage division in mouse. Proc Natl Acad Sci U S A 109(2):490–494. https://doi.org/10.1073/pnas.1110368109
Magill ST, Cambronne XA, Luikart BW, Lioy DT, Leighton BH, Westbrook GL, Mandel G, Goodman RH (2010) microRNA-132 regulates dendritic growth and arborization of newborn neurons in the adult hippocampus. Proc Natl Acad Sci U S A 107(47):20382–20387. https://doi.org/10.1073/pnas.1015691107
Martin CG, Kim H, Yun S, Livingston W, Fetta J, Mysliwiec V, Baxter T, Gill JM (2017) Circulating miRNA associated with posttraumatic stress disorder in a cohort of military combat veterans. Psychiatry Res 251:261–265. https://doi.org/10.1016/j.psychres.2017.01.081
Meewisse ML, Reitsma JB, de Vries GJ, Gersons BP, Olff M (2007) Cortisol and post-traumatic stress disorder in adults: systematic review and meta-analysis. Br J Psychiatry 191:387–392. https://doi.org/10.1192/bjp.bp.106.024877
Mehta D, Gonik M, Klengel T, Rex-Haffner M, Menke A, Rubel J, Mercer KB, Putz B, Bradley B, Holsboer F, Ressler KJ, Muller-Myhsok B, Binder EB (2011) Using polymorphisms in FKBP5 to define biologically distinct subtypes of posttraumatic stress disorder: evidence from endocrine and gene expression studies. Arch Gen Psychiatry 68(9):901–910. https://doi.org/10.1001/archgenpsychiatry.2011.50
Memczak S, Jens M, Elefsinioti A, Torti F, Krueger J, Rybak A, Maier L, Mackowiak SD, Gregersen LH, Munschauer M, Loewer A, Ziebold U, Landthaler M, Kocks C, le Noble F, Rajewsky N (2013) Circular RNAs are a large class of animal RNAs with regulatory potency. Nature 495(7441):333–338. http://www.nature.com/nature/journal/v495/n7441/abs/nature11928.html#supplementary-information
Michopoulos V, Powers A, Gillespie CF, Ressler KJ, Jovanovic T (2017) Inflammation in fear- and anxiety-based disorders: PTSD, GAD, and beyond. Neuropsychopharmacology 42(1):254–270. https://doi.org/10.1038/npp.2016.146
Mitchell PS, Parkin RK, Kroh EM, Fritz BR, Wyman SK, Pogosova-Agadjanyan EL, Peterson A, Noteboom J, O’Briant KC, Allen A, Lin DW, Urban N, Drescher CW, Knudsen BS, Stirewalt DL, Gentleman R, Vessella RL, Nelson PS, Martin DB, Tewari M (2008) Circulating microRNAs as stable blood-based markers for cancer detection. Proc Natl Acad Sci U S A 105(30):10513–10518. https://doi.org/10.1073/pnas.0804549105
Nair VS, Pritchard CC, Tewari M, Ioannidis JP (2014) Design and analysis for studying microRNAs in human disease: a primer on -Omic technologies. Am J Epidemiol 180(2):140–152. https://doi.org/10.1093/aje/kwu135
Neigh GN, Ali FF (2016) Co-morbidity of PTSD and immune system dysfunction: opportunities for treatment. Curr Opin Pharmacol 29:104–110. https://doi.org/10.1016/j.coph.2016.07.011
Norberg MM, Krystal JH, Tolin DF (2008) A meta-analysis of D-cycloserine and the facilitation of fear extinction and exposure therapy. Biol Psychiatry 63(12):1118–1126. https://doi.org/10.1016/j.biopsych.2008.01.012
Park NJ, Zhou H, Elashoff D, Henson BS, Kastratovic DA, Abemayor E, Wong DT (2009) Salivary microRNA: discovery, characterization, and clinical utility for oral cancer detection. Clin Cancer Res 15(17):5473–5477. https://doi.org/10.1158/1078-0432.CCR-09-0736
Pepe MS, Feng Z, Janes H, Bossuyt PM, Potter JD (2008) Pivotal evaluation of the accuracy of a biomarker used for classification or prediction: standards for study design. J Natl Canc Inst 100(20):1432–1438. https://doi.org/10.1093/jnci/djn326
Peschansky VJ, Wahlestedt C (2014) Non-coding RNAs as direct and indirect modulators of epigenetic regulation. Epigenetics 9(1):3–12. https://doi.org/10.4161/epi.27473
Pritchard CC, Cheng HH, Tewari M (2012) MicroRNA profiling: approaches and considerations. Nat Rev Genet 13(5):358–369. https://doi.org/10.1038/nrg3198
Raabe FJ, Spengler D (2013) Epigenetic risk factors in PTSD and depression. Front Psych 4:80. https://doi.org/10.3389/fpsyt.2013.00080
Ramalingam P, Palanichamy JK, Singh A, Das P, Bhagat M, Kassab MA, Sinha S, Chattopadhyay P (2014) Biogenesis of intronic miRNAs located in clusters by independent transcription and alternative splicing. RNA 20(1):76–87. https://doi.org/10.1261/rna.041814.113
Rassoulzadegan M, Grandjean V, Gounon P, Vincent S, Gillot I, Cuzin F (2006) RNA-mediated non-mendelian inheritance of an epigenetic change in the mouse. Nature 441(7092):469–474. https://doi.org/10.1038/nature04674
Rodgers AB, Morgan CP, Bronson SL, Revello S, Bale TL (2013) Paternal stress exposure alters sperm microRNA content and reprograms offspring HPA stress axis regulation. J Neurosci 33(21):9003–9012. https://doi.org/10.1523/JNEUROSCI.0914-13.2013
Rong H, Liu TB, Yang KJ, Yang HC, Wu DH, Liao CP, Hong F, Yang HZ, Wan F, Ye XY, Xu D, Zhang X, Chao CA, Shen QJ (n.d.) MicroRNA-134 plasma levels before and after treatment for bipolar mania. J Psychiatr Res 45(1):92–95. https://doi.org/10.1016/jjpsychires201004.028
Rose NR, Klose RJ (2014) Understanding the relationship between DNA methylation and histone lysine methylation. Biochim Biophys Acta 1839(12):1362–1372. https://doi.org/10.1016/j.bbagrm.2014.02.007
Sabbagh JJ, O’Leary JC, Blair LJ, Klengel T, Nordhues BA, Fontaine SN, Binder EB, Dickey CA (2014) Age-associated epigenetic Upregulation of the FKBP5 gene selectively impairs stress resiliency. PLoS One 9(9):e107241. https://doi.org/10.1371/journal.pone.0107241
Sato F, Tsuchiya S, Meltzer SJ, Shimizu K (2011) MicroRNAs and epigenetics. FEBS J 278(10):1598–1609. https://doi.org/10.1111/j.1742-4658.2011.08089.x
Satrom P, Snove O, Rossi JJ (2007) Epigenetics and microRNAs. Pediatr Res 61(5 Part 2):17R–23R
Schmidt U, Holsboer F, Rein T (2011) Epigenetic aspects of posttraumatic stress disorder. Dis Markers 30(2–3):77–87. https://doi.org/10.3233/DMA-2011-0749
Schmidt MV, Paez-Pereda M, Holsboer F, Hausch F (2012) The prospect of FKBP51 as a drug target. ChemMedChem 7(8):1351–1359. https://doi.org/10.1002/cmdc.201200137
Schmidt U, Herrmann L, Hagl K, Novak B, Huber C, Holsboer F, Wotjak CT, Buell DR (2013) Therapeutic action of fluoxetine is associated with a reduction in prefrontal cortical miR-1971 expression levels in a mouse model of posttraumatic stress disorder. Front Psych 4:66. https://doi.org/10.3389/fpsyt.2013.00066
Schouten M, Aschrafi A, Bielefeld P, Doxakis E, Fitzsimons CP (2013) MicroRNAs and the regulation of neuronal plasticity under stress conditions. Neuroscience 241:188–205. https://doi.org/10.1016/j.neuroscience.2013.02.065
Scott GK, Mattie MD, Berger CE, Benz SC, Benz CC (2006) Rapid alteration of microRNA levels by histone deacetylase inhibition. Cancer Res 66(3):1277–1281. https://doi.org/10.1158/0008-5472.can-05-3632
Sherin JEN, Charles B (2011) Post-traumatic stress disorder: the neurobiological impact of psychological trauma. Dialogues Clin Neurosci 13(3):263
Sherin JE, Nemeroff CB (2011) Post-traumatic stress disorder: the neurobiological impact of psychological trauma. Dialogues Clin Neurosci 13(3):263–278
Shin LM, Rauch SL, Pitman RK (2006) Amygdala, medial prefrontal cortex, and hippocampal function in PTSD. Ann N Y Acad Sci 1071:67–79. https://doi.org/10.1196/annals.1364.007
Sinkkonen L, Hugenschmidt T, Berninger P, Gaidatzis D, Mohn F, Artus-Revel CG, Zavolan M, Svoboda P, Filipowicz W (2008) MicroRNAs control de novo DNA methylation through regulation of transcriptional repressors in mouse embryonic stem cells. Nat Struct Mol Biol 15(3):259–267. http://www.nature.com/nsmb/journal/v15/n3/suppinfo/nsmb.1391_S1.html
Smalheiser NR, Lugli G, Rizavi HS, Torvik VI, Turecki G, Dwivedi Y (2012) MicroRNA expression is down-regulated and reorganized in prefrontal cortex of depressed suicide subjects. PLoS One 7(3):e33201. https://doi.org/10.1371/journal.pone.0033201
Smith ME (2005) Bilateral hippocampal volume reduction in adults with post-traumatic stress disorder: a meta-analysis of structural MRI studies. Hippocampus 15(6):798–807. https://doi.org/10.1002/hipo.20102
Smith B, Treadwell J, Zhang D, Ly D, McKinnell I, Walker PR, Sikorska M (2010) Large-scale expression analysis reveals distinct microRNA profiles at different stages of human neurodevelopment. PLoS One 5(6):e11109. https://doi.org/10.1371/journal.pone.0011109
Taylor D, Gercel-Taylor C (2013) The origin, function, and diagnostic potential of RNA within extracellular vesicles present in human biological fluids. Front Genet 4(142). https://doi.org/10.3389/fgene.2013.00142
Turchinovich A, Samatov T, Tonevitsky A, Burwinkel B (2013) Circulating miRNAs: cell-cell communication function? Front Genet 4(119). https://doi.org/10.3389/fgene.2013.00119
Valadi H, Ekstrom K, Bossios A, Sjostrand M, Lee JJ, Lotvall JO (2007) Exosome-mediated transfer of mRNAs and microRNAs is a novel mechanism of genetic exchange between cells. Nat Cell Biol 9(6):654–659. http://www.nature.com/ncb/journal/v9/n6/suppinfo/ncb1596_S1.html
Van den Hove DL, Kompotis K, Lardenoije R, Kenis G, Mill J, Steinbusch HW, Lesch KP, Fitzsimons CP, De Strooper B, Rutten BP (2014) Epigenetically regulated microRNAs in Alzheimer’s disease. Neurobiol Aging 35(4):731–745. https://doi.org/10.1016/j.neurobiolaging.2013.10.082
Venkatesh S, Workman JL (2015) Histone exchange, chromatin structure and the regulation of transcription. Nat Rev Mol Cell Biol 16(3):178–189. https://doi.org/10.1038/nrm3941
Vetere G, Barbato C, Pezzola S, Frisone P, Aceti M, Ciotti M, Cogoni C, Ammassari-Teule M, Ruberti F (2014) Selective inhibition of miR-92 in hippocampal neurons alters contextual fear memory. Hippocampus 24(12):1458–1465. https://doi.org/10.1002/hipo.22326
Vickers KC, Palmisano BT, Shoucri BM, Shamburek RD, Remaley AT (2011) MicroRNAs are transported in plasma and delivered to recipient cells by high-density lipoproteins. Nat Cell Biol 13(4):423–433. https://doi.org/10.1038/ncb2210
Volk N, Paul ED, Haramati S, Eitan C, Fields BK, Zwang R, Gil S, Lowry CA, Chen A (2014) MicroRNA-19b associates with Ago2 in the amygdala following chronic stress and regulates the adrenergic receptor beta 1. J Neurosci 34(45):15070–15082. https://doi.org/10.1523/JNEUROSCI.0855-14.2014
Volk N, Pape JC, Engel M, Zannas AS, Cattane N, Cattaneo A, Binder EB, Chen A (2016) Amygdalar MicroRNA-15a is essential for coping with chronic stress. Cell Rep 17(7):1882–1891. https://doi.org/10.1016/j.celrep.2016.10.038
Wagner J, Riwanto M, Besler C, Knau A, Fichtlscherer S, Roxe T, Zeiher AM, Landmesser U, Dimmeler S (2013) Characterization of levels and cellular transfer of circulating lipoprotein-bound microRNAs. Arterioscler Thromb Vasc Biol 33(6):1392–1400. https://doi.org/10.1161/ATVBAHA.112.300741
Wang G, van der Walt JM, Mayhew G, Li YJ, Zuchner S, Scott WK, Martin ER, Vance JM (2008) Variation in the miRNA-433 binding site of FGF20 confers risk for Parkinson disease by overexpression of alpha-synuclein. Am J Hum Genet 82(2):283–289. https://doi.org/10.1016/j.ajhg.2007.09.021
Wang K, Zhang S, Weber J, Baxter D, Galas DJ (2010) Export of microRNAs and microRNA-protective protein by mammalian cells. Nucleic Acids Res 38:7248. https://doi.org/10.1093/nar/gkq601
Wang RY, Phang RZ, Hsu PH, Wang WH, Huang HT, Liu IY (2013) In vivo knockdown of hippocampal miR-132 expression impairs memory acquisition of trace fear conditioning. Hippocampus 23(7):625–633. https://doi.org/10.1002/hipo.22123
Weber JA, Baxter DH, Zhang S, Huang DY, Huang KH, Lee MJ, Galas DJ, Wang K (2010) The microRNA spectrum in 12 body fluids. Clin Chem 56(11):1733–1741. https://doi.org/10.1373/clinchem.2010.147405
Wingo AP, Almli LM, Stevens JS, Klengel T, Uddin M, Li Y, Bustamante AC, Lori A, Koen N, Stein DJ, Smith AK, Aiello AE, Koenen KC, Wildman DE, Galea S, Bradley B, Binder EB, Jin P, Gibson G, Ressler KJ (2015) DICER1 and microRNA regulation in post-traumatic stress disorder with comorbid depression. Nat Commun 6:10106. https://doi.org/10.1038/ncomms10106
Winston CN, Goetzl EJ, Akers JC, Carter BS, Rockenstein EM, Galasko D, Masliah E, Rissman RA (2016) Prediction of conversion from mild cognitive impairment to dementia with neuronally derived blood exosome protein profile. Alzheimers Dement (Amst) 3:63–72. https://doi.org/10.1016/j.dadm.2016.04.001
Yang X, WW D, Li H, Liu F, Khorshidi A, Rutnam ZJ, Yang BB (2013) Both mature miR-17-5p and passenger strand miR-17-3p target TIMP3 and induce prostate tumor growth and invasion. Nucleic Acids Res 41(21):9688–9704. https://doi.org/10.1093/nar/gkt680
Yehuda R (2001) Biology of posttraumatic stress disorder. J Clin Psychiatry 62(Suppl 17):41–46
Yehuda R (2006) Advances in understanding neuroendocrine alterations in PTSD and their therapeutic implications. Ann N Y Acad Sci 1071:137–166. https://doi.org/10.1196/annals.1364.012
Yehuda R, Bierer LM, Schmeidler J, Aferiat DH, Breslau I, Dolan S (2000) Low cortisol and risk for PTSD in adult offspring of holocaust survivors. Am J Psychiatry 157(8):1252–1259. https://doi.org/10.1176/appi.ajp.157.8.1252
Yehuda R, Daskalakis NP, Bierer LM, Bader HN, Klengel T, Holsboer F, Binder EB (2016) Holocaust exposure induced intergenerational effects on FKBP5 methylation. Biol Psychiatry 80(5):372–380. https://doi.org/10.1016/j.biopsych.2015.08.005
Young EA, Breslau N (2004) Cortisol and catecholamines in posttraumatic stress disorder: an epidemiologic community study. Arch Gen Psychiatry 61(4):394–401
Zamore PD (2002) Ancient pathways programmed by small RNAs. Science 296(5571):1265–1269. https://doi.org/10.1126/science.1072457
Zannas AS, Provencal N, Binder EB (2015) Epigenetics of posttraumatic stress disorder: current evidence, challenges, and future directions. Biol Psychiatry 78(5):327–335. https://doi.org/10.1016/j.biopsych.2015.04.003
Zhao H, Shen J, Medico L, Wang D, Ambrosone CB, Liu S (2010) A pilot study of circulating miRNAs as potential biomarkers of early stage breast cancer. PLoS One 5(10):e13735. https://doi.org/10.1371/journal.pone.0013735
Zhao H, Yao R, Cao X, Wu G (2011) Neuroimmune modulation following traumatic stress in rats: evidence for an immunoregulatory cascade mediated by c-Src, miRNA222 and PAK1. J Neuroinflammation 8(1):159. https://doi.org/10.1186/1742-2094-8-159
Zheng D, Sabbagh JJ, Blair LJ, Darling AL, Wen X, Dickey CA (2016) MicroRNA-511 binds to FKBP5 mRNA, which encodes a chaperone protein, and regulates neuronal differentiation. J Biol Chem 291(34):17897–17906. https://doi.org/10.1074/jbc.M116.727941
Zhou J, Nagarkatti P, Zhong Y, Ginsberg JP, Singh NP, Zhang J, Nagarkatti M (2014) Dysregulation in microRNA expression is associated with alterations in immune functions in combat veterans with post-traumatic stress disorder. PLoS One 9(4):e94075. https://doi.org/10.1371/journal.pone.0094075
Zimmermann P, Bruckl T, Nocon A, Pfister H, Binder EB, Uhr M, Lieb R, Moffitt TE, Caspi A, Holsboer F, Ising M (2011) Interaction of FKBP5 gene variants and adverse life events in predicting depression onset: results from a 10-year prospective community study. Am J Psychiatry 168(10):1107–1116. https://doi.org/10.1176/appi.ajp.2011.10111577
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2017 Springer International Publishing AG
About this chapter
Cite this chapter
Snijders, C. et al. (2017). MicroRNAs in Post-traumatic Stress Disorder. In: Vermetten, E., Baker, D.G., Risbrough, V.B. (eds) Behavioral Neurobiology of PTSD. Current Topics in Behavioral Neurosciences, vol 38. Springer, Cham. https://doi.org/10.1007/7854_2017_32
Download citation
DOI: https://doi.org/10.1007/7854_2017_32
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-319-94823-2
Online ISBN: 978-3-319-94824-9
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)