Abstract
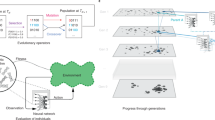
Dynamic Local Search [1] has been applied to the evolution of interactions between protein-like structures. These are composed of a randomly selected sequence of amino acids that are linked together to form linear polymers in three dimensions. The objective function chosen for optimisation is the potential energy given by a Toy protein model. Proteins fold, move and interact with other chains to minimise their objective function at a given rate, F,ote, depending on the sum of the rates for re-organisation of their structures. The interaction between different proteins gives a whole range of local attraction/repulsion regimes that result in new structures with new bonds, broken bonds and recursive loops.
Preview
Unable to display preview. Download preview PDF.
Similar content being viewed by others
References
Amin, S., Fernández-Villacañas, J.L.: Dynamic Local Search. Proc. Galesia97 conference, Glasgow, September 1997
Lau, K.F., Dill, A.: Theory for protein mutability and biogenesis. Proc. Nat. Acad. Sci. USA 87 1990 638–642
Unger, R., Moult, J.: Genetic Algorithms for Protein Folding Simulations. J. Mol. Biol. 231 1993 75–81
Jaeger, J.A., Turner, D.H., Zucker, M.: Improved predictions of secondary structures for RNA. Proc. Nat. Acad. Sci. USA 86 1989 7706–10
Quian, N., Sejnowski, T.J.: Predicting the Secondary structure of Globular proteins using Neural Network models. J. Mol. Biol. 202 1988 865–884
Fariselli, P., Compiani, M., Casadio, R.: Predicting Secondary structures of Membrane proteins with Neural Networks. Euro. Biophys. J. 22 1993 41–51
Calabretta, R., Nolfi, S., Parisi, D.: An Artificial Life model for predicting the Tertiary Structure of unknown proteins that emulates the folding process. Proc. of Third European Conference of Artificial Life, Granada 1995
Stillinger, F.H., Head-Gordon, T., Hirsfield, C.L.: Toy model for protein folding. Physical Review E 48 2 1993 1469
Fernández-Villacañas, J.L., Fatah, J., Amin, S.: Evolution of Protein Interactions. Proc. BCEC'97 Bio-Computing and Emergent Behaviour, Skövde, Sweden, September 1997
Berstein, F.C., Koetzle, T.F., Williams, G.J.B., Meyer, E.F., Brice, M.D., Rodgers, J.R., Kennard, O., Shimanouchi, T., Tasumi, M.: The protein data bank: a computer-based archival file for macromolecular structures. J. Mol. Biol. 112 1977 535–542 *** DIRECT SUPPORT *** A0008D07 00008
Author information
Authors and Affiliations
Editor information
Rights and permissions
Copyright information
© 1998 Springer-Verlag Berlin Heidelberg
About this paper
Cite this paper
Fernández-Villacañas, J.L., Fatah, J.M., Amin, S. (1998). Computing with evolving proteins. In: Rolim, J. (eds) Parallel and Distributed Processing. IPPS 1998. Lecture Notes in Computer Science, vol 1388. Springer, Berlin, Heidelberg. https://doi.org/10.1007/3-540-64359-1_690
Download citation
DOI: https://doi.org/10.1007/3-540-64359-1_690
Published:
Publisher Name: Springer, Berlin, Heidelberg
Print ISBN: 978-3-540-64359-3
Online ISBN: 978-3-540-69756-5
eBook Packages: Springer Book Archive