Abstract
Aims
Extensively-drug-resistant Pseudomonas aeruginosa constitutes a serious threat to patients suffering from Cystic Fibrosis (CF). In these patients, P. aeruginosa lung infection is commonly treated with aminoglycosides, but treatments are largely unsuccessful due a variety of resistance mechanisms. Here we investigate the prevalence of resistance to gentamicin, amikacin and tobramycin and the main aminoglycoside resistance genes found in P. aeruginosa strains isolated at a regional CF centre.
Results
A total number of 147 randomly selected P. aeruginosa strains isolated from respiratory samples sent by the Marche regional Cystic Fibrosis Centre to the Microbiology lab, were included in this study. Of these, 78 (53%) were resistant to at least one of the three aminoglycosides tested and 27% were resistant to all three antibiotics, suggesting a major involvement of a chromosome-encoded mechanism, likely MexXY-OprM efflux pump overexpression. A specific pathogenic clone (found in 7/78 of the aminoglycoside resistant strains) carrying ant(2″)-Ia was isolated over time from the same patient, suggesting a role for this additional resistance gene in the antibiotic unresponsiveness of CF patients.
Conclusions
The MexXY-OprM efflux pump is confirmed as the resistance determinant involved most frequently in P. aeruginosa aminoglycoside resistance of CF lung infections, followed by the ant(2″)-Ia-encoded adenylyltransferase. The latter may prove to be a novel target for new antimicrobial combinations against P. aeruginosa.
Co-first authors: Gianmarco Mangiaterra and Nicholas Cedraro
Access provided by Autonomous University of Puebla. Download chapter PDF
Similar content being viewed by others
Keywords
- Acquired resistance determinants
- Aminoglycoside resistance
- Cystic fibrosis
- Efflux pumps
- Pseudomonas aeruginosa
1 Introduction
Extensively drug-resistant (XDR) Pseudomonas aeruginosa is a major cause of mortality in immunocompromised subjects, particularly in Cystic Fibrosis (CF) patients, whose chronic lung infections are mainly caused by this microorganism and are commonly treated with antimicrobial combinations that include tobramycin. Aminoglycosides (AMGs) are included in the gold standard treatment for P. aeruginosa lung infections and several studies have confirmed the efficacy of tobramycin (Stanojevic et al. 2014; Buttini et al. 2018). However, the development of AMG resistance is a growing threat hampering infection eradication (Costello et al. 2018).
AMG resistance relies on a broad variety of mechanisms (Poole 2005). First of all, the MexXY-OprM efflux pump (EP) is the main cause of the failure of routine antibiotic treatments against P. aeruginosa lung infection; in particular, it is involved in adaptive resistance (Morita et al. 2012), which enables pathogen survival in the presence of AMGs. The proneness of P. aeruginosa to accumulate mutations in the mexXY regulatory gene mexZ, results in EP overexpression and heightened resistance (Frimodt-Møller et al. 2018). Biofilm production hampers antibiotic action, further contributing to AMG resistance (Müller et al. 2018); notably, the genes involved in resistance development are upregulated in biofilm-embedded cells (Soto 2013; Hall et al. 2018), as reported for ndvB, encoding for cyclic glucans responsible of AMGs sequestration in the periplasmic space (Mah et al. 2003). Another recently described AMG resistance mechanism involves mutations in the fusA1 gene, leading to single amino acid substitutions in the elongation factor G (EF-G1A), resulting in a lower drug affinity for the ribosome (Bolard et al. 2018).
Moreover, P. aeruginosa acquires new resistance determinants through horizontal gene transfer, which induces the formation of gene mosaics; the great variability of its genome results in a marked heterogeneity of AMG-resistant clones (Partridge et al. 2018). Enzymes that modify both the drug and its target are involved in the spread of high-level resistance (Poole 2011).
In the past few years, the search for new compounds capable of counteracting antibiotic resistance has mainly been directed at the modulation of EPs, a constitutive mechanism conferring resistance against several different antibiotics, which are often upregulated in P. aeruginosa (Mangiaterra et al. 2017; Laudadio et al. 2019). Another key area of investigation are other resistance mechanisms characterizing XDR P. aeruginosa.
We studied the prevalence of AMG resistance among the P. aeruginosa strains isolated from the CF patients managed by a regional referral centre in Marche (central Italy), by analysing the level of resistance to gentamycin, amikacin and tobramycin – the antibiotics routinely tested in the microbiology lab of the CF centre – and the resistance genes involved most frequently. In particular, we studied the frequency of transferable chromosome-independent resistance determinants and compared it with the frequency of chromosome-encoded resistances not involved in horizontal gene transfer events. We also compared the spread of the resistance mechanisms specific to each antibiotic with those providing resistance to all AMGs.
2 Material and Methods
2.1 Bacterial Strains, Media and Antibiotics
A total number of 147 randomly selected anonymized P. aeruginosa strains, sent for isolation by the Marche regional Cystic Fibrosis Centre to the Microbiology lab of “Ospedali Riuniti” Hospital (Ancona, IT), were included in the study. The strains had been collected from April 2014 to March 2015 (P. aeruginosa C1-C51) and from October 2015 to January 2016 (P. aeruginosa AR1-AR96) (Supplementary Table 1).
Additional strains included in the study were P. aeruginosa PAO1 and PA14, kindly provided by Prof. Olivier Jousson of the Integrated Biology Centre, University of Trento (Trento, IT), P. aeruginosa PAO1, carrying plasmid pHERD30T, kindly provided by Prof. Paul Williams of the Centre of Biomolecular Sciences, University of Nottingham (UK), and P. aeruginosa ATCC 27853, from the collection of the Microbiology section of the Department of Life and Environmental Sciences, Polytechnic University of Marche (Ancona, IT). All strains were cultured in Luria Bertani (LB) broth and Pseudomonas cetrimide agar plates and stored as stock cultures in LB broth supplemented with 20% glycerol at −80 °C.
All media were purchased from Oxoid (Oxoid S.p.A., Rodano, Milano, IT); the antibiotics were obtained from Sigma-Aldrich (Saint Louis, MO, USA).
2.2 Antibiotic Susceptibility Tests
The P. aeruginosa strains were screened for their resistance to tobramycin (TOB), gentamicin (GEN) and amikacin (AMK) by a routine antibiotic susceptibility test (Sensititre™ Complete Automated AST System, Thermo Fisher Scientific, Waltham, MA, USA). The resistant phenotype was confirmed by determination of the Minimal Inhibitory Concentration (MIC) either by agar dilution or by broth microdilution according to European Committee on Antimicrobial Susceptibility Testing (EUCAST) guidelines (2003) using P. aeruginosa ATCC 27853 as the reference strain. The results were interpreted according to the EUCAST epidemiological cut-off (ECOFF) values (2016). Strains showing intermediate susceptibility were considered resistant. The following antibiotic concentration ranges were used: GEN, 0.5–32 μg/ml; AMK, 1–64 μg/ml; TOB, 0.25–16 μg/ml.
2.3 Detection of Aminoglycoside Resistance Genes
Two chromosome-encoded resistance determinants (mexY and ndvB) and 4 additional AMG resistance determinants (aac(3)-Ia, aph(3′)-IIa, ant(2″)-Ia and rmtA) were sought by PCR assays. Bacterial DNA was obtained from crude cell lysates (Hynes et al. 1992) and 5 μl were used in each PCR reaction together with 1.25 U Dream-Taq Polymerase (Thermo Fisher Scientific), 1x PCR Buffer, 0.2 mM dNTPs and 0.5 μM of each primer. The target genes and the specific primer pairs used are reported in Table 1. P. aeruginosa PAO1 and PA14 DNA was used as positive control in PCRs targeting mexY and ndvB, respectively; P. aeruginosa PAO1 harbouring the plasmid pHERD30T, which carries the aac(3)-Ia gene, was used as a positive control in the relevant PCR assays. Amplicons rmtA, aph(3′)-IIa and ant(2″)-Ia of the right size were purified (Gene Elute PCR Cleanup kit, Sigma-Aldrich) and sequenced using BigDye Terminator v.1.1 Cycle Sequencing kit according to the manufacturer’s instructions. The sequences were analysed on an ABI Prism 310 Genetic Analyzer (Applied Biosystems, Foster City, CA, USA), and used as positive controls in PCRs targeting the corresponding gene. RNase-free water (Thermo Fisher Scientific) was used as the negative control. The PCR products were checked by 1.5% agarose gel electrophoresis.
2.4 Pulsed Field Gel Electrophoresis Typing
Pulsed Field Gel Electrophoresis (PFGE) was performed as described by Seifert et al. (2005), with some modifications. Briefly, agarose plugs (low-melting 1.6% agarose) were digested with 30 U of SpeI (Thermo Fisher Scientific) and the restriction fragments were separated using a CHEF MAPPER system (BioRad, Hercules, CA, USA). Running conditions were as follows: switch angle 120°, voltage 6 V/cm, temperature 14 °C, run time 18 h, initial and final switching time 5.8 s and 30 s, respectively. The low-range PFG marker (Amersham Biosciences, Little Chalfont, UK) was used as a molecular weight marker and PFGE patterns were analysed as described previously (Biavasco et al. 2007). Dendrograms were drawn using the TreeCon software.
2.5 Statistical Analysis
The frequency of P. aeruginosa strains resistant to each of the three antibiotics was evaluated by the chi square test. Values were considered statistically significant when p was <0.05.
3 Results
3.1 Detection of Aminoglycoside Resistance in CF P. aeruginosa Strains
The screening by Sensititre™ of 147 CF P. aeruginosa strains for resistance to gentamicin, amikacin and tobramycin by routine susceptibility tests found that 78 (53%) were resistant to at least one antibiotic. These 78 strains were subjected to further analyses. Testing of their resistance level by agar dilution (Supplementary Table 1) demonstrated that 66 were resistant to gentamicin, 66 were resistant to amikacin (each accounting for 84.62%) and 27 were resistant to tobramycin (34.62%). The frequency of strains resistant to gentamicin or amikacin was significantly (p < 0.01) higher than that of tobramycin-resistant isolates (Fig. 1).
In particular, resistance to all 3 antibiotics (26.92%) was statistically more common than resistance to gentamicin (7.69%) or tobramycin (2.56%) alone (p < 0.05 and p < 0.01, respectively), whereas resistance to gentamicin+amikacin was significantly (p < 0.01) more frequent (44.87%) than tobramycin resistance (2.56%) (Fig. S1). There were no strains resistant to amikacin+tobramycin.
3.2 Detection of Aminoglycoside Resistance Genes
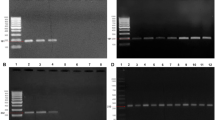
The 78 strains resistant to gentamicin, amikacin and/or tobramycin were analysed for their AMG resistance genes by PCR assays targeting two chromosome-encoded (mexY and ndvB) genes and four additional (rmtA, aac(3)-Ia, ant(2″)-Ia and aph(3′)-IIa) genes (Supplementary Table 1).
mexY was detected in all strains and ndvB in all strains but one (C36), aac(3)-Ia and ant(2″)-Ia were detected respectively in 4 (C40, AR15, AR39 and AR66) and 7 (C44, C45, AR45, AR49, AR86, AR88 and AR89) isolates, whereas all strains were negative for aph(3′)-IIa and rmtA. Sequencing of ant(2″)-Ia amplicon of P. aeruginosa AR86 demonstrated a similarity of 99% to the sequence of P. aeruginosa strain PB350 (Accession no. CP025055.1). There were 11 strains carrying aac(3)-Ia or ant(2″)-Ia (Table 2). Their frequency is reported in Fig. 2.
Frequency (log scale) of the aminoglycoside resistance determinants aac(3)-Ia and ant(2″)-Ia in the 78 CF P. aeruginosa strains resistant to one or more aminoglycosides. The number of strains carrying each resistance gene is reported as a proportion of the P. aeruginosa isolates resistant to the same antibiotic(s). GEN gentamicin, AMK amikacin, TOB tobramycin
As expected, all P. aeruginosa isolates carrying aac(3)-Ia were resistant to gentamicin alone (16.7%) or to gentamicin+amikacin (8.57%). ant(2″)-Ia was detected in 50% of strains resistant to tobramycin or tobramycin+gentamicin and in 10% of strains resistant to amikacin, in 9.52% of those resistant to tobramycin+amikacin+gentamicin and in 2.86% of those resistant to gentamicin+amikacin.
The gentamicin, amikacin and tobramycin MIC values of the 11 strains carrying the additional resistance genes aac(3)-Ia or ant(2″)-Ia were further investigated by broth microdilution using a wider range of antibiotic concentrations and the specific resistance phenotype checked against the presence of aac(3)-Ia or ant(2″)-Ia (Table 2).
Notably, the two P. aeruginosa strains (C44 and AR86) with the highest tobramycin MIC (256 μg/ml) harboured ant(2″)-Ia, as well as the strain with the highest gentamycin MIC (>2048 μg/ml).
3.3 PFGE Typing of the Aminoglycoside-Resistant P. aeruginosa Strains Carrying aac(3)-Ia or ant(2″)-Ia
The 11 P. aeruginosa strains carrying aac(3)-Ia or ant(2″)-Ia were typed by PFGE (Fig. 3), which highlighted two main clusters with a similarity <55%. One cluster encompassed the seven isolates encoding ant(2″)-Ia (top) and the other the four isolates encoding aac(3)-Ia (bottom). Within the former cluster, five isolates, of which four were from the same patient (Table 2), belonged to the same clone (100% similarity) and displayed a similarity <65% to the other 2 isolates. The 4 strains of the second cluster belonged to 4 different pulsotypes and shared a low similarity (about 60%) although 2 of them, represented by C40 and AR39, seem more related each other showing a similarity of 75%.
4 Discussion
A major issue in the control and management of CF P. aeruginosa lung infection is bacterial unresponsiveness to antibiotic treatment. This is due to a variety of causes, including chromosome-encoded resistance, biofilm development and the increasing diffusion of XDR strains (Colque et al. 2020). AMGs are frontline antibiotics, especially tobramycin, which is included in the eradication protocols of chronic lung infections (Ratjen et al. 2019). This work aimed to investigate the prevalence of resistance to tobramycin, gentamicin and amikacin (the three AMGs routinely tested in CF samples) and of the additional resistance genes rmtA, ant(2″)-Ia, aac(3)-Ia and aph(3′)-IIa in the CF P. aeruginosa strains isolated from sputum samples collected at the Marche Cystic Fibrosis centre. The high prevalence (53%) of AMG-resistant strains among these isolates highlights the need for measures to contain the spread of P. aeruginosa antibiotic resistance and for novel therapeutic approaches. Resistance to gentamicin and amikacin was the most frequent (44.87%) AMG resistance phenotype and was followed by resistance to all three AMGs (26.92%), to amikacin (12.82%), to gentamicin (7.69%), to gentamicin and tobramycin (5.13%) and to tobramycin alone (2.56%); therefore the proportion of strains resistant to two or all three AMGs was higher (76.92%) than that of strains resistant to a single antibiotic (23.08%). These findings suggest the major involvement of a cross-resistance mechanism capable of counteracting different AMGs. The most likely seems to be MexXY-OprM EP upregulation; this has been described as the main AMG resistance mechanism in P. aeruginosa (López-Causapé et al. 2018) through mutations of its regulatory gene mexZ, although mutations of hot-spot genes, like fusA1, could also be responsible for the multi-resistant phenotype. The lack of additional resistance genes in the strains showing multi-AMG resistance in our study, supports these views. Notably, the susceptibility to tobramycin of 44.87% of strains resistant to gentamicin and amikacin and lacking additional AMG resistance genes, could be explained by mutations in the MexY inner membrane channel not affecting specifically the tobramycin binding site; this would be in line with our modelling results, which suggest that these polymorphisms can occur as a consequence of mexY single nucleotide substitutions (unpublished data). None of our 147 strains was resistant to amikacin and tobramycin and susceptible to gentamicin, reflecting the absence among them of resistance mechanisms capable of counteracting the antibacterial activity of amikacin and tobramycin without affecting gentamicin effectiveness.
Tobramycin has proved to be a better therapeutic option than gentamicin and amikacin, with 34.62% of resistant strains compared to 84.62% of the other two AMGs (Fig. 1). These data are in agreement with those reported by Mustafa et al. (2016) and support the use of tobramycin, rather than other AMGs, in the eradication protocols of early P. aeruginosa CF lung infection (Mostofian et al. 2019).
This study involved the examination of six genes to evaluate the chromosome-dependent (mexY and ndvB) or -independent (rmtA, ant(2″)-Ia, aac(3)-Ia and aph(3′)-IIa) nature of AMG resistance in our strains, and the assessment of the prevalence of antibiotic specific (ant(2)”-Ia, aac(3)-Ia and aph(3′)-IIa) or aspecific (mexY, ndvB and rmtA) resistance mechanisms. As expected, mexY and ndvB were detected in all AMG-resistant strains, the only exception being P. aeruginosa C36, which showed no ndvB amplification, likely due to mutations in the sequences targeted by the ndvB primer pair. Since ndvB is involved in resistance of biofilm-growing P. aeruginosa (Mah et al. 2003) and we used planktonic cultures in the antibiotic susceptibility assays, the overexpression of the mexXY-OprM gene cluster seems responsible for the AMG resistance of P. aeruginosa isolates lacking additional resistance genes and is consistent with the above considerations about the prevalence of multi-AMG resistance phenotypes. MexXY-OprM is involved in the adaptive low-level resistance of CF P. aeruginosa (Morita et al. 2012); high-level resistance may evolve as a consequence of the gradual accrual of mutations in the mexXY-oprM regulator mexZ, which has often been reported in patients in chronic P. aeruginosa infections (Prickett et al. 2017). In such patients, the expression of the P. aeruginosa mexXY-oprM gene cluster progressively increases, also as a consequence of adaptation to the stressful conditions of the CF lung environment (Martin et al. 2018). This can explain the variable resistant phenotypes observed in the P. aeruginosa strains lacking the additional resistance determinants aac(3)-Ia and/or ant(2″)-Ia. Further investigation of the mexY expression level of these strains is warranted. We are currently evaluating the correlation between these phenotypes and mexY expression level.
Of the additional AMG resistance genes sought in this study – rmtA, aac(3)-Ia, aph(3′)-IIa and ant(2″)-Ia – only aac(3)-Ia and ant(2″)-Ia (Poole 2011) were detected and were found in only 11/78 strains, in line with the notion that AMG resistance in CF P. aeruginosa isolates is mainly a consequence of mutational events (López-Causapé et al. 2018). As expected, the four strains harbouring aac(3)-Ia were all resistant to gentamicin and susceptible to tobramycin (Ramirez and Tolmasky 2010), whereas two (P. aeruginosa AR88 and AR89) of the seven strains carrying ant(2″)-Ia, which confers resistance both to gentamicin and tobramycin (Poole 2005), were susceptible to the latter (MIC <4 μg/ml), likely due to the lack or downregulation of ant(2″)-Ia expression. These results are consistent with the hypervariability of CF P. aeruginosa (Qin et al. 2018).
PFGE typing assays of the 11 strains carrying aac(3)-Ia or ant(2″)-Ia demonstrated that the four aac(3)-Ia-carrying strains were unrelated, thus suggesting the spread of the GEN resistance gene among different P. aeruginosa clones through horizontal gene transfer (Kiddee et al. 2013). In contrast, five of the seven tobramycin-resistant isolates carrying the ant(2″)-Ia gene showed 100% similarity, suggesting the spread of a single clone. Since the 147 P. aeruginosa isolates were collected randomly and anonymously, we retrospectively investigated whether they came from the same patient. Indeed, four of them had been collected from patient B over an eight-month period (Table 2), thus highlighting failed eradication of P. aeruginosa, which induced symptom relapse. On the other hand, recovery of the same clone from different patients suggests a clonal spread of the tobramycin-resistant strain. The selective pressure exerted by the repeated use of tobramycin to counteract P. aeruginosa lung infection in CF patients (Smith et al. 2017) can explain the persistence and spread of this clone among patients referring to the CF centre. The role of additional resistance genes in P. aeruginosa persistence has been described by Mózes et al. (2014), who suggested a relationship between endemic P. aeruginosa clones and their carriage of integron-borne AMG resistance determinants. Ostensibly the fitness cost due to the additional gene is largely offset by the favourable conditions found inside the host.
In conclusion, P. aeruginosa AMG resistance is a major threat for CF patients, whose lung infections are routinely treated with these drugs. Our results show the spread of AMG resistance among the patients managed by the Marche regional Cystic Fibrosis centre, who showed a high prevalence of strains resistant to gentamicin, amikacin and tobramycin. The origin of resistance seems to be largely mutational, probably related to the MexXY-OprM EP or to other chromosome-encoded determinants. This suggests that MexXY-OprM EP inhibitors suitable for synergistic combinations with AMGs should urgently be developed to treat P. aeruginosa lung infections (Lamers et al. 2013; Aron and Opperman 2016; Laudadio et al. 2019). Our findings also suggest that monitoring additional AMG resistance genes, particularly ant(2″)-Ia, in early P. aeruginosa lung infection could contribute to a more effective management of CF patients.
References
Aron Z, Opperman TJ (2016) Optimization of a novel series of pyranopyridine RND efflux pump inhibitors. Curr Opin Microbiol 33:1–6
Beaudoin T, Zhang L, Hinz AJ et al (2012) The biofilm-specific antibiotic resistance gene ndvB is important for expression of ethanol oxidation genes in Pseudomonas aeruginosa biofilms. J Bacteriol 194:3128–3136
Biavasco F, Foglia C, Paoletti G et al (2007) VanA-type enterococci from humans, animals, and food: species distribution, population structure, Tn1546 typing and location, and virulence determinants. Appl Environ Microbiol 73:307–3319
Bolard A, Plésiat P, Jeannot K (2018) Mutations in gene fusA1 as a novel mechanism of aminoglycoside resistance in clinical strains of Pseudomonas aeruginosa. Antimicrob Agents Chemother 62:e01835–e01817
Buttini F, Balducci AG, Colombo G et al (2018) Dose administration maneuvers and patient care in tobramycin dry powder inhalation therapy. Int J Pharm 548:182–191
Colque CA, Albarracín Orio AG, Feliziani S et al (2020) Hypermutator Pseudomonas aeruginosa exploits multiple genetic pathways to develop multidrug resistance during long-term infections in the airways of cystic fibrosis patients. Antimicrob Agents Chemother 64:e02142-19
Costello SE, Deshpande LM, Davis AP et al (2018) Aminoglycoside-modifying enzymes and 16S ribosomal RNA methyltransferases-encoding genes among a global collection of gram-negative isolates. J Glob Antimicrob Resist 16:278–285
European Committee for Antimicrobial Susceptibility Testing (EUCAST) (2003) Determination of minimum inhibitory concentrations (MICs) of antibacterial agents by broth dilution. EUCAST discussion document E. Dis 5.1. Clin Microbiol Infect 9:1–7
European Committee on Antimicrobial Susceptibility Testing (EUCAST). Breakpoint tables for interpretation of MICs and zone diameters. EUCAST, Basel. Accessed 2016
Frimodt-Møller J, Rossi E, Haagensen JAJ et al (2018) Mutations causing low level antibiotic resistance ensure bacterial survival in antibiotic-treated hosts. Sci Rep 8:12512
Hall CW, Hinz AJ, Gagnon LB et al (2018) Pseudomonas aeruginosa biofilm antibiotic resistance gene ndvB expression requires the RpoS stationary-phase sigma factor. Appl Environ Microbiol 84:e02762–e02717
Hynes WL, Ferretti JJ, Gilmore MS et al (1992) PCR amplification of streptococcal DNA using crude cell lysates. FEMS Microbiol Lett 73:139–142
Kiddee A, Henghiranyawong K, Yimsabai J et al (2013) Nosocomial spread of class 1 integron-carrying extensively drug-resistant Pseudomonas aeruginosa isolates in a Thai hospital. Int J Antimicrob Agents 42:301–306
Lamers RP, Cavallari JF, Burrows LL (2013) The efflux inhibitor phenylalanine-arginine beta-naphthylamide (PAβN) permeabilizes the outer membrane of gram-negative bacteria. PLoS One 8:e60666
Laudadio E, Cedraro N, Mangiaterra G et al (2019) Natural alkaloid Berberine activity against Pseudomonas aeruginosa MexXY-mediated aminoglycoside resistance: In Silico and in Vitro studies. J Nat Prod 82:1935–1944
López-Causapé C, Cabot G, Del Barrio-Tofiño E et al (2018) The versatile mutational Resistome of Pseudomonas aeruginosa. Front Microbiol 9:685
Mah TF, Pitts B, Pellock B et al (2003) A genetic basis for Pseudomonas aeruginosa biofilm antibiotic resistance. Nature 426:306–310
Mangiaterra G, Laudadio E, Cometti M et al (2017) Inhibitors of multidrug efflux pumps of Pseudomonas aeruginosa from natural sources: an in silico high-throughput virtual screening and in vitro validation. Med Chem Res 26:414–430
Martin LW, Robson CL, Watts AM (2018) Expression of Pseudomonas aeruginosa antibiotic resistance genes varies greatly during infections in cystic fibrosis patients. Antimicrob Agents Chemother 62:e01789–e01718
Michalska AD, Sacha PT, Ojdana D et al (2014) Prevalence of resistance to aminoglycosides and fluoroquinolones among Pseudomonas aeruginosa strains in a University Hospital in Northeastern Poland. Braz J Microbiol 45:1455–1458
Morita Y, Tomida J, Kawamura Y (2012) MexXY multidrug efflux system of Pseudomonas aeruginosa. Front Microbiol 3:408
Mostofian F, Alkadri J, Tang K et al (2019) A real world evaluation of the long-term efficacy of strategies to prevent chronic Pseudomonas Aeruginosa pulmonary infection in children with cystic fibrosis. Int J Infect Dis 85:92–97
Mózes J, Szűcs I, Molnár D et al (2014) A potential role of aminoglycoside resistance in endemic occurrence of Pseudomonas Aeruginosa strains in lower airways of mechanically ventilated patients. Diagn Microbiol Infect Dis 78:79–84
Müller L, Murgia X, Siebenbürger L et al (2018) Human airway mucus alters susceptibility of Pseudomonas aeruginosa biofilms to tobramycin, but not colistin. J Antimicrob Chemother 73:2762–2769
Mustafa MH, Chalhoub H, Denis O et al (2016) Antimicrobial susceptibility of Pseudomonas Aeruginosa isolated from cystic fibrosis patients in northern Europe. Antimicrob Agents Chemother 60:6735–6741
Oh H, Stenhoff J, Jalal S et al (2003) Role of efflux pumps and mutations in genes for topoisomerases II and IV in fluoroquinolone-resistant Pseudomonas aeruginosa strains. Microb Drug Resist 9:323–328
Partridge SR, Kwong SM, Firth N et al (2018) Mobile genetic elements associated with antimicrobial resistance. Clin Microbiol Rev 31:e00088–e00017
Poole K (2005) Aminoglycoside Resistance in Pseudomonas aeruginosa. Antimicrob Agents Chemother 49:479–487
Poole K (2011) Pseudomonas aeruginosa: resistance to the max. Front Microbiol 2:65
Prickett MH, Hauser AR, McColley SA et al (2017) Aminoglycoside resistance of Pseudomonas Aeruginosa in cystic fibrosis results from convergent evolution in the mexZ gene. Thorax 72:40–47
Qin X, Zhou C, Zerr DM et al (2018) Heterogeneous antimicrobial susceptibility characteristics in Pseudomonas aeruginosa isolates from cystic fibrosis patients. mSphere 3:e00615–e00617
Ramirez MS, Tolmasky ME (2010) Aminoglycoside modifying enzymes. Drug Resist Updat 13:151–171
Ratjen F, Moeller A, McKinney ML et al (2019) Eradication of early P. aeruginosa infection in children <7 years of age with cystic fibrosis: the early study. J Cyst Fibros 18:78–85
Seifert H, Dolzani L, Bressan R et al (2005) Standardization and interlaboratory reproducibility assessment of pulsed-field gel electrophoresis-generated fingerprints of Acinetobacter baumannii. J Clin Microbiol 43:4328–4335
Smith WD, Bardin E, Cameron L et al (2017) Current and future therapies for Pseudomonas aeruginosa infection in patients with cystic fibrosis. FEMS Microbiol Lett 364(14):1–9
Soto SM (2013) Role of efflux pumps in the antibiotic resistance of bacteria embedded in a biofilm. Virulence 4:223–229
Stanojevic S, Waters V, Mathew JL et al (2014) Effectiveness of inhaled tobramycin in eradicating Pseudomonas aeruginosa in children with cystic fibrosis. J Cyst Fibros 13:172–178
Woegerbauer M, Zeinzinger J, Springer B et al (2004) Prevalence of the aminoglycoside phosphotransferase genes aph(3′)-IIIa and aph(3′)-IIa in Escherichia coli, Enterococcus faecalis, Enterococcus faecium, Pseudomonas aeruginosa, Salmonella enterica subsp. enterica and Staphylococcus aureus isolates in Austria. J Med Microbiol 63:210–217
Yamane K, Doi Y, Yokoyama K et al (2004) Genetic environments of the rmtA gene in Pseudomonas aeruginosa clinical isolates. Antimicrob Agents Chemother 48:2069–2074
Acknowledgments
We are grateful to Dr. Esther Manso for providing the CF P. aeruginosa strains, to Dr. Francesca Andreoni for sequencing analysis, to Prof. Luigi Ferrante for the statistical analysis, and to Marina Lombardi and Natascia Gracciotti for their technical assistance.
Author Disclosure Statement
No competing financial interests exist.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
1 Electronic Supplementary Material (S)
Fig. S1
Frequency of strains resistant to gentamicin (GEN), amikacin (AMK) and tobramycin (TOB) or to their combinations among the 78 aminoglycoside-resistant CF P. aeruginosa isolates. * p < 0.05; ** p < 0.01 (JPG 47 kb)
Supplementary Table 1
Gentamycin (GEN), amikacin (AMK) and tobramycin (TOB) MICs and the relevant resistance genes detected in the 78 CF P. aeruginosa strains resulted resistant to at least one of the three antibiotics by the routine antibiotic susceptibility test. The breakpoints (EUCAST 2016) are reported below. Breakpoints: GEN and TOB: 4 ≤ =S, >4 = R; AMK:8 ≤ =S, >16 = R (DOCX 27 kb)
Rights and permissions
Copyright information
© 2020 Springer Nature Switzerland AG
About this chapter
Cite this chapter
Mangiaterra, G., Cedraro, N., Citterio, B., Simoni, S., Vignaroli, C., Biavasco, F. (2020). Diffusion and Characterization of Pseudomonas aeruginosa Aminoglycoside Resistance in an Italian Regional Cystic Fibrosis Centre. In: Donelli, G. (eds) Advances in Microbiology, Infectious Diseases and Public Health. Advances in Experimental Medicine and Biology(), vol 1323. Springer, Cham. https://doi.org/10.1007/5584_2020_570
Download citation
DOI: https://doi.org/10.1007/5584_2020_570
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-030-71201-3
Online ISBN: 978-3-030-71202-0
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)