Abstract
Direct lineage reprogramming is the conversion of one specialized cell type to another without the need for a pluripotent intermediate. To date, a wide variety of cell types have been successfully generated using direct reprogramming, both in vitro and in vivo. These newly converted cells have the potential to replace cells that are lost to disease and/or injury. In this chapter, we will focus on direct reprogramming in the central nervous system. We will review current progress in the field with regards to all the major neural cell types and explore how cellular heterogeneity, both in the starter cell and target cell population, may have implications for direct reprogramming. Finally, we will discuss new technologies that will improve our understanding of the reprogramming process and aid the development of more specific and efficient future CNS-based reprogramming strategies.
Access provided by Autonomous University of Puebla. Download chapter PDF
Similar content being viewed by others
Keywords
- Cellular reprogramming
- Direct lineage conversion
- Cellular heterogeneity
- Neurological disease/Injury
- Central nervous system
1 Introduction
Historically, it was believed that cell fate was fixed after the completion of development (Heins et al. 2002; Barker et al. 2018; Faiz and Nagy 2013; Vierbuchen and Wernig 2011). However, discoveries including cell fusion, somatic nuclear transfer, and most recently reprogramming to pluripotency (or the generation of induced pluripotent stem cells, iPSCs) have shown that cell fate is flexible (Faiz and Nagy 2013; Vierbuchen and Wernig 2011; Gurdon 1962; Chen et al. 2015; Graf and Enver 2009; Blau et al. 1983, 1985; Xie et al. 2004; Takahashi and Yamanaka 2006). In this review, we will focus on direct lineage reprogramming, which is the conversion of one specialized cell type to another (Graf and Enver 2009; Xu et al. 2015; Wang and Zhang 2018; Gascón et al. 2017a; Masserdotti et al. 2016) (Fig. 1). This was first demonstrated by Davis and colleagues, who showed that overexpression of MyoD resulted in the conversion of fibroblasts to myoblasts (Davis et al. 1987). More recently, a number of studies have demonstrated successful conversion of various other cell types, both in vitro and in vivo (for review, see (Barker et al. 2018; Chen et al. 2015; Xu et al. 2015; Wang and Zhang 2018; Gascón et al. 2017a; Masserdotti et al. 2016)). This ground-breaking technology has had a significant impact on the field of regenerative medicine, as directly reprogrammed cells could be used to replace those lost or damaged to disease or injury (Barker et al. 2018; Faiz and Nagy 2013; Chen et al. 2015; Graf and Enver 2009; Takahashi and Yamanaka 2006; Xu et al. 2015; Wang and Zhang 2018; Gascón et al. 2017a; Masserdotti et al. 2016).
Direct lineage reprogramming. Direct lineage reprogramming is the conversion of one specialized cell type (Cell A) to another (Cell B) without the need for a pluripotent intermediate. It can be initiated by a variety of methods (small molecules, microRNAs), but is typically achieved by the overexpression of transcription factors. Illustrated by Kayla Hoffman-Rogers
Direct lineage conversion uses the delivery of specific factors to induce the conversion of cells without the need for a pluripotent intermediate (Graf and Enver 2009; Xu et al. 2015; Wang and Zhang 2018; Gascón et al. 2017a; Masserdotti et al. 2016). Typically, transcription factors have been used, but the feasibility of using small molecules (Hu et al. 2015; Li et al. 2015), microRNAs (Yoo et al. 2011; Victor et al. 2014), and CRISPRa (Chakraborty et al. 2014; Black et al. 2016) (Fig. 1) has also been demonstrated. To date, most studies have identified reprogramming factors based on their role in specifying a target cell fate during development, and/or uniquely high gene expression in a target cell. For example, Najm and colleagues used microarray data from different central nervous system (CNS) cells to identify a pool of genes that were exclusively upregulated in oligodendrocytes (Najm et al. 2013). These genes were then tested for their ability to convert fibroblasts to oligodendrocytes (Najm et al. 2013).
Many studies have focused on identifying “core” factors that are needed for cellular conversion using a reductionist-additive approach (Ninkovic and Götz 2018). In this paradigm, one factor is removed at a time until the “necessary” factor(s) are found (Ninkovic and Götz 2018). Additional factors are then added back until a desired phenotype or efficiency is achieved (Ninkovic and Götz 2018). For example, following confirmation that a cocktail of eleven reprogramming factors was able to reprogram fibroblasts to motor neurons, Son and colleagues removed one transcription factor at a time and analyzed its effect on the conversion (Son et al. 2011). This allowed them to determine that Ascl1 or Lhx3 were crucial for fibroblast to neuron conversion (Son et al. 2011). Then, to determine the optimal combination of transcription factors for a motor neuron phenotype, they added back single transcription factors and identified a “core set” of seven (Ascl1/Brn2/Myt1l/Lhx3/Hb9/Isl1/Ngn2) (Son et al. 2011). This approach suggests that an end goal is to achieve reprogramming with the smallest number of factors. Indeed, the seminal study by Davis and colleagues used only MyoD – highlighting the feasibility of a single factor for direct lineage reprogramming (Davis et al. 1987). One transcription factor for reprogramming may be favorable for future clinical applications, both in terms of feasibility of delivery and patient safety and tolerability. While it has been argued that single-factor reprogramming results in immature cell phenotypes (Morris 2016), neuronal reprogramming strategies using only one factor have resulted in the generation of mature and functional neurons, albeit at times with a slower maturation rate (Chanda et al. 2014; Zhu et al. 2018; Heinrich et al. 2010; Guo et al. 2014).
Interestingly, single-factor lineage reprogramming highlights the ability of certain reprogramming factors to behave as “pioneers” (Ninkovic and Götz 2018). Pioneer factors can bind to closed areas of chromatin and recruit supporting transcription factors that may be needed to initiate the reprogramming process (Ninkovic and Götz 2018). Further, it has been suggested that the feasibility and efficiency of conversion using single factors may be due to their pioneer activity (Ninkovic and Götz 2018). For example, the pioneer factor Ascl1, may endogenously recruit other factors beneficial for fibroblast to neuron conversion, such as Brn2 and Myt1l (Ninkovic and Götz 2018). This ability to bind to closed areas of chromatin demonstrates one way in which a starting cell state can be overridden; as in development, the genes regulating alternate cell fates are epigenetically repressed via chromatin modifications (Ninkovic and Götz 2018). Although a valuable insight into how reprogramming is initiated, many of the mechanisms that drive direct reprogramming have yet to be fully elucidated. It has been suggested that this is a complex process, dependent on many variables, including chromatin remodeling (Ninkovic and Götz 2018; Wapinski et al. 2017) and metabolic changes (Gascón et al. 2016, 2017b), amongst others (see (Xu et al. 2015; Wang and Zhang 2018; Gascón et al. 2017a, b; Masserdotti et al. 2015, 2016; Gascón et al. 2016) for a comprehensive review).
While the mechanisms of reprogramming remain unclear, the applicability of direct reprogramming technology is unmistakable. Direct lineage conversion has been used in many tissue systems and provides a novel therapeutic option for drug-resistant diseases or diseases with no current treatment options (Xu et al. 2015; Berninger 2010). In this review we will use the neural lineage as a model system to explore direct lineage reprogramming. Most studies have focused on direct reprogramming to neurons (reviewed in (Chen et al. 2015; Xu et al. 2015; Wang and Zhang 2018; Gascón et al. 2017a; Masserdotti et al. 2016)), because of the significant loss or injury to these cells in most neurological conditions. However, other neural lineage cells, for example, oligodendrocytes, may also be of interest. We will discuss the progress and current state of the field of direct lineage reprogramming with regards to all the major CNS cell types. We will explore how cellular heterogeneity, both in the starter cell population and the target cell type, may have implications for direct reprogramming. Finally, we will discuss new technologies that will improve our understanding of direct reprogramming and development of future conversion strategies.
2 Direct Reprogramming to a Neural Cell Fate
2.1 Overview
The first report of direct reprogramming to cells of the neural lineage used the transcription factor Pax6 to convert astrocytes to neurons in vitro (Heins et al. 2002). Subsequent studies showed that the delivery of other transcription factors, such as Ascl1 (Chanda et al. 2014), Brn2 (Zhu et al. 2018) and Ngn2 (Heinrich et al. 2010), could also convert astrocytes to neurons in vitro. Direct conversion has also been used to generate other neural cells such as oligodendrocytes (Najm et al. 2013; Yang et al. 2013; Mokhtarzadeh Khanghahi et al. 2018) and astrocytes (Caiazzo et al. 2015; Tian et al. 2016). It has also been shown that a wide variety of cell types, including those of a non-neural lineage, can be converted to the neural lineage. Fibroblasts and hepatocytes, two examples of non-neural cells, were successfully reprogrammed to neurons using a combination of Brn2/Mytl1/Ascl1 (Vierbuchen et al. 2010; Marro et al. 2011). There are both advantages and disadvantages in using cells that belong to non-neural lineages as a source population for reprogramming. Veritably, it broadens the potential scope of direct reprogramming, as it does not limit choice of a starting cell type. Conversely, neural lineage cells, such as astrocytes, may already have relevant epigenetic marks and active transcription factors, that may result in easier reprogramming (Faiz and Nagy 2013; Ninkovic and Götz 2018). Thus, future studies must include a functional comparison of cells that are generated from neural versus non-neural starter populations.
Following initial in vitro studies, a number of reports demonstrated in vivo reprogramming in the brain and spinal cord. This is of particular interest for brain repair, as it enables the targeted generation of new cells at the site of injury and circumvents the need for transplantation of exogenous cells and the associated risks, namely immune-rejection and the potential for cell mutagenesis from long-term cell culture (Faiz and Nagy 2013; Xu et al. 2015; Gascón et al. 2017a). It also provides an alternative to strategies using endogenous neural stem cells that reside within the brain and spinal cord. Attempts to generate neurons from these neural stem cells have resulted in low differentiation into the proper mature neuronal phenotypes, and poor long-term survival (Barker et al. 2018; Gascón et al. 2017a; Arvidsson et al. 2002; Thored et al. 2007).
In 2005, Buffo and colleagues demonstrated for the first time in the CNS that the manipulation of transcription factors could alter cell fate in vivo (Buffo et al. 2005). They converted NG2 glia into cells of a neuronal phenotype by inhibiting the expression of Olig2 (Buffo et al. 2005). This inhibition was achieved through the specific delivery of the dominant negative form of Olig2 to NG2 glia (Buffo et al. 2005). In vivo direct conversion has now been shown in the healthy brains of both young and old mice (Niu et al. 2013; Rouaux and Arlotta 2013). Of clinical relevance, the success of in vivo direct reprogramming has also been demonstrated in a number of models of CNS injury and disease, including stroke (Faiz et al. 2015), stab wound injury (Chen et al. 2015; Guo et al. 2014; Heinrich et al. 2014), spinal cord injury (Su et al. 2014), Alzheimer’s disease (Chen et al. 2015; Guo et al. 2014) and Parkinson’s disease (Rivetti di Val Cervo et al. 2017). Interestingly, it has been suggested that aspects of the injured/diseased environment, such as the increase of beneficial growth factors, increased plasticity of glial cells and increased glycolysis may actually enhance the reprogramming process (Gascón et al. 2017a; Guo et al. 2014; Grande et al. 2013). These disease-induced changes could explain why some reprogramming paradigms have encountered success in an injury context, but no conversion (or a significantly reduced conversion) was observed when the same transcription factor(s) were delivered to the uninjured brain (Heinrich et al. 2014; Grande et al. 2013). Conversely, it has also been noted that an increased production of reactive oxygen species (ROS) during injury could be deleterious to newly generated cells and explain the discrepancy in conversion success between in vitro and in vivo studies (Gascón et al. 2017a). A better understanding of the mechanisms that underlie each particular injury or disease model will allow for reprogramming strategies that are tailored and optimized for different applications.
2.2 Target Cell Type
Many neurological disorders or conditions have at their core, a significant loss or injury to the cells of the CNS. However, not all disorders implicate the same cells and as such, it is important to generate specific cell types that are needed for a particular disorder. The versatility of direct lineage reprogramming technology is clear – studies have shown the generation of all the main cell types of the CNS, including certain subtypes and progenitors.
2.2.1 Neurons
Generating Neurons In Vitro
Neurons are affected in a wide variety of neurological conditions, and thus direct lineage reprogramming strategies have mainly been focused on regenerating these cells. Since their seminal Pax6 study, work from Magdalena Götz’s lab has also demonstrated that a combination of Ascl1/Dlx2 or Ngn2 results in the conversion of astrocytes to GABAergic and glutamatergic neurons, respectively (Vierbuchen and Wernig 2011; Xu et al. 2015; Heinrich et al. 2010). Simultaneously, work done by Vierbuchen and colleagues established the ability of the combination of Ascl1/Brn2/Mytl1 (referred to as BAM factors) to induce glutamatergic neurons from fibroblasts (Vierbuchen et al. 2010). The conversion of glial cells (both astrocytes and NG2 glia) to neurons using NeuroD1 by Gong Chen’s lab further demonstrated that transcription factors involved in later stages of neuronal development could also be used to regenerate neurons (Guo et al. 2014).
A number of other starter cell types have also been successfully converted to neurons. Non-neural cell types, such as pericytes, have been reprogrammed to glutamatergic and GABAergic cells (Karow et al. 2018) and the BAM factors have been used to reprogram hepatocytes to glutamatergic-like neuronal cells (Marro et al. 2011). Additionally, it has been shown that microglia can be converted to functional neurons with the delivery of NeuroD1 alone (Matsuda et al. 2019).
Importantly, the type of neuron lost or affected in a particular disease is often of a specific subtype (i.e.: dopaminergic neurons in Parkinson’s disease and motor neurons in Amyotrophic Lateral Sclerosis), and differs across various neurological conditions (Faiz and Nagy 2013; Chen et al. 2015; Xu et al. 2015; Wang and Zhang 2018; Masserdotti et al. 2016). As such, the generation of a random assortment of neuronal subtypes, or the ability to generate only one specific subtype would likely be of minimal therapeutic benefit. For example, generating cholinergic neurons in Alzheimer’s disease is likely to confer more benefit than in Parkinson’s disease, where dopaminergic neurons are needed. Direct reprogramming must therefore reliably generate subtype specific cell types appropriate for the neurological deficit in question (Faiz and Nagy 2013; Chen et al. 2015; Xu et al. 2015; Wang and Zhang 2018; Masserdotti et al. 2016). Accordingly, in vitro studies have shown the generation of dopaminergic (Rivetti di Val Cervo et al. 2017; Kim et al. 2011; Caiazzo et al. 2011; Sheng et al. 2012), motor (Son et al. 2011), serotonergic (Vadodaria et al. 2016), and cholinergic (Liang et al. 2018; Liu et al. 2013) neurons, amongst others (see (Masserdotti et al. 2016) for in depth review) using specific combinations of transcription factors.
In summary, direct lineage reprogramming in vitro is clearly feasible, customizable and reliable in generating new neurons. However, in vitro lineage conversion still requires transplantation into the brain.
Generating Neurons In Vivo
One of the most exciting features of direct reprogramming is the ability to target endogenous cells at their source. Thus, in vivo studies generating novel populations of neurons are of particular interest to the field. Work performed by a number of groups has shown the reliable generation of new neurons in vivo using direct reprogramming in healthy and injured environments, and has been extensively reviewed elsewhere.(Chen et al. 2015; Xu et al. 2015; Wang and Zhang 2018; Gascón et al. 2017a; Masserdotti et al. 2016) What is lacking and of significant interest however, is a systematic comparison of different transcription factors and delivery strategies in various models of disease and injury (Gascón et al. 2017a). Although the transcription factors used in these studies (Sox2 (Niu et al. 2013; Heinrich et al. 2014), BAM factors (Torper et al. 2013), NeuroD1 (Guo et al. 2014) and Ascl1/Lmx1a/Nurr1 (Torper et al. 2015)) correspond to in vitro studies, there is variation with regards to the delivery system used. It has been proposed that the choice of delivery system may affect the reprogramming paradigm, as there is variance in their temporal kinetics (Gascón et al. 2017a). As such, clear conclusions on the “best” direct reprogramming paradigm for a particular starting cell type, target cell type or disease state cannot yet be made with certainty. Nonetheless, these newly generated neurons are capable of surviving, maturing and integrating into the pre-existing neural circuitry, as shown by electrophysiological and functional assays (Guo et al. 2014; Niu et al. 2013; Heinrich et al. 2014; Torper et al. 2013, 2015).
One hurdle that remains with regards to in vivo neuronal reprogramming is subtype specific neuronal regeneration. Success seen in in vitro studies of neuronal subtype generation has not been replicated to the same extent in vivo, even with the use of the same transcription factors (Chen et al. 2015; Xu et al. 2015; Wang and Zhang 2018; Gascón et al. 2017a; Masserdotti et al. 2016). The reasons for this are unclear, but as discussed above, could be attributed to the deleterious environment that results from injury (Gascón et al. 2017a). A more complex in vivo environment may require multiple transcription factors and/or a combination of both transcription factors and small molecules or microRNA to generate specific neuronal sub-types. In fact, Rivetti di Val Cervo and colleagues successfully obtained a novel population of dopaminergic neurons from astrocytes in vivo when they used a combination of both NeuroD1/Ascl1/Lmx1a and the microRNA miR218 (Rivetti di Val Cervo et al. 2017).
Neuron to Neuron Reprogramming
While most studies have focused on converting non-neuronal cells to neurons, reports of neuron to neuron reprogramming show that there is cell fate flexibility even within this population. In the cortex, Rouaux and Arlotta were able to successfully convert layer 2/3 callosal projection neurons into layer 5/6 corticofugal projections using only the transcription factor Fezf2 (Rouaux and Arlotta 2013). More recently, Niu and colleagues used a combination of Sox2/Nurr1/Foxa2/Lmx1a paired with valproic acid (VPA) to reprogram striatal neurons to dopaminergic neurons (Niu et al. 2018). Interestingly, they showed that these newly induced dopaminergic cells arise directly from the striatal neurons, without passing through a progenitor stage (Niu et al. 2018). These studies beg the question of whether a shared identity (i.e. neuron) between the starting and target cell is an important consideration for easily generating specific neuronal subtypes in vivo.
2.2.2 Oligodendrocytes
Oligodendrocytes play crucial roles in maintaining proper cell signaling in the CNS and many diseases result from their widespread loss or malfunction. Oligodendrocyte death and subsequent de-myelination is characteristic to the pathology of multiple sclerosis (Lassmann et al. 2012; Reich et al. 2018; Sawcer et al. 2014), and a reduction in myelin is seen in multi-system atrophy (Burn and Jaros 2001). Oligodendrocytes have also been implicated in Alzheimer’s disease. Although traditionally thought of as a grey matter disease, Alzheimer’s disease presents with white matter disruption, impaired myelination patterns and decreased oligodendrocyte and oligodendrocyte progenitor gene expression (Desai et al. 2009, 2010; Roth et al. 2005). Interestingly, a mouse model of Alzheimer’s disease showed that impaired myelination and decreased CNPase and MBP expression precedes the onset of tau and amyloid pathology (Desai et al. 2009). Finally, white matter damage and oligodendrocyte dysfunction have been proposed as a risk factor and predictor of stroke (Kuller et al. 2004) and of schizophrenia (Cassoli et al. 2015).
Given the significant implication of oligodendrocytes in disease, strategies to restore or replenish damaged or lost oligodendrocytes are needed. Yet, there is a clear disparity in the number of studies investigating direct reprogramming to neurons versus oligodendrocytes. Only three studies to date have looked at using direct reprogramming to generate new populations of oligodendrocytes and their precursors. Work done by Najm and colleagues, as well as Yang and colleagues produced oligodendrocyte progenitor cells (OPCs) and oligodendrocytes from fibroblasts in vitro using combinations of transcription factors involved in OPC development and oligodendrocyte function (Sox10/Olig2/Nkx6.2 and Sox10/Olig2/Zfp536, respectively) (Najm et al. 2013; Yang et al. 2013). The oligodendrocytes generated from both these studies expressed characteristic OPC and oligodendrocyte markers and showed myelination capability in transplantation experiments (Najm et al. 2013; Yang et al. 2013). More recently, Khanghahi and colleagues reported that both in vitro and in vivo delivery of Sox10 alone to astrocytes in cuprizone induced de-myelinated mice results in the generation of new oligodendrocyte-like cells (Mokhtarzadeh Khanghahi et al. 2018). Cells transduced in vitro expressed markers of OPC and oligodendrocyte lineage and were transplanted into the corpus callosum of the cuprizone mice (Mokhtarzadeh Khanghahi et al. 2018).
Although promising, there are significantly fewer studies of oligodendrocyte reprogramming in comparison with neuronal reprogramming. Future work examining oligodendrocyte generation and the optimal factors involved are warranted.
2.2.3 Astrocytes
Most reprogramming studies have focused on astrocytes as the starter cell type, rather than the target cell type. Nonetheless, a few reports have shown the feasibility of generating astrocytes. From a pool of 14 genes involved in determining astrocyte fate, Caiazzo and colleagues found that the combination of NFIA/NFIB/Sox9 could successfully reprogram fibroblasts to astrocytes (Caiazzo et al. 2015). Similarly, work by Tian and colleagues demonstrated that a cocktail of 6 small molecules generated functional astrocytes from fibroblasts (Tian et al. 2016). Given the recent knowledge that a subset of astrocytes, A2 cells, are neuroprotective and conducive to recovery following injury, future studies examining conversion of a starter cell to a beneficial astrocyte subtype, such as the A2 phenotype, may be of interest (Liddelow and Barres 2017; Liddelow et al. 2017; Zamanian et al. 2012; Toft-Hansen et al. 2011).
2.2.4 Stem/Progenitor Cells
To date, studies have generated both neural stem cells and glial progenitors that can give rise to mature neurons and glia. Researchers in the Wernig lab demonstrated that the use of two transcription factors, FoxG1 and Brn2, that are important for neural stem cell (NSC) fate, could convert fibroblasts into tripotent NSCs (Lujan et al. 2012). Not only were these NSCs capable of further differentiation into functional neurons, astrocytes and oligodendrocytes, but they also demonstrated clear proliferative capacity, capable of being passaged up to 17 times, without loss of function (Lujan et al. 2012). Additionally, other studies by Han and colleagues, as well as Ring and colleagues have shown conversion of fibroblasts to NSCs using a combination of Brn2/Sox2/Klf4/c-Myc/E47 and Sox2 alone, respectively (Xu et al. 2015; Han et al. 2012; Ring et al. 2012). Similarly, astrocytes have also been successfully converted to NSPCs through delivery of both single factors (OCT4, SOX2 or NANOG) (Corti et al. 2012) and combination of factors (Foxg1/Sox2/Brn2) (Ma et al. 2018). Finally, OPCs have also been generated in the Tesar lab using a combination of the transcription factors Sox10/Olig2/Nkx6.2 (Najm et al. 2013). The pathophysiology of a particular disease may determine whether it is advantageous to induce progenitor populations or mature cell types. Given this, future studies comparing the functional outcomes of direct reprogramming to progenitors versus mature cells will be of interest.
2.2.5 Cell Heterogeneity
It is now clear that neurons are not the only cell type in the CNS with delineated subtypes comprising different roles, with recent work demonstrating intra-cellular differences in oligodendrocyte populations. Using single-cell RNA sequencing (sc RNA-seq), Marques and colleagues found unique transcriptome profiles according to the age and region of oligodendrocytes and progenitors in mice (Marques et al. 2016). In addition, they noted a novel population of cells (ITPR2+) hypothesized to be involved in periods of rapid myelination (Marques et al. 2016). Furthermore, they noted that varying regions of the CNS were associated with differing forms of mature oligodendrocytes, such as MOL6 oligodendrocytes specific to the S1 cortex and corpus callosum (Marques et al. 2016). Differences in mature oligodendrocytes in the CNS are also present in the context of disease. Work by Jäkel and colleagues showed that not only is there a similar heterogeneity of oligodendrocytes in humans, but that MS patients had a unique loss of certain mature oligodendrocyte populations (OLIG1+) when compared to controls (Jäkel et al. 2019). These findings will be of particular importance when considering transcription factor cocktails used to create functional, myelinating oligodendrocytes and how the transcription factor combinations may vary based on disease need.
2.3 Starting Cell Type
When performing direct lineage reprogramming, genetic systems with cell specific promoters can allow for targeting of a precise cellular population (Wang and Zhang 2018). Traditionally, starter cell populations have been chosen based on their developmental closeness to the target cell type (Gascón et al. 2017a; Masserdotti et al. 2016; Waddington 1957). However, developmental closeness may not be the only, or even “best” reason for choosing a particular starting cell type. In the context of injury or disease, it may be more relevant to choose a starting cell based on the role of that cell at the time of reprogramming. For example, cells that contribute to ongoing neuronal death and therefore disease pathology may be the most clinically relevant choice for reprogramming.
2.3.1 Developmental Closeness
The Waddington model, used to explain normal cell fate determination, denotes a linear differentiation and restriction pattern of cell type during development (Waddington 1957). It was hypothesized that more closely related starter and target cells would be easier to convert and reprogram (Gascón et al. 2017a; Masserdotti et al. 2016). Initial studies using starting cell populations that belonged to a non-neuronal lineage, such as fibroblasts, required a combination of transcription factors (Son et al. 2011; Vierbuchen et al. 2010; Kim et al. 2011; Caiazzo et al. 2011; Sheng et al. 2012) for conversion to neurons. In contrast, starting cell populations within the neural lineage, such as astrocytes, could be successfully converted with just one transcription factor (Gascón et al. 2017a; Heinrich et al. 2010; Guo et al. 2014; Niu et al. 2013). However, more recent work has demonstrated the use of single factors to convert non-neural cells neurons. Chanda and colleagues demonstrated the generation of neurons from fibroblasts using only Ascl1 (Chanda et al. 2014) and MYT1L alone has been shown to reprogram pericytes into mature cholinergic neurons (Liang et al. 2018). Given these findings, a new model of reprogramming has been proposed – the Cook’s Island model (Masserdotti et al. 2016; Sieweke 2015). In this analogy, the starting cell is a boat and target cell types are the various islands to which the boat can travel (Masserdotti et al. 2016; Sieweke 2015). The boat may face various challenges or hurdles depending on the proximity of the island, but all are potentially accessible (Masserdotti et al. 2016; Sieweke 2015).
2.3.2 Cellular Heterogeneity
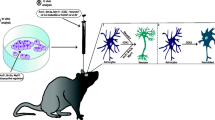
Cell heterogeneity within the starter cell population is of particular interest to direct lineage reprogramming. One specific subtype of astrocyte, for instance, may be especially conducive to generating a particular subtype of neuron (Fig. 2). Conversely, as we broaden our scope of potential cells that can be generated by reprogramming, it will be important that those generated are of the correct subtype for the particular disease or injury at hand.
Cellular heterogeneity. Heterogeneity of both the starting and target cell populations is an important consideration for direct lineage reprogramming. Diversity of the starting cell population may determine what types of target cell types are generated. All subsets of starting cells may give rise to one only type of target cell (a). Alternately, specific subtypes of starting cells may only give rise to specific subtypes of target cells (b). Or, only one type of starting cell may generate all target cell subtypes (c). Illustrated by Kayla Hoffman-Rogers
Astrocytes
Recent work by Liddelow and colleagues has shown that astrocytes have at least two defined functional states in the context of disease/injury, termed A1 and A2 (Liddelow and Barres 2017; Liddelow et al. 2017). A1 cells are present in many disease states, including Alzheimer’s Disease, Huntington’s Disease, Parkinson’s Disease and Multiple Sclerosis (Liddelow and Barres 2017; Liddelow et al. 2017). Furthermore, A1 astrocytes lose many normal astrocytic functions, such as phagocytic capacity and the promotion of synaptic formation and become toxic, killing neurons and oligodendrocytes, and impairing oligodendrocyte progenitor cell (OPC) differentiation (Liddelow and Barres 2017; Liddelow et al. 2017). Conversely, A2 cells are thought to be neuroprotective (Liddelow and Barres 2017). They upregulate a number of neurotrophic factors, cytokines and thrombospondins that may help repair and rebuild lost synapses (Liddelow and Barres 2017; Zamanian et al. 2012). In addition, it has also been postulated that there are many more astrocyte subtypes that have yet to be characterized (Liddelow and Barres 2017).
Given the heterogeneity of astrocytes, it may prove advantageous to reprogram astrocyte subtypes that are detrimental to disease outcome or progression (such as A1 cells), over reprogramming protective subtypes (such as A2 cells) that could lead to worse disease outcomes (Liddelow and Barres 2017; Liddelow et al. 2017; Zamanian et al. 2012; Toft-Hansen et al. 2011). Furthermore, it would be worthwhile investigating whether A1 neurotoxic astrocytes could be reprogrammed to their more beneficial A2 counterparts. This has been suggested in work done by Gong Chen’s lab, which noted that astrocytes transduced in their NeuroD1 mediated astrocyte to neuron paradigm showed a reduction in A1 gene expression prior to their conversion to neurons (Zhang et al. 2018).
Microglia
Microglia have also been shown to have at least two distinct subtypes, termed M1 and M2, with more recent work suggesting that many sub-classes, or even a continuum of microglial states may exist (Liddelow and Barres 2017; Boche et al. 2013; Tang and Le 2016). These subtypes pertain to activation states that correspond with particular functions: M1 microglia are pro-inflammatory and potentially damaging to neighboring cells, whereas M2 microglia are involved in tissue repair and are immunosuppressive (Liddelow and Barres 2017; Boche et al. 2013). Interestingly, this activation pattern is thought to be dependent on the particular injury or disease state (Boche et al. 2013). In fact, work by Tang and Le have shown that changes in M1 and M2 microglia phenotype correspond to different stages of disease (Tang and Le 2016). This knowledge may be of particular relevance in future clinical applications of direct reprogramming, allowing for tailored paradigms based off disease progression.
2.4 Direct Reprogramming: Readouts
In order to ensure the clinical relevance and feasibility of direct reprogramming, there is a need to generate mature cells that can integrate into existing host circuitry, have long-term survival and perform proper cell functions (Barker et al. 2018; Xu et al. 2015; Wang and Zhang 2018; Gascón et al. 2017a; Berninger 2010; Yang et al. 2011). If the cells generated fail to meet these conditions, it is unlikely that they could be utilized as a novel therapy for neurological diseases.
2.4.1 Characterization of Target Cells
To characterize newly reprogrammed cells, many studies have examined the expression of cell type specific proteins and patterns of global gene expression that correspond to native cells (Barker et al. 2018; Faiz and Nagy 2013; Xu et al. 2015; Gascón et al. 2017a; Masserdotti et al. 2016; Yang et al. 2011). Some studies have also used a lack of gene/protein expression, of cells of unwanted lineages or of cells of the starting population to be indicative of proper cell conversion. For example, Niu and colleagues demonstrated that reprogrammed dopaminergic neurons expressed cell-type specific markers [DDC (DOPA Decarboxylase), VMAT2 (Vesicular monoamine transporter 2) and DAT (Dopamine transporter)], but also confirmed that the reprogrammed cells did not express markers associated with other neuronal subtypes (cholinergic or glutamatergic, using CHAT and GLUT1, respectively) (Niu et al. 2018).
Functional assays specific to the desired cell type are also important (Barker et al. 2018; Xu et al. 2015; Wang and Zhang 2018; Gascón et al. 2017a; Berninger 2010; Yang et al. 2011) (Fig. 3). For neurons, there are both general and subtype specific means of assessing neuronal function and integration (Yang et al. 2011). Patch-clamp recording can demonstrate whether reprogrammed neurons exhibit electrophysiological characteristics of neurons – their ability to fire action potentials and their synaptic patterns (Chen et al. 2015; Wang and Zhang 2018; Yang et al. 2011). Most studies to date have demonstrated mature, electrically active neurons, both in vivo and in vitro. The firing patterns of reprogrammed cells can also be compared to the expected firing patterns of native cells to assess similarity in function, as was done by Niu and colleagues (Niu et al. 2018). Furthermore, fluorescent reporters can be used to trace reprogrammed cells and assess the extent of their integration into host circuitry (Chen et al. 2015; Torper et al. 2015). For example, Torper and colleagues noted that in a mouse model of Parkinson’s disease, newly generated neurons did not migrate to alternate regions of the CNS, but rather integrated locally (Torper et al. 2015).
Functional target cells. The goal of direct lineage reprogramming is to generate functional cells for repair or regeneration. Newly generated cells must recapitulate the function of their endogenous counterparts (blue neurons, red oligodendrocytes). After astrocyte to neuron conversion, new neurons (yellow) must fire action potentials, form synapses and integrate into the exiting host circuitry (blue neurons) (a). Similarly, new oligodendrocytes (yellow) must generate myelin and ensheath existing neurons like native oligodendrocytes (red) (b). Illustrated by Kayla Hoffman-Rogers
For oligodendrocytes, the ability to myelinate is key. To characterize the reprogrammed OPCs or oligodendrocytes, studies have used mouse models of demyelination or impaired myelination (Najm et al. 2013; Yang et al. 2013; Mokhtarzadeh Khanghahi et al. 2018; Chernoff 1981; Blakemore 1972; Matsushima and Morell 2001). The Shiverer mouse, for example, lacks myelin basic protein (MBP) and consequently, the ability to form compact myelin (Chernoff 1981). It has been used in studies such as those done by Najm and colleagues to demonstrate the generation of MBP+ myelin following transplantation of OPCs that were directly reprogrammed from fibroblasts (Najm et al. 2013).
2.4.2 Functional Outcomes
Ultimately, the goal of direct reprogramming is to be a clinical treatment option for neurological diseases. Therefore, it is crucial to employ animal models of disease to determine whether reprogramming can induce functional recovery, slow disease progression or reverse disease progression/impairments all together (Chen et al. 2015; Xu et al. 2015). To date, there has been limited study of the outcomes of reprogramming with regards to disease progression or prevention in disease models. The first report of functional recovery was by Rivetti di Val Cervo and colleagues in a unilateral 6-hydroxydopamine (6-OHDA) mouse model of Parkinson’s disease (Rivetti di Val Cervo et al. 2017). Following astrocyte reprogramming to dopaminergic neurons, newly generated neurons were capable of rescuing gait deficits and dopamine-deficient circling behaviors (Rivetti di Val Cervo et al. 2017). A second report by Chen and colleagues, showed improvement in motor and fear memory deficits following ischemia (Chen et al. 2018).
2.4.3 Application of New Technologies
Many exciting and novel technologies have recently emerged that will benefit our understanding of the reprogramming process and cellular outcomes. A new tool for analyzing reprogrammed cell identity is the CellNet database (Xu et al. 2015). It identifies gene regulatory networks (GRNs) in reprogrammed cells, and therefore enables confirmation of reprogramming factor expression in target cells and the comparison of gene expression profiles of experimental and naive cells (Xu et al. 2015; Cahan et al. 2014; Morris et al. 2014). Most striking, however, is the utility of CellNet in predicting how reprogramming paradigms could be improved, which is based on incorrect expression of GRNs (Xu et al. 2015; Morris et al. 2014).
In a pioneering study by Cadwell and colleagues, electrophysiological and single cell RNA-seq readouts were combined to create Patch-seq technology (Cadwell et al. 2016). This technique results in the simultaneous acquisition of cell transcriptomes (sc RNA-seq) and electrophysiological information (Patch-clamp readings), thereby correlating changes in cell function and the transcriptome within a single cell (Cadwell et al. 2016). This is of particular interest, as changes in cell function can be predicted based on particular transcriptome patterns or modifications (Cadwell et al. 2016). This technology has already been used to identify and predict populations of highly functional reprogrammed neurons generated from iPSCs (Bardy et al. 2016). The application of Patch-seq in direct reprogramming studies is thus greatly warranted, as it would enable a more tailored approach for the creation of specific cell types by identifying reprogramming factors that produce bonafide reprogrammed cells.
CRISPR is another new technology that can be used to determine genes that are involved in cell fate changes and therefore, elucidate optimal transcription factor combinations for direct reprogramming paradigms (Liu et al. 2018). This strategy has unveiled novel factor(s) that result in the conversion of fibroblasts to neurons, such as Ezh2/Mecom, which were not traditionally thought to be key proneural genes, (Liu et al. 2018). As the field strives for subtype specific cell generation, as well as tailorable and translatable therapies, utilizing the power of CRISPR may be of great interest.
Finally, optogenetics and pharmacogenetics provide novel means by which to target populations of cells and manipulate their activity (Deisseroth 2011; Amamoto and Arlotta 2014; Steinbeck and Studer 2015). This technology can be used to specifically analyze whether newly generated cells directly contribute to functional recovery (Amamoto and Arlotta 2014; Steinbeck and Studer 2015). It can also be employed to potentiate the activity of reprogrammed cells (Amamoto and Arlotta 2014; Steinbeck and Studer 2015). Indeed, in a study done by Dell’Anno and colleagues, a DREADD pharmacogenic system was used to selectively activate reprogrammed dopaminergic neurons to enhance their activity (Dell’Anno et al. 2014). Researchers noted that upon activation of these cells, dopamine levels were increased, even up to 5 weeks following reprogramming, supporting the use of chemogenetics as an adjunct strategy to direct reprogramming (Dell’Anno et al. 2014).
3 Conclusions and Future Directions
The field of reprogramming is still in its infancy. Although significant progress has been made in our understanding of direct reprogramming in the CNS since the pioneering work done by Heins and colleagues (2002), new research into the mechanisms underlying direct reprogramming will allow us to tailor better strategies for brain repair. Studies that will determine the optimal starting cell types that are needed for the generation of functional target cells, as well as experiments that systematically compare the efficacy of different reprogramming paradigms are needed. Moreover, research into the impact of cellular heterogeneity in reprogramming will result in better designed reprogramming strategies that cater to a specific disease or injury state. Our progress will only become faster with the implementation of the novel, cutting technologies, such as CellNet and Patch-seq. Given the integrative and multi-faceted nature of direct reprogramming, it seems only fitting to employ an interdisciplinary approach as we move forward.
Abbreviations
- 6-OHDA:
-
6-hydroxydopamine
- Ascl1:
-
achaete-scute family bHLH transcription factor 1
- BAM factors:
-
combination of the transcription factors Ascl1, Brn2 and Mytl1
- Brn2:
-
POU Class 3 Homeobox 2
- CHAT:
-
Choline O-Acetyltransferase
- c-Myc:
-
cellular Myc
- CNP:
-
2′,3’-Cyclic Nucleotide 3’ Phosphodiesterase
- CNS:
-
central nervous system
- CRISPR:
-
clustered regularly interspaced short palindromic repeats
- CRISPRa:
-
CRISPR activation
- DAT:
-
Dopamine transporter
- DDC:
-
DOPA Decarboxylase
- Dlx2:
-
Distal-Less Homeobox 2
- DREADD:
-
Designer Receptors Exclusively Activated by Designer Drugs
- E47:
-
transcription factor 3
- Ezh2:
-
Enhancer Of Zeste 2 Polycomb Repressive Complex 2 Subunit
- Fezf2:
-
FEZ Family Zinc Finger 2
- Foxa2:
-
Forkhead Box A2
- FoxG1:
-
forkhead box G1
- GABA:
-
Gamma-amino butyric acid
- GLUT1:
-
glucose transporter protein type 1
- GRN:
-
gene regulatory network
- Hb9:
-
Motor Neuron And Pancreas Homeobox 1
- iPSC:
-
induced pluripotent stem cell
- Isl1:
-
Insulin gene enhancer protein ISL-1
- ITPR2:
-
Inositol 1,4,5-Trisphosphate Receptor Type 2
- Klf4:
-
Kruppel Like Factor 4
- Lhx3:
-
LIM Homeobox 3
- Lmx1a:
-
LIM Homeobox Transcription Factor 1 Alpha
- MBP:
-
myelin basic protein
- Mecom:
-
MDS1 And EVI1 Complex Locus
- miRNA:
-
microRNA
- MOL6:
-
mature oligodendrocytes expressing Grm3 (Glutamate Metabotropic Receptor 3) and Jph4 (Junctophilin 4)
- MS:
-
multiple sclerosis
- MyoD:
-
myogenic differentiation 1
- Myt1l:
-
myelin transcription factor 1 like protein
- NANOG:
-
Nanog Homeobox
- NeuroD1:
-
Neurogenic Differentiation Factor 1
- NFIA:
-
Nuclear Factor I A
- NFIB:
-
Nuclear Factor I B
- NG2 glia:
-
Neural/glial antigen 2 expressing glial cells
- Ngn2:
-
Neurogenin 2
- Nkx6.2:
-
NK6 Homeobox 2
- NSC:
-
neural stem cell
- NSPC:
-
neural stem and progenitor cells
- Nurr1:
-
Nuclear receptor related 1 protein
- OCT4:
-
octamer-binding transcription factor 4
- Olig1:
-
Oligodendrocyte Transcription Factor 1
- Olig2:
-
Oligodendrocyte Transcription Factor 2
- OPC:
-
oligodendrocyte progenitor cell
- Pax6:
-
Paired Box 6
- ROS:
-
reactive oxygen species
- S1 cortex:
-
primary somatosensory cortex
- sc RNA-seq:
-
single cell RNA sequencing
- Sox10:
-
SRY-Box 10
- Sox2:
-
SRY-Box 2
- Sox9:
-
SRY-box 9
- VMAT2:
-
Vesicular monoamine transporter 2
- VPA:
-
valproic acid
- Zfp536:
-
Zinc Finger Protein 536
References
Amamoto R, Arlotta P (2014) Development-inspired reprogramming of the mammalian central nervous system. Science (New York, NY) 343(6170):1239882. https://doi.org/10.1126/science.1239882
Arvidsson A, Collin T, Kirik D, Kokaia Z, Lindvall O (2002) Neuronal replacement from endogenous precursors in the adult brain after stroke. Nat Med 8(9):963–970. https://doi.org/10.1038/nm747
Bardy C, van den Hurk M, Kakaradov B, Erwin JA, Jaeger BN, Hernandez RV, Eames T, Paucar AA, Gorris M, Marchand C et al (2016) Predicting the functional states of human iPSC-derived neurons with single-cell RNA-seq and electrophysiology. Mol Psychiatry 21(11):1573–1588. https://doi.org/10.1038/mp.2016.158
Barker RA, Götz M, Parmar M (2018) New approaches for brain repair—from rescue to reprogramming. Nature 557(7705):329. https://doi.org/10.1038/s41586-018-0087-1
Berninger B (2010) Making neurons from mature glia: a far-fetched dream? Neuropharmacology 58(6):894–902. https://doi.org/10.1016/j.neuropharm.2009.11.004
Black JB, Adler AF, Wang H-G, D’Ippolito AM, Hutchinson HA, Reddy TE, Pitt GS, Leong KW, Gersbach CA (2016) Targeted epigenetic remodeling of endogenous loci by CRISPR/Cas9-based transcriptional activators directly converts fibroblasts to neuronal cells. Cell Stem Cell 19(3):406–414. https://doi.org/10.1016/j.stem.2016.07.001
Blakemore WF (1972) Observations on oligodendrocyte degeneration, the resolution of status spongiosus and remyelination in cuprizone intoxication in mice. J Neurocytol 1(4):413–426. https://doi.org/10.1007/BF01102943
Blau HM, Chiu CP, Webster C (1983) Cytoplasmic activation of human nuclear genes in stable heterocaryons. Cell 32(4):1171–1180
Blau HM, Pavlath GK, Hardeman EC, Chiu CP, Silberstein L, Webster SG, Miller SC, Webster C (1985) Plasticity of the differentiated state. Science (New York, NY) 230(4727):758–766
Boche D, Perry VH, Nicoll JAR (2013) Review: activation patterns of microglia and their identification in the human brain. Neuropathol Appl Neurobiol 39(1):3–18. https://doi.org/10.1111/nan.12011
Buffo A, Vosko MR, Ertürk D, Hamann GF, Jucker M, Rowitch D, Götz M (2005) Expression pattern of the transcription factor Olig2 in response to brain injuries: implications for neuronal repair. Proc Natl Acad Sci U S A 102(50):18183–18188. https://doi.org/10.1073/pnas.0506535102
Burn DJ, Jaros E (2001) Multiple system atrophy: cellular and molecular pathology. Mol Pathol 54(6):419–426
Cadwell CR, Palasantza A, Jiang X, Berens P, Deng Q, Yilmaz M, Reimer J, Shen S, Bethge M, Tolias KF et al (2016) Electrophysiological, transcriptomic and morphologic profiling of single neurons using Patch-seq. Nat Biotechnol 34(2):199–203. https://doi.org/10.1038/nbt.3445
Cahan P, Li H, Morris SA, Lummertz da Rocha E, Daley GQ, Collins JJ (2014) CellNet: network biology applied to stem cell engineering. Cell 158(4):903–915. https://doi.org/10.1016/j.cell.2014.07.020
Caiazzo M, Dell’Anno MT, Dvoretskova E, Lazarevic D, Taverna S, Leo D, Sotnikova TD, Menegon A, Roncaglia P, Colciago G et al (2011) Direct generation of functional dopaminergic neurons from mouse and human fibroblasts. Nature 476(7359):224–227. https://doi.org/10.1038/nature10284
Caiazzo M, Giannelli S, Valente P, Lignani G, Carissimo A, Sessa A, Colasante G, Bartolomeo R, Massimino L, Ferroni S et al (2015) Direct conversion of fibroblasts into functional astrocytes by defined transcription factors. Stem Cell Rep 4(1):25–36. https://doi.org/10.1016/j.stemcr.2014.12.002
Cassoli JS, Guest PC, Malchow B, Schmitt A, Falkai P, Martins-de-Souza D (2015) Disturbed macro-connectivity in schizophrenia linked to oligodendrocyte dysfunction: from structural findings to molecules. NPJ Schizophr 1:15034. https://doi.org/10.1038/npjschz.2015.34
Chakraborty S, Ji H, Kabadi AM, Gersbach CA, Christoforou N, Leong KW (2014) A CRISPR/Cas9-based system for reprogramming cell lineage specification. Stem Cell Rep 3(6):940–947. https://doi.org/10.1016/j.stemcr.2014.09.013
Chanda S, Ang CE, Davila J, Pak C, Mall M, Lee QY, Ahlenius H, Jung SW, Südhof TC, Wernig M (2014) Generation of induced neuronal cells by the single reprogramming factor ASCL1. Stem Cell Rep 3(2):282–296. https://doi.org/10.1016/j.stemcr.2014.05.020
Chen G, Wernig M, Berninger B, Nakafuku M, Parmar M, Zhang C-L (2015) In vivo reprogramming for brain and spinal cord repair. eNeuro 2(5). https://doi.org/10.1523/ENEURO.0106-15.2015, https://www.ncbi.nlm.nih.gov/pmc/articles/PMC4699832/
Chen Y, Ma N, Pei Z, Wu Z, Do-Monte FH, Huang P, Yellin E, Chen M, Yin J, Lee G, Minier A, Hu Y, Bai Y, Lee K, Quirk G, Chen G (2018) Functional repair after ischemic injury through high effciency in situ astrocyte-to-neuron conversion. bioRxiv. https://doi.org/10.1101/294967
Chernoff GF (1981) Shiverer: an autosomal recessive mutant mouse with myelin deficiency. J Hered 72(2):128
Corti S, Nizzardo M, Simone C, Falcone M, Donadoni C, Salani S, Rizzo F, Nardini M, Riboldi G, Magri F et al (2012) Direct reprogramming of human astrocytes into neural stem cells and neurons. Exp Cell Res 318(13–16):1528–1541. https://doi.org/10.1016/j.yexcr.2012.02.040
Davis RL, Weintraub H, Lassar AB (1987) Expression of a single transfected cDNA converts fibroblasts to myoblasts. Cell 51(6):987–1000
Deisseroth K (2011) Optogenetics. Nat Methods 8(1):26–29. https://doi.org/10.1038/nmeth.f.324
Dell’Anno MT, Caiazzo M, Leo D, Dvoretskova E, Medrihan L, Colasante G, Giannelli S, Theka I, Russo G, Mus L et al (2014) Remote control of induced dopaminergic neurons in parkinsonian rats. J Clin Invest 124(7):3215–3229. https://doi.org/10.1172/JCI74664
Desai MK, Sudol KL, Janelsins MC, Mastrangelo MA, Frazer ME, Bowers WJ (2009) Triple-transgenic Alzheimer’s disease mice exhibit region-specific abnormalities in brain myelination patterns prior to appearance of amyloid and tau pathology. Glia 57(1):54–65. https://doi.org/10.1002/glia.20734
Desai MK, Mastrangelo MA, Ryan DA, Sudol KL, Narrow WC, Bowers WJ (2010) Early oligodendrocyte/myelin pathology in Alzheimer’s disease mice constitutes a novel therapeutic target. Am J Pathol 177(3):1422–1435. https://doi.org/10.2353/ajpath.2010.100087
Faiz M, Nagy A (2013) Induced pluripotent stem cells and disorders of the nervous system: progress, problems, and prospects. Neuroscientist 19(6):567–577. https://doi.org/10.1177/1073858413493148
Faiz M, Sachewsky N, Gascón S, Bang KWA, Morshead CM, Nagy A (2015) Adult neural stem cells from the subventricular zone give rise to reactive astrocytes in the cortex after stroke. Cell Stem Cell 17(5):624–634. https://doi.org/10.1016/j.stem.2015.08.002
Gascón S, Murenu E, Masserdotti G, Ortega F, Russo GL, Petrik D, Deshpande A, Heinrich C, Karow M, Robertson SP et al (2016) Identification and successful negotiation of a metabolic checkpoint in direct neuronal reprogramming. Cell Stem Cell 18(3):396–409. https://doi.org/10.1016/j.stem.2015.12.003
Gascón S, Masserdotti G, Russo GL, Götz M (2017a) Direct neuronal reprogramming: achievements, hurdles, and new roads to success. Cell Stem Cell 21(1):18–34. https://doi.org/10.1016/j.stem.2017.06.011
Gascón S, Ortega F, Götz M (2017b) Transient CREB-mediated transcription is key in direct neuronal reprogramming. Neurogenesis (Austin) 4(1):e1285383. https://doi.org/10.1080/23262133.2017.1285383
Graf T, Enver T (2009) Forcing cells to change lineages. Nature 462(7273):587–594. https://doi.org/10.1038/nature08533
Grande A, Sumiyoshi K, López-Juárez A, Howard J, Sakthivel B, Aronow B, Campbell K, Nakafuku M (2013) Environmental impact on direct neuronal reprogramming in vivo in the adult brain. Nat Commun 4:2373. https://doi.org/10.1038/ncomms3373
Guo Z, Zhang L, Wu Z, Chen Y, Wang F, Chen G (2014) In vivo direct reprogramming of reactive glial cells into functional neurons after brain injury and in an Alzheimer’s disease model. Cell Stem Cell 14(2):188–202. https://doi.org/10.1016/j.stem.2013.12.001
Gurdon JB (1962) Adult frogs derived from the nuclei of single somatic cells. Dev Biol 4(2):256–273. https://doi.org/10.1016/0012-1606(62)90043-X
Han DW, Tapia N, Hermann A, Hemmer K, Höing S, Araúzo-Bravo MJ, Zaehres H, Wu G, Frank S, Moritz S et al (2012) Direct reprogramming of fibroblasts into neural stem cells by defined factors. Cell Stem Cell 10(4):465–472. https://doi.org/10.1016/j.stem.2012.02.021
Heinrich C, Blum R, Gascón S, Masserdotti G, Tripathi P, Sánchez R, Tiedt S, Schroeder T, Götz M, Berninger B (2010) Directing astroglia from the cerebral cortex into subtype specific functional neurons. PLoS Biol 8(5):e1000373. https://doi.org/10.1371/journal.pbio.1000373
Heinrich C, Bergami M, Gascón S, Lepier A, Viganò F, Dimou L, Sutor B, Berninger B, Götz M (2014) Sox2-mediated conversion of NG2 glia into induced neurons in the injured adult cerebral cortex. Stem Cell Rep 3(6):1000–1014. https://doi.org/10.1016/j.stemcr.2014.10.007
Heins N, Malatesta P, Cecconi F, Nakafuku M, Tucker KL, Hack MA, Chapouton P, Barde Y-A, Götz M (2002) Glial cells generate neurons: the role of the transcription factor Pax6. Nat Neurosci 5(4):308–315. https://doi.org/10.1038/nn828
Hu W, Qiu B, Guan W, Wang Q, Wang M, Li W, Gao L, Shen L, Huang Y, Xie G et al (2015) Direct conversion of normal and Alzheimer’s disease human fibroblasts into neuronal cells by small molecules. Cell Stem Cell 17(2):204–212. https://doi.org/10.1016/j.stem.2015.07.006
Jäkel S, Agirre E, Falcão AM, van Bruggen D, Lee KW, Knuesel I, Malhotra D, Ffrench-Constant C, Williams A, Castelo-Branco G (2019) Altered human oligodendrocyte heterogeneity in multiple sclerosis. Nature 23:1. https://doi.org/10.1038/s41586-019-0903-2
Karow M, Camp JG, Falk S, Gerber T, Pataskar A, Gac-Santel M, Kageyama J, Brazovskaja A, Garding A, Fan W et al (2018) Direct pericyte-to-neuron reprogramming via unfolding of a neural stem cell-like program. Nat Neurosci 21(7):932–940. https://doi.org/10.1038/s41593-018-0168-3
Kim J, Su SC, Wang H, Cheng AW, Cassady JP, Lodato MA, Lengner CJ, Chung C-Y, Dawlaty MM, Tsai L-H et al (2011) Functional integration of dopaminergic neurons directly converted from mouse fibroblasts. Cell Stem Cell 9(5):413–419. https://doi.org/10.1016/j.stem.2011.09.011
Kuller LH, Longstreth WT, Arnold AM, Bernick C, Bryan RN, Beauchamp NJ (2004) Cardiovascular Health Study Collaborative Research Group. White matter hyperintensity on cranial magnetic resonance imaging: a predictor of stroke. Stroke 35(8):1821–1825. https://doi.org/10.1161/01.STR.0000132193.35955.69
Lassmann H, van Horssen J, Mahad D (2012) Progressive multiple sclerosis: pathology and pathogenesis. Nat Rev Neurol 8(11):647–656. https://doi.org/10.1038/nrneurol.2012.168
Li X, Zuo X, Jing J, Ma Y, Wang J, Liu D, Zhu J, Du X, Xiong L, Du Y et al (2015) Small-molecule-driven direct reprogramming of mouse fibroblasts into functional neurons. Cell Stem Cell 17(2):195–203. https://doi.org/10.1016/j.stem.2015.06.003
Liang X-G, Tan C, Wang C-K, Tao R-R, Huang Y-J, Ma K-F, Fukunaga K, Huang M-Z, Han F (2018) Myt1l induced direct reprogramming of pericytes into cholinergic neurons. CNS Neurosci Ther 24(9):801–809. https://doi.org/10.1111/cns.12821
Liddelow SA, Barres BA (2017) Reactive astrocytes: production, function, and therapeutic potential. Immunity 46(6):957–967. https://doi.org/10.1016/j.immuni.2017.06.006
Liddelow SA, Guttenplan KA, Clarke LE, Bennett FC, Bohlen CJ, Schirmer L, Bennett ML, Münch AE, Chung W-S, Peterson TC et al (2017) Neurotoxic reactive astrocytes are induced by activated microglia. Nature 541(7638):481–487. https://doi.org/10.1038/nature21029
Liu M-L, Zang T, Zou Y, Chang JC, Gibson JR, Huber KM, Zhang C-L (2013) Small molecules enable neurogenin 2 to efficiently convert human fibroblasts into cholinergic neurons. Nat Commun 4:2183. https://doi.org/10.1038/ncomms3183
Liu Y, Yu C, Daley TP, Wang F, Cao WS, Bhate S, Lin X, Still C, Liu H, Zhao D et al (2018) CRISPR activation screens systematically identify factors that drive neuronal fate and reprogramming. Cell Stem Cell 23(5):758–771.e8. https://doi.org/10.1016/j.stem.2018.09.003
Lujan E, Chanda S, Ahlenius H, Südhof TC, Wernig M (2012) Direct conversion of mouse fibroblasts to self-renewing, tripotent neural precursor cells. Proc Natl Acad Sci 109(7):2527–2532. https://doi.org/10.1073/pnas.1121003109
Ma K, Deng X, Xia X, Fan Z, Qi X, Wang Y, Li Y, Ma Y, Chen Q, Peng H et al (2018) Direct conversion of mouse astrocytes into neural progenitor cells and specific lineages of neurons. Transl Neurodegener 7:29. https://doi.org/10.1186/s40035-018-0132-x
Marques S, Zeisel A, Codeluppi S, van Bruggen D, Mendanha Falcão A, Xiao L, Li H, Häring M, Hochgerner H, Romanov RA et al (2016) Oligodendrocyte heterogeneity in the mouse juvenile and adult central nervous system. Science (New York, NY) 352(6291):1326–1329. https://doi.org/10.1126/science.aaf6463
Marro S, Pang ZP, Yang N, Tsai M-C, Qu K, Chang HY, Südhof TC, Wernig M (2011) Direct lineage conversion of terminally differentiated hepatocytes to functional neurons. Cell Stem Cell 9(4):374–382. https://doi.org/10.1016/j.stem.2011.09.002
Masserdotti G, Gillotin S, Sutor B, Drechsel D, Irmler M, Jørgensen HF, Sass S, Theis FJ, Beckers J, Berninger B et al (2015) Transcriptional mechanisms of proneural factors and REST in regulating neuronal reprogramming of astrocytes. Cell Stem Cell 17(1):74–88. https://doi.org/10.1016/j.stem.2015.05.014
Masserdotti G, Gascón S, Götz M (2016) Direct neuronal reprogramming: learning from and for development. Development 143(14):2494–2510. https://doi.org/10.1242/dev.092163
Matsuda T, Irie T, Katsurabayashi S, Hayashi Y, Nagai T, Hamazaki N, Adefuin AMD, Miura F, Ito T, Kimura H et al (2019) Pioneer Factor NeuroD1 rearranges transcriptional and epigenetic profiles to execute microglia-neuron conversion. Neuron 101(3):472–485.e7. https://doi.org/10.1016/j.neuron.2018.12.010
Matsushima GK, Morell P (2001) The neurotoxicant, cuprizone, as a model to study demyelination and remyelination in the central nervous system. Brain Pathol (Zurich, Switzerland) 11(1):107–116
Mokhtarzadeh Khanghahi A, Satarian L, Deng W, Baharvand H, Javan M (2018) In vivo conversion of astrocytes into oligodendrocyte lineage cells with transcription factor Sox10; Promise for myelin repair in multiple sclerosis. PLoS One 13(9):e0203785. https://doi.org/10.1371/journal.pone.0203785
Morris SA (2016) Direct lineage reprogramming via pioneer factors; a detour through developmental gene regulatory networks. Development 143(15):2696–2705. https://doi.org/10.1242/dev.138263
Morris SA, Cahan P, Li H, Zhao AM, San Roman AK, Shivdasani RA, Collins JJ, Daley GQ (2014) Dissecting engineered cell types and enhancing cell fate conversion via CellNet. Cell 158(4):889–902. https://doi.org/10.1016/j.cell.2014.07.021
Najm FJ, Lager AM, Zaremba A, Wyatt K, Caprariello AV, Factor DC, Karl RT, Maeda T, Miller RH, Tesar PJ (2013) Transcription factor-mediated reprogramming of fibroblasts to expandable, myelinogenic oligodendrocyte progenitor cells. Nat Biotechnol 31(5):426–433. https://doi.org/10.1038/nbt.2561
Ninkovic J, Götz M (2018) Understanding direct neuronal reprogramming-from pioneer factors to 3D chromatin. Curr Opin Genet Dev 52:65–69. https://doi.org/10.1016/j.gde.2018.05.011
Niu W, Zang T, Zou Y, Fang S, Smith DK, Bachoo R, Zhang C-L (2013) In vivo reprogramming of astrocytes to neuroblasts in the adult brain. Nat Cell Biol 15(10):1164–1175. https://doi.org/10.1038/ncb2843
Niu W, Zang T, Wang L-L, Zou Y, Zhang C-L (2018) Phenotypic reprogramming of striatal neurons into dopaminergic neuron-like cells in the adult mouse brain. Stem Cell Rep 11(5):1156–1170. https://doi.org/10.1016/j.stemcr.2018.09.004
Reich DS, Lucchinetti CF, Calabresi PA (2018) Multiple sclerosis. N Engl J Med 378(2):169–180. https://doi.org/10.1056/NEJMra1401483
Ring KL, Tong LM, Balestra ME, Javier R, Andrews-Zwilling Y, Li G, Walker D, Zhang WR, Kreitzer AC, Huang Y (2012) Direct reprogramming of mouse and human fibroblasts into multipotent neural stem cells with a single factor. Cell Stem Cell 11(1):100–109. https://doi.org/10.1016/j.stem.2012.05.018
Rivetti di Val Cervo P, Romanov RA, Spigolon G, Masini D, Martín-Montañez E, Toledo EM, La Manno G, Feyder M, Pifl C, Ng Y-H et al (2017) Induction of functional dopamine neurons from human astrocytes in vitro and mouse astrocytes in a Parkinson’s disease model. Nat Biotechnol 35(5):444–452. https://doi.org/10.1038/nbt.3835
Roth AD, Ramírez G, Alarcón R, Von Bernhardi R (2005) Oligodendrocytes damage in Alzheimer’s disease: beta amyloid toxicity and inflammation. Biol Res 38(4):381–387
Rouaux C, Arlotta P (2013) Direct lineage reprogramming of post-mitotic callosal neurons into corticofugal neurons in vivo. Nat Cell Biol 15(2):214–221. https://doi.org/10.1038/ncb2660
Sawcer S, Franklin RJM, Ban M (2014) Multiple sclerosis genetics. Lancet Neurol 13(7):700–709. https://doi.org/10.1016/S1474-4422(14)70041-9
Sheng C, Zheng Q, Wu J, Xu Z, Sang L, Wang L, Guo C, Zhu W, Tong M, Liu L et al (2012) Generation of dopaminergic neurons directly from mouse fibroblasts and fibroblast-derived neural progenitors. Cell Res 22(4):769–772. https://doi.org/10.1038/cr.2012.32
Sieweke MH (2015) Waddington’s valleys and Captain Cook’s islands. Cell Stem Cell 16(1):7–8. https://doi.org/10.1016/j.stem.2014.12.009
Son EY, Ichida JK, Wainger BJ, Toma JS, Rafuse VF, Woolf CJ, Eggan K (2011) Conversion of mouse and human fibroblasts into functional spinal motor neurons. Cell Stem Cell 9(3):205–218. https://doi.org/10.1016/j.stem.2011.07.014
Steinbeck JA, Studer L (2015) Moving stem cells to the clinic: potential and limitations for brain repair. Neuron 86(1):187–206. https://doi.org/10.1016/j.neuron.2015.03.002
Su Z, Niu W, Liu M-L, Zou Y, Zhang C-L (2014) In vivo conversion of astrocytes to neurons in the injured adult spinal cord. Nat Commun 5:3338. https://doi.org/10.1038/ncomms4338
Takahashi K, Yamanaka S (2006) Induction of pluripotent stem cells from mouse embryonic and adult fibroblast cultures by defined factors. Cell 126(4):663–676. https://doi.org/10.1016/j.cell.2006.07.024
Tang Y, Le W (2016) Differential roles of M1 and M2 microglia in neurodegenerative diseases. Mol Neurobiol 53(2):1181–1194. https://doi.org/10.1007/s12035-014-9070-5
Thored P, Wood J, Arvidsson A, Cammenga J, Kokaia Z, Lindvall O (2007) Long-term neuroblast migration along blood vessels in an area with transient angiogenesis and increased vascularization after stroke. Stroke 38(11):3032–3039. https://doi.org/10.1161/STROKEAHA.107.488445
Tian E, Sun G, Sun G, Chao J, Ye P, Warden C, Riggs AD, Shi Y (2016) Small-molecule-based lineage reprogramming creates functional astrocytes. Cell Rep 16(3):781–792. https://doi.org/10.1016/j.celrep.2016.06.042
Toft-Hansen H, Füchtbauer L, Owens T (2011) Inhibition of reactive astrocytosis in established experimental autoimmune encephalomyelitis favors infiltration by myeloid cells over T cells and enhances severity of disease. Glia 59(1):166–176. https://doi.org/10.1002/glia.21088
Torper O, Pfisterer U, Wolf DA, Pereira M, Lau S, Jakobsson J, Björklund A, Grealish S, Parmar M (2013) Generation of induced neurons via direct conversion in vivo. Proc Natl Acad Sci 110(17):7038–7043. https://doi.org/10.1073/pnas.1303829110
Torper O, Ottosson DR, Pereira M, Lau S, Cardoso T, Grealish S, Parmar M (2015) In vivo reprogramming of striatal NG2 glia into functional neurons that integrate into local host circuitry. Cell Rep 12(3):474–481. https://doi.org/10.1016/j.celrep.2015.06.040
Vadodaria KC, Mertens J, Paquola A, Bardy C, Li X, Jappelli R, Fung L, Marchetto MC, Hamm M, Gorris M et al (2016) Generation of functional human serotonergic neurons from fibroblasts. Mol Psychiatry 21(1):49–61. https://doi.org/10.1038/mp.2015.161
Victor MB, Richner M, Hermanstyne TO, Ransdell JL, Sobieski C, Deng P-Y, Klyachko VA, Nerbonne JM, Yoo AS (2014) Generation of human striatal neurons by microRNA-dependent direct conversion of fibroblasts. Neuron 84(2):311–323. https://doi.org/10.1016/j.neuron.2014.10.016
Vierbuchen T, Wernig M (2011) Direct lineage conversions: unnatural but useful? Nat Biotechnol 29(10):892–907. https://doi.org/10.1038/nbt.1946
Vierbuchen T, Ostermeier A, Pang ZP, Kokubu Y, Südhof TC, Wernig M (2010) Direct conversion of fibroblasts to functional neurons by defined factors. Nature 463(7284):1035–1041. https://doi.org/10.1038/nature08797
Waddington CH (1957) The strategy of the genes; a discussion of some aspects of theoretical biology. Routledge, London. https://doi.org/10.4324/9781315765471
Wang L-L, Zhang C-L (2018) Engineering new neurons: in vivo reprogramming in mammalian brain and spinal cord. Cell Tissue Res 371(1):201–212. https://doi.org/10.1007/s00441-017-2729-2
Wapinski OL, Lee QY, Chen AC, Li R, Corces MR, Ang CE, Treutlein B, Xiang C, Baubet V, Suchy FP et al (2017) Rapid chromatin switch in the direct reprogramming of fibroblasts to neurons. Cell Rep 20(13):3236–3247. https://doi.org/10.1016/j.celrep.2017.09.011
Xie H, Ye M, Feng R, Graf T (2004) Stepwise reprogramming of B cells into macrophages. Cell 117(5):663–676
Xu J, Du Y, Deng H (2015) Direct lineage reprogramming: strategies, mechanisms, and applications. Cell Stem Cell 16(2):119–134. https://doi.org/10.1016/j.stem.2015.01.013
Yang N, Ng YH, Pang ZP, Südhof TC, Wernig M (2011) Induced neuronal (iN) cells: how to make and define a neuron. Cell Stem Cell 9(6):517–525. https://doi.org/10.1016/j.stem.2011.11.015
Yang N, Zuchero JB, Ahlenius H, Marro S, Ng YH, Vierbuchen T, Hawkins JS, Geissler R, Barres BA, Wernig M (2013) Generation of oligodendroglial cells by direct lineage conversion. Nat Biotechnol 31(5):434–439. https://doi.org/10.1038/nbt.2564
Yoo AS, Sun AX, Li L, Shcheglovitov A, Portmann T, Li Y, Lee-Messer C, Dolmetsch RE, Tsien RW, Crabtree GR (2011) MicroRNA-mediated conversion of human fibroblasts to neurons. Nature 476(7359):228–231. https://doi.org/10.1038/nature10323
Zamanian JL, Xu L, Foo LC, Nouri N, Zhou L, Giffard RG, Barres BA (2012) Genomic analysis of reactive astrogliosis. J Neurosci 32(18):6391–6410
Zhang L, Lei Z, Guo Z, Pei Z, Chen Y, Zhang F, Cai A, Mok YK, Lee G, Swaminathan V et al (2018) Reversing glial scar Back to neural tissue through NeuroD1-mediated astrocyte-to-neuron conversion. bioRxiv 7:261438. https://doi.org/10.1101/261438
Zhu X, Zhou W, Jin H, Li T (2018) Brn2 alone is sufficient to convert astrocytes into neural progenitors and neurons. Stem Cells Dev 27(11):736–744. https://doi.org/10.1089/scd.2017.0250
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2019 Springer Nature Switzerland AG
About this chapter
Cite this chapter
Bajohr, J., Faiz, M. (2019). Direct Lineage Reprogramming in the CNS. In: Turksen, K. (eds) Cell Biology and Translational Medicine, Volume 6. Advances in Experimental Medicine and Biology(), vol 1212. Springer, Cham. https://doi.org/10.1007/5584_2019_374
Download citation
DOI: https://doi.org/10.1007/5584_2019_374
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-030-32822-1
Online ISBN: 978-3-030-32823-8
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)