Abstract
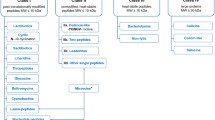
A survey is given of the main classes of bacteriocins, produced by lactic acid bacteria: I. lantibiotics II. small heat-stable non-lanthionine containing membrane-active peptides and III. large heat-labile proteins. First, their mode of action is detailed, with emphasis on pore formation in the cytoplasmatic membrane. Subsequently, the molecular genetics of several classes of bacteriocins are described in detail, with special attention to nisin as the most prominent example of the lantibiotic-class. Of the small non-lanthionine bacteriocin class, the Lactococcus lactococcins, and the Lactobacillus sakacin A and plantaricin A-bacteriocins are discussed. The principles and mechanisms of immunity and resistance towards bacteriocins are also briefly reported. The biosynthesis of bacteriocins is treated in depth with emphasis on response regulation, post-translational modification, secretion and proteolytic activation of bacteriocin precursors. To conclude, the role of the leader peptides is outlined and a conceptual model for bacteriocin maturation is proposed.
Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only
Preview
Unable to display preview. Download preview PDF.
Similar content being viewed by others
References
De Vuyst L, Vandamme EJ (eds) (1994) Bacteriocins of Lactic Acid Bacteria: Microbiology, Genetics and Applications. Blackie Academic & Professional, London
Rose AH (1982) Fermented Foods. Academic Press, New York
Reed G (1983) Food and Feed Production with Microorganisms. Verlag Chemie, Deerfield Beach, Florida
Steinkraus KH (1983) Handbook of Indigenous Fermented Foods. Marcel Dekker, New York
Wood BJB (1985) Microbiology of Fermented Foods. Elsevier, London
Gilliland SE (1986a) Bacterial Starter Cultures for Foods. CRC Press, Boca Raton, Florida
Buckenhüskes HJ (1993) Selection criteria for lactic acid bacteria to be used as starter cultures for various food commodities. FEMS Microbiol Rev 12: 253–271
Gilliland SE (1986b) Role of starter culture bacteria in food preservation. In: Gilliland SE (ed) Bacterial Starter Cultures for Foods. CRC Press, Boca Raton, Florida, pp 175–185
Lindgren SE, Dobrogosz WJ (1990) Antagonistic activities of lactic acid bacteria in food and feed fermentations. FEMS Microbiol Rev 12: 207–220
Schillinger U (1990) Bacteriocins of lactic acid bacteria. In: Bills DD, Kung SD (eds) Biotechnology and Food Safety. Burrerworth-Heinemann, Boston, pp 55–74
Vandenbergh PA (1993) Lactic acid bacteria, their metabolic products and interference with microbial growth. FEMS Microbiol Rev 12: 221–237
Lloyd AG, Drake JJP (1975) Problems posed by essential food preservatives. Br Med bull. 31: 214–219
Lewus CB, Kaiser A, Montville TJ (1991) Inhibition of food-borne bacterial pathogens by bacteriocins from lactic acid bacteria isolated from meat. Appl Environ Microbiol 57:1683–1688
Marteau P, Rambeaud J-C (1993) Potential of using lactic acid bacteria for therapy and immunomodulation in man. FEMS Microbiol Rev 12: 207–220
Gerritse K, Posno M, Schellekens M, Boersma WJA, Claassen E (1990) Oral administration of TNP-Lactobacillus conjugates in mice: a model for evaluation of mucosal and systemic immune responses and memory formation elicited by transformed lactobacilli. Res Microbiol 141: 955–962
Norton PM, Wells JM, Brown HWG, Macpherson AM, Le Page RWF (1997) Protection against tetanus toxin in mice nasally immunized with recombant Lactobacillus lactis expressing tetanus toxin fragment C. Vaccine 15: 616–649
Tagg JR, Dajani AS, Wannamaker LW (1976) Bacteriocins of Gram-positive bacteria. Bacteriol Rev 40: 722–756
Gratia A (1925) Sur un remarquable example d’antagonisme entre souches de colibacille. CR Soc Biol 93:1040–1041
Frédericq P (1948) Actions antibiotiques reciproques chez les Enterobacteriaceae. Rev Bel Pathol Med Exp 19:1–107
Jack RW, Tagg JR, Ray B (1995) Bacteriocins of Gram-positive bacteria. Microbiol Rev 59:171–200
Klaenhammer TR (1993) Genetics of bacteriocins produced by lactic acid bacteria. FEMS Microbiol Rev 12: 39–86
Jung G, Sahl H-G (1991) Nisin and Novel Lantibiotics, ESCOM Science Publishers BV., Leiden
Baba T, Schneewind O (1998) Instruments of microbial warfare: bacteriocin synthesis, toxicity and immunity. Trends Microbiol 6: 66–71
Rogers LA (1928) The inhibitory effect of Streptococcus lactis on Lactobacillus bulgaricus. J Bacteriol 16: 321–325
Whitehead HR (1933) A substance inhibiting bacterial growth, produced by certain strains of lactic streptococci. Biochem J 27:1793–1800
Mattick ATR, Hirsch A (1947) Further observations on an inhibitory substance (nisin) from lactic streptococci. Lancet 2: 5–7
Hurst A (1981) Nisin.Adv Appl Microbiol 27: 85–123
Nes IF, Diep DB, Håvarstein LS, Brurberg MB, Eijsink V, Holo H (1996) Biosynthesis of bacteriocins in lactic acid bacteria. Antonie van Leeuwenhoek
Kaletta C, Entian K-D (1989) Nisin, a peptide antibiotic: cloning and sequencing of the nisA gene and posttranslational processing of its peptide product. J Bacteriol 171:1597–1601
Piard JC, Delorme F, Giraffa G, Commissaire J, Desmazeaud M (1990) Evidence for a bacteriocin produced by lactococcus lactis CNRZ 481. Neth Milk Dairy J 44: 143–158
Møfrtveldt CI, Nes IF (1990) Plasmid-associated bacteriocin production by a Lactobacillus sake strain. J Gen Microbiol 136:1601–1607
Horn N, Swindell S, Dodd H, Gasson M (1991) Nisin biosynthesis genes are encoded by a novel conjugative transposon. Mol Gen Genet 228:129–135
Møfrtveldt CI, Nissen-Meyer J, Sletten K, Nes IF (1991) Purification and amino acid sequence of lactocin S, a bacteriocin produced by Lactobacillus sake L45. Appl Environ Microbiol 57:1829–1834
Harris LJ, Fleming HP, Klaenhammer TR (1992) Developments in nisin research. Food Res Int 25: 57–66
Stoffels G, Nes IF, Gudmundsdottir A (1992) Isolation and properties of a bacteriocinproducing Carnobacterium piscicola isolated from fish. J Appl Bacteriol 73: 309–316
Stoffels G, Nissen-Meyer J, Gudmundsdottir A, Sletten K, Holo H, Nes IF (1992) Purification and characterization of a new bacteriocin isolated from a Carnobacterium sp. Appl Environ Microbiol 58:1417–1422
Rauch PJG, Beerthuyzen MM, De Vos WM (1990) Nucleotide sequence of IS904 from Lactococcus lactis subsp. lactis strain NIZO R5. Nucleic Acids Res 18: 4253–4254
Paik SH, Chakicherla A, Hansen JN (1998) Identification and Characterization of the Structural and Transporter Genes for, and the Chemical and Biological Properties of Sublancin 168, a Novel lantibiotic Produced by Bacillus subtilis 168. J Biol Chem 273: 23134–23142
Jung G (1991) Lantibiotics: a survey. In: Jung G, Sahl H-G (eds) Nisin and Novel Lantibiotics. ESCOM Science, Leiden, pp 1–34
De Vos WM,K uipers OP, van der Meer JR, Siezen RJ (1995b) Maturation pathway of nisin and other lantibiotics: post-translational modified antimicrobial peptides exported by Gram-positive bacteria. Mol Microbiol 17: 427–437
Havarstein LS, Holo H, Nes IF (1994) The leader peptide of colicin V shares consensus sequences with leader peptides that are common among peptide bacteriocins produced by gram positive bacteria. Microbiology 140: 2383–2389
Hastings JW, Sailer M, Johnson K, Rou KK, Vederas JC, Stiles ME (1991) Characterization of leucocin A UAL 187 and cloning of the bacteriocin gene from Leuconostoc gelidum. J Bacteriol 173: 7491–7500
Holck A, Axelsson L, Birkeland S-E, Aukrust T, Blom H (1992) Purification and amino acid sequence of sakacin A, a bacteriocin from Lactobacillus sake Lb706. J Gen Microbiol 138: 2715–2720
Lozano JCN, Meyer JN, Sletten K, Pelaz C, Nes IF (1992) Purification and amino acid sequence of a bacteriocin produced by Pedicococcus acidilactici. J Gen Microbiol138: 1985–1990
Tichaczek PS, Nissen-Meyer J, Nes IF, Vogel RF, Hammes WP (1992) Characterization of the bacteriocin curvacin A from Lactobacillus curvatus LTH 1174 and sakacin P from L.sa ke LTH673. Syst Appl Microbiol 15: 460–468
van Belkum MJ, Hayema BJ, Jeeninga RE, Kok J, Venema G (1991a) Organization and nucleotide sequence of two lactococcal bacteriocin operons. Appl Environ Microbiol 57: 492–498
Nissen-Meyer J, Holo H, Håvarstein LS, Sletten K, Nes IF (1992) A novel lactococcal bacteriocin whose activity depends on the complementary action of two peptides. J Bacteriol 174: 5686–5692
Allison GE, Frémaux C, Klaenhammer TR (1994) Expansion of bacteriocin activity and host range upon complementation of two peptides encoded within the lactacin F operon. J Bacteriol 176: 2235–2241
Diep DB, Håvarstein LS, Nes IF (1996) Characterization of the locus responsible for the bacteriocin production in Lactobacillus plantarum Cll. J Bacteriol 178: 4472–4483
Leer RL, van der Vossen JMBM, van Giezen M, van Noort JM, Pouwels PH (1995) Genetic analysis of acidocin B, a novel bacteriocin produced by lactobacillus acidophilus. Microbiology 141:1629–1635
Worobo RW, van Belkum MJ, Sailer M, Roy KL, Vederas JC, Stiles ME (1995) A signal peptide-dependent bacteriocin from Carnobacterium divergens. J Bacteriol 177: 3143–3149
Metha AM, Patel KA, Dave PJ (1983) Isolation and purification of an inhibitory protein from Lactobacillus acidophilus ACT. Microbiology 37: 37–43
Joerger MC, Klaenhammer TR (1986) Characterization and purification of helveticin J and evidence for a chromosomally determined, bacteriocin produced by Lactobacillus helveticus 481. J Bacteriol 167: 439–446
Joerger MC, Klaenhammer TR (1990) Cloning, expression and nucleotide sequence of the Lactobacillus helveticus 481 gene encoding the bacteriocin helveticin J JBacteriol 172: 6339–6347
Toba T, Yoshioka E, Itoh T (1991) Acidophilucin A, a new heat-labile bacteriocin produced by Lactobacillus acidophilus LAPT 1060. Lett Appl Microbiol 12:106–108
Vaughan EE, Daly C, Fitzgerald GF (1992) Identification and characterization of helveticin V-1829, a bacteriocin produced by Lactobacillus helveticus 1829. J Appl Bacteriol 73: 299–308
Upreti GC, Hinsdill RD (1973) Isolation and characterization of a bacteriocin from a homofermentative Lactobacillus. Antimicrob Agents Chemother 4: 487–494
Upreti GC, Hinsdill RD (1975a) Isolation and characterization of a bacteriocin from a homofermentative Lactobacillus. Antimicrob Agents Chemother 7:139–145
Lewus CB, Sun S, Montville TJ (1992) Production of an amylase-sensitive bacteriocin by an atypical Leuconostoc paramesenteroides strain. Appl Environ Microbiol 58: 143–149
Jiménez-Dïaz R, Rios-Sánchez RM, Desmazeaud M, Ruiz-Barba JL, Piard J-C (1993) Plantaricin S and T, two new bacteriocins produced by Lactobacillus plantarum LPCO10 isolated from a green olive fermentation. Appl Environ Microbiol 59:1416–1424
Schved F, Lalazar A, Henis Y, Juven BJ (1993) Purification, partial characterization and plasmid linkage of pediocin SJ-1, a bacteriocin produced by Pediococcus acidilacitici. J Appl Bacteriol 74: 67–77
Venema K, Venema G, Kok J (1995) Lactococcal bacteriocins: mode of action and immunity. Trends Microbiol 3: 299–304
Davey GP (1981) Mode of action of diplococcin, a bacteriocin from Streptococcus cremoris 346. NZJ Dairy Sci Technol 16:187–190
Zajdel JK, Ceglowski P, Dobrzanski WT (1985) Mechanism of action of lactostrepcin 5, a bacteriocin produced by Streptococcus cremoris. Appl Environ Microbiol 49: 969–974
van Belkum MJ, Kok J, Venema G, Holo H, Nes IF, Konings WN and Abee T (1991b) The bacteriocin lactococcin A specifically increases permeability of lactococcal cytoplasmic membranes in a voltage-independent, protein-mediated manner. J Bacteriol 173: 7934–7941
Bhunia AK, Johnson MC, Ray B, Kalchayanand N (1991) Mode of action of pediocin AcH from Pediococcus acidilactici H on sensitive bacterial strains. J Appl Bacteriol 70: 25–33
Sahl H-G (1991) Pore formation in bacterial membranes by cationic lantibiotics. In: Jung G, Sahl H-G (eds) Nisin and Novel Lantibiotics. ESCOM Science, Leiden, pp 347–358
Upreti GC, Hinsdill RD (1975b) Production and mode of action of lactocin 27 bacteriocin from a homofermentative Lactobacillus. Antimicrob Agents Chemother 7:139–145
Hastings JW, Stiles ME (1991) Antibiosis of Leuconostoc gelidum isolated from meat. J Appl Bacteriol 70:127–134
Kok J, Holo H, van Belkum MJ, Haandrikman AJ, Nes IF (1993) Non-nisin bacteriocins in lactococci: biochemistry, genetics and mode of action. In: Hoover D, Steenson L (eds) Bacteriocins of Lactic Acid Bacteria. Academic Press, New York, pp 121–150
Venema K, Abee T, Haandrikman AJ, Leenhouts KJ, Kok J, Konings WN, Venema G (1993) Mode of action of lactococcin B, a thiol-activated bacteriocin from Lactococcus lactis. Appl Environ Microbiol 59:1041–1048
Moll G, Ubbink-Kok T, Hildeng-Hauge H, Nissen-Meyer J, Nes IF, Konings WN, Driessen AJM (1996) Lactococcin G is a potassium ion-conducting, two-component bacteriocin. J Bacteriol 178: 600–605
Christensen DP, Hutkins RW (1992) Collapse of the proton motive force in Listeria monocytogenes caused by a bacteriocin produced by Pediococcus acidilactici. Appl Environ Microbiol 58: 3312–3315
Ray B, Hoover DG (1993) Pediocins. In: Hoover DG, Steenson LR (eds) Bacteriocins of Lactic Acid Bacteria. Academic Press, New York, pp 181–210
Chikindas ML, Garcia-Garcera MJ, Driessen AJM, Ledeboer AM, Nissen-Meyer N, Nes IF, Abee T, Konings WN, Venema G (1993) Pediocin PA-1, a bacteriocin from Pediococcus acidilactici PAC1.0, forms hydrophylic pores in the cytoplasmic membrane of target cells. Appl Environ Microbiol 59: 3577–3584
Kaiser ET (1987) Design of amphiphilic peptides. In: Alan R (ed) Protein Engineering. Liss, New York, pp 193–199
Frémaux C, Ahn C, Klaenhammer TR (1993) Molecular analysis of the lactacin F operon. Appl Environ Microbiol 59: 3906–3915
Salomon RA, Farias RN (1993) The FhuA protein is involved in microcin 25 uptake. J Bacteriol 175: 7741–7742
Salomon RA, Farias RN (1995) The peptide antibiotic microcin 25 is imported through the TonB pathway and the SbmA protein. J Bacteriol 177: 3323–3325
Moeck GS, Fasly Bazzaz BS, Gras MF, Ravi TS, Ratcliffe MJH, Coulton JW (1994) Genetic insertion and exposure of a reporter epitope in the ferrichrome-iron receptor of Escherichia coli K-12. J Bacteriol 176: 4250–4259
Gross E, J, Morell J (1971) The stucture of nisin. J Am Chem Soc 93: 4634–4635
Sahl H-G, Jack RW, Bierbaum G (1995) Biosynthesis and biological activities of lantibiotics with unique post-translational modifications. Eur J Biochem 230: 827–833
Mulders JW, Boerrigter LJ, Rollema HS, Siezen RJ, De Vos WM (1991) Identification and characterization of the lantibiotic nisin Z, a natural nisin variant. Eur J Biochem 201: 581–584
Banerjee S, Hansen JN (1988) Structure and expression of a gene encoding the precursor of subtilin, a small protein antibiotic. J Biol Chem 263: 9508–9514
Reis M, Sahl H-G (1991) Genetic analysis of the producer self-protection mechanism (‘immunity’) against Pep5. In: Jung G, Sahl H-G (eds) Nisin and Novel Lantibiotics. ESCOM, Leiden, pp 320–331
Chung YJ, Steen MT, Hansen JN (1992) The subtilin gene of Bacillus subtilis ATCC 6633 is encoded in an operon that contains a homologue of the hemolysin B transport protein. J Bacteriol 174:1417–1422
Klein C, Kaletta C, Schnell N, Entian K-D (1992) Analysis of genes involved in the biosynthesis of the lantibiotic subtilin. Appl Environ Microbiol 58:132–142
Schnell N, Engelke G,A ugustin J, Rosenstein R,U ngermann V, Götz F, Entian K-D (1992) Analysis of genes involved in the biosynthesis of the lantibiotic epidermin. Eur J Biochem 204: 57–68
Klein C, Kaletta C, Entian K-D (1993) Biosynthesis of the lantibiotic subtilin is regulated by a histidine kinase/response regulator system. Appl Environ Microbiol 59: 296–303
Schnell N, Entian K-D, Schneider U, Götz F, Zähner H, Kellner R, Jung G (1988) Prepeptide sequence of epidermin, a ribosomally synthesized antibiotic with four sulphide-rings. Nature 333: 276–278
Steen MT, Chung YJ, Hansen JN (1991) Characterization of the nisin gene as a part of a polycistronic operon in the chromosome of Lactococcus lactis ATCC 11454. Appl Environ Microbiol 57:1181–1188
Kuipers OP, Beerthuyzen MM, Siezen JR, De Vos W (1993) Characterization of the nisin gene cluster nisABTCIPR of Lactococcus lactis. Requirement of expression of the nisA and nisI genes for development of nisin immunity. Eur J Biochem 216: 281–291
Siezen RJ, Kuipers OP, de Vos WM (1996) Comparison of lantibiotics geneclu sters and encoded proteins. Antonie van Leeuwenhoek 69:171–184
Gasson MJ (1984) Transfer of sucrose fermenting ability, nisin resistance and nisin production in Streptococcus lactis 712. FEMS Microbiol Lett 21: 7–10
Dodd HM, Horn N, Gasson MJ (1990) Analysis of the genetic determinant for production of the petpide antibiotic nisin. J Gen Microbiol 136: 555–566
Rodriguez JM, Dodd HM (1996) Genetic determinants for the biosynthesis of nisin, a bacteriocin produced by Lactobacillus lactis. Microbiologia SEM 12: 61–74
Rauch PJG, De Vos WM (1992) Characterization of the novel nisin-sucrose conjugative transposon Tn5276 and its insertion in Lactococcus lactis. J Bacteriol 174:1280–1287
Rauch PJG, De Vos WM (1994) Identification and characterization of the genes involved in excision of the Lactococcus lactis conjugative transposon Tn5276. J Bacteriol 176: 2165–2171
Poyart-Salmeron C, Trieu-Cuot P, Carlier C, Courvalin P (1989) Molecular characterization of two proteins involved in the excision of the conjugative transposon Tn1545: homologies with other site-specific recombinases. EMBO J 8: 2425–2433
Engelke G, Gutowski-Eckel Z, Hammelmann M, Entian K-D (1992) Biosynthesis of the lantibiotic nisin: genomic organization and membrane localization of the NisB protein. Appl Environ Microbiol 58: 3730–3743
van der Meer FR, Polman J, Beerthuyzen MM, Siezen RJ, Kuipers OP, De Vos W (1993) Characterization of the Lactococcus lactis nisin A operon genes nisP, encoding a subtilisin-like serine protease involved in precursor processing and nisR, encoding a regulatory protein involved in nisin biosynthesis. J Bacteriol 174: 2152–2159
van der Meer FR, Rollema HS, Siezen RJ, Beerthuyzen MM, Kuipers OP, De Vos WM (1994) Influence of amino acid substitutions in the nisin leader peptide on biosynthesis and secretion of nisin by Lactococcus lactis. J Biol Chem 269: 3555–3562
De Vos WM, Beerthuyzen MM, Luesink EL, Kuipers OP (1995a) Genetics of the nisin operon and the sucrose-nisin conjugative transposon Tn5276. In: Ferretti JJ, Gilmore MS, Klaenhammer TR (eds) Genetics of Streptococci, Enterococci and Lactococci. Karger, New York, pp 617–625
Siegers K, Entian K-D (1995) Genes involved in immunity to the lantibiotic nisin produced by Lactococcus lactis 6F3. Appl Environ Microbiol 61:1081–1089
Fath MJ, Kolter R (1993) ABC exporters: bacterial exporters. Microbiol Rev 57: 995–1017
Qiao M, Saris PE (1996) Evidence for a role of Nis T in transport of the lantibiotic nisin produced by Lactococcus lactis N8. FEMS Microbil Lett 144: 89–93
Engelke G, Gutowski-Eckel Z, Kiesau P, Siegers K, Hammelmann M, Entian K-D (1994) Regulation of nisin biosynthesis and immunity in Lactococcus lactis 6F3. Appl Environ Microbiol 60: 814–825
Stock JB, Ninfa AJ, Stock AM (1989) Protein phosphorylation and regulation of adaptive response in bacteria. Microbiol Rev 53: 450–490
Garido MC, Herrero M, Kolter R, Moreno F (1988) The export of the DNA replication inhibitor microcin B17 provides immunity for the host cell. EMBO J 7:1853–1862
Klein C, Entian K-D (1994) Genes involved in self-protection against the antibiotic subtilin produced by Bacillus subtilis ATCC 6633. Appl Environ Microbiol 60: 2793–2801
Pugsley AP (1988) Protein secretion across the outer membrane of Gram-negative bacteria. In: Das RA, Robins PW (eds) Protein transfer and organelle biogenesis. Academic Press, Orlando, pp 607–652
Song HY, Cramer WA (1991) Membrane topography of ColEI gene products: the immunity protein. J Bacteriol 173: 2935–2943
Immonen T, Ye S, Ra R, Quia M, Paulin L, Saris PEJ (1991) The codon usage of the nisZ operon in Lactococcus lactis N8 suggests a non-lactococcal origin of the conjugative nisin-sucrose transposon. DNA sequence 5: 203–218
De Vos WM, Simmons GFM (1994) Gene cloning and expression systems in lactococci. In: Gasson MJ, De Vos WM (eds) Genetics and Biotechnology of Lactic Acid Bacteria. Blackie Academic & Professional, Glasgow, pp 52–97
Ra SR, Qiao M, Immonen T, Pujana I, Saris PEJ (1996) Genes responsible for nisin synthesis, regulation and immunity form a regulon of two operons and are induced by nisin in Lactococcus lactis N8. Microbiology 142:1281–1288
Ra SR, Saris PEJ (1995) Characterization of procaryotic mRNAs by RT-PCR. Biotechniques 18: 792–795
Piard JC, Muriana PM, Desmazeaud PJ, Klaenhammer TR (1992) Purification and partial characterization of lacticin 481, a lanthionine-containing bacteriocin produced by Lactococcus lactis subsp. lactis CNRZ 481. Appl Environ Microbiol 58: 279–284
Rince A, Dufour A, Le Pogam S, Thuault D, Bourgeois CM, Le Pennec JP (1994) Cloning, expression and nucleotide sequence of genes involved in production of lactococcin DR, a bacteriocin from lactococcus lactis subsp. lactis. Appl Environ Microbiol 60:1652–1657
Rince A, Dufour A, Uguen P, Le Pennec JP, Haras D (1997) Characterization of the lacticin 481 operon: the Lactococcus lactis genes IctF, IctE and IctG encode a pupative ABC transporter involved in bacteriocin immunity. Appl Environ Microbiol 63: 4252–4260
Nes IF, Tagg JR (1996) Novel lantibiotics and their prepeptides. Antonie van Leeuwenhoek 69: 89–97
Skaugen M, Abildgaard CI, Nes IF (1997) Organization and expression of a gene cluster involved in the biosynthesis of the lantibiotic lactocin S.Mol Gen Genet 253: 674–686
Gilmore MS, Segarra RA, Booth MC, Bogie CP, Hall LR, Clewell DB (1994) Genetic structure of the Enterococcus faecalis plasmid pAD1-encoded cytolytic toxin system and its relationship to lantibiotic determinants. J Bacteriol 176: 7335–7344
Siezen RJ, De Vos WM, Leunissen JAM, Dijkstra BW (1995) Homology modelling and protein engineering strategy of subtilases, the family of subtilisin-like serine proteinases. Protein Eng 4: 719–737
Frémaux C, Héchard Y, Cienatiempo Y (1995) Mesentericin Y105 gene cluster in Leuconostoc mesenteroides Y105. Microbiology 141:1637–1645
Holo H, Nilssen Ø, Nes IF (1991) Lactococcin A, a new bacteriocin from Lactococcus lactis subsp. cremoris. Isolation and characterization of the protein and its gene. J Bacteriol 173: 3879–3887
Stoddard GW, Petzel JP, van Belkum MJ, Kok J, McKay LL (1992) Molecular analysis of the lactococcin A gene cluster from Lactococcus lactis subsp. lactis Biovar diacetylactis WM4. Appl Environ Microbiol 58:1952–1961
van Belkum MJ (1994) Lactococcins, bacteriocins of Lactococcus lactis. In: De Vuyst L, Vandamme EJ (eds) Bacteriocins of Lactic Acid Bacteria: Microbiology, Genetics and Applications. Blackie Academic & Professional, London, pp 301–318
Nes IF, Håvarstein LS, Holo H (1995) Genetics of non-lantibiotic bacteriocins. In: Ferretti JJ, Gilmore MS, Klaenhammer TR (eds) Genetics of Streptococci, Enterococci and Lactococci. Karger, New York, pp 645–651
Van Belkum MJ, Stiles ME (1995) Molecular characterization of the genes involved in the production of the bacteriocin leucocin A from Leuconostoc gelidum. Appl Environ Microbiol 61: 3573–3579
Marugg JD, Gonzalez CF, Kunka BS, Ledeboer AM, Pucci MF, Toonen MY, Walker SA, Zoetmulder LCM, Vandenberg PA (1992) Cloning, expression and nucleotide sequence of genes involved in production of pediocin PA-1, a bacteriocin from Pediococcus acidilactici PACl.0. Appl Environ Microbiol 58: 2360–2367
Bukhtiyarova M, Yang R, Ray B (1994) Analysis of the pediocin AcH gene cluster from plasmid pSMB74 and its expression in a pediocin-negative strain. Appl Environ Microbiol 60: 3405–3408
Venema K, Kok J,M arugg JD, Toonen MY, Ledeboer AM, Venema G, Chikindas L (1995) Functional analysis of the pediocin operon of Pediococcus acidilactici PAC 1.0 PedB is the isnmunity protein and PedD is the precursor processing enzyme. Mol Microbiol 17: 515–522
Axelsson L, Holck A (1995) The genes involved in production of and immunity to sakacin A, a bacteriocin from Lactobacillus sake Lb706. J Bacteriol 177: 2125–2137
Diep DB, Håvarstein LS, Nissen-Meyer J, Nes IF (1994) The gene encoding plantaricin A, a bacteriocin from Lactobacillus plantarum C ll, is located on the same transcription unit as an agr-like regulatory system. Appl Environ Microbiol 60:160–166
van Belkum MJ, Hayema BJ, Jeeninga RE, Kok J, Venema G (1989) Cloning of two bacteriocin genes from a lactococcal bacteriocin plasmid. Appl Environ Microbiol 55:1187–1191
van Belkum MJ, Kok J, Venema G (1992) Cloning, sequencing and expression in Escherichia coli of IcnB, a third bacteriocin determinant from the lactococcal bacteriocin plasmid p9B4-6. Appl Environ Microbiol 58:572–577
Delepelaire P, Wandersman C (1990) Protein secretion in Gram-negative bacteria. J Biol Chem 265:17118–17125
Harmon KS, McKay LL (1987) Restriction enzyme analysis of lactose and bacteriocin plasmids from Streptococcus lactis subsp. diacetylactis WM4 and cloning of BclII fragments coding for bacteriocin production. Appl Environ Microbiol 53:1171–1174
Randall LL, Hardy SJS, Thom JR (1987) Export of protein: a biochemical view. Annu Rev Microbiol 41: 507–541
Wagner W, Vogel M, Goebel W (1983) Transport of hemolysin across the outer membrane of Escherichia coli requires two functions. J Bacteriol 27:1793–1800
Gilson L, Mahanty HK, Kolter R (1990) Genetic analysis of an MDR-like export system: the secretion of colicin V. EMBO J 9: 3875–3884
Delepelaire P, Wandersman C (1990) Protein secretion in Gramnegative bacteria. J Biol Chem 265:17118–17125
Strathdee CA, Lo RY (1989) Cloning, nucleotide sequence and characterization of genes encoding the secretion function of the Pasteurella haemolytica leukotoxin determinant. J Bacteriol 171: 916–928
Letoffe S, Delepelaire P, Wandersman C (1990) Protease secretion by Erwinia chrysanthemi: the specific secretion functions are analogous to those of Escherichia coli A-hemolysin. EMBO J 9:1375–1382
Hui FM, Morrison DA (1991) Genetic transformation in Streptococcus pneumoniae: nucleotide sequence analysis shows comA, a gene required for competence induction, to be a member of the bacterial ATP-dependent transport protein family. J Bacteriol 173: 372–381
Diep DB, Håvarstein LS, Nes IF (1995) A bacteriocin-like peptide induces bacteriocin synthesis in Lactobacillus plantarum Cll. Food Microbiol 60:160–166
Håvarstein LS, Diep DB, Nes IF (1995) A family of bacteriocin ABC transporters carry out proteolytic processing of their substrates concomitant with export. Mol Microbiol 16: 229–240
Axelsson L, Holck A, Birkeland S-T, Aukrust T, Blom H (1993) Cloning and nucleotide sequence of a gene from Lactobacillus sake Lb706 necessary for sakacin A production and immunity. Appl Environ Microbiol 59: 2868–2875
Kornblum J, Kreiswirth B, Projan SJ, Ross H, Novick RP (1990) agr: a polycistronic locus regulating exoprotein synthesis in Staphylococcus aureus. In: Novick RP (ed) Molecular Biology of the Staphylococci. VCH Publishers, New York, pp 373–402
Morfeldt CI, Janzon L, Arvidson S, Löfdahl S (1988) Cloning of a chromosomal locus (exp) which regulates the expression of several exoprotein genes in Staphylococcus aureus. Mol Gen Genet. 211:1601–1607
Peng H, Novick RP, Kreitswirth B, Kornblum J, Schlievert P (1988) Cloning, characterization and sequencing of an accessory gene regulator (agr) in Staphylococcus aureus. J Bacteriol 170: 4365–4372
Muriana PM, Klaenhammer TR (1991b) Cloning, phenotypic expression and DNA sequence of the gene for lactacin F, an antimicrobial peptide produced by lactobacillus spp. J Bacteriol 173:1779–1788
Nissen-Meyer J, Larsen AG, Sletten K, Daeschel M, Nes IF (1993b) Purification and characterization of plantaricin A, a Lactobacillus plantarum bacteriocin whose activity depends on the action of two peptides. J Gen Microbiol 139:1973–1978
Saris EJ, Immonen T, Reis M, Sahl H (1996) Immunity of lantibiotics. Antonie van Leeuwenhoeh 69:151–159
Immonen Y, Ye S, Ra R, Qiao M, Paulin L, Saris P (1995) The codon usage of the nisin Z operon in Lactococcus lactis N8 suggests a non-lactococcal origin of the conjugative nisin-sucrose transposon. Sequence 5: 203–218
Qiao M, Immonen T, Koponen O, Saris PEJ (1995) The cellular location and effect on nisin immunity of the NisI protein from Lactococcus lactis N8 expressed in Escherichia coli and L.lact is. FEMS Microbiol Lett 131: 75–80
Reis M, Eschbach-Bludau M, Iglesias-Wind MI, Kupke T, Sahl H-G (1994) Producer immunity towards the lantibiotic Pep5: identification of the immunity gene pepI and localization and functional analysis of its gene product. Appl Environ Microbiol 60: 2876–2883
Meyer C, Beirbaum G, Heidrich C, Reis M, Suling J, Iglesias-Wind MI, Kempter C, Molitor E, Sahl HG (1995) Nucleotide sequence of the lantibiotic Pep 5 biosynthetic gene cluster and functional analysis of PepP and PepC. Evidence for a role in PepC in thioether formation. Eur J Biochem 232: 478–489
Quadri LEN, Sailer M, Roy KL, Vederas JC, Stiles ME (1994) Chemical and genetic characterization of bacteriocins produced by Carnobacterium piscicola LV17B. J Mol Biol 269:12204–12211
Tichaczek PS, Vogel RF, Hammes WP (1993) Cloning and sequencing of curA encoding curvacin A, the bacteriocin produced by Lactobacillus curvatus LTH1174. Arch Microbiol 160: 279–283
Tichaczek PS, Vogel RF, Hammes WP (1994) Cloning and sequencing of sakP encoding sakacinP, the bacteriocin produced by Lactobacillus sake LTH673. Microbiology 140: 361–367
Nissen-Meyer J, Håvarstein LS, Holo H, Sletten K, Nes IF (1993a) Association of the lactococcin A immunity factor with the cell membrane: purification and characterization of the immunity factor. J Gen Microbiol 139:1503–1509
Kok J, Venema K, Venema G (1995) Analysis of lactococcin secretion and immunity in Lactococcus lactis. In: Ferretti JJ, Gilmore MS, Klaenhammer TR (eds) Genetics of Streptococci, Enterococci and Lactococci. Karger, New York, pp 653–659
Peschel A, Götz F (1996) Analysis of the Staphylococcus epidermidis genes epiF,-E and-G involved in epidermin immunity. J Bacteriol 178: 531–536
Kerpolla RE, Shyamala VK, Klebba P, Ferro-Luzzi Ames G (1991) The membrane-bound proteins of periplasmic permeases form a complex. J Biol Chem 266: 9857–9865
Panagiotidis CH,R eyes M, Sievertsen A, Boos W, Shuman HA (1993) Characterization of the structural requirements for assembly and nucleotide binding of an ATP-binding cassette transporter. J Biol Chem 268: 23685–23696
Froseth BR, Herman RE, McKay LL (1988) Cloning of nisin resistance determinant and replication origin on 7.6-kilobase EcoRI fragment of pNP40 from Streptococcus lactis subsp. diacetylactis DRC3. Appl Environ Microbiol 54: 2136–2139
Froseth BR, McKay LL (1991) Molecular characterization of the nisin resistance region of Lactococcus lactis subsp. lactis biovar diacetylactis DRC3. Appl Environ Microbiol 57: 804–811
Jarvis B, Farr J (1971) Partial purification, specificity and mechanism of action of the nisin-inactivating enzyme from Bacillus cereus. Biochim Biophys Acta 227: 232–240
Hansen JN (1993) The molecular biology of nisin and its structural analogues. In: Hoover D, Steenson L (eds) Bacteriocins of Lactic Acid Bacteria. Academic Press, New York, pp 93–120
Bouret RB, Borkovich KA, Simon MI (1991) Signal transduction pathways involving protein phosphorylation in prokaryotes. Annu Rev Biochem 60: 401–441
Parkinson JS, Kofoid EC (1992) Communication modules in bacterial signaling proteins. Annu Rev Gen 26: 71–112
Kleerebezem M, Quadri IE, Kuipers OP, de Vos WM (1997) Quorum sensing by peptide pheromones and two-component signal-transductionsystems in Gram-positive bacteria. Mol Microbiol 24: 895–904
Huo L, Martin KJ, Schleif R (1988) Alternative loops regulate the arabinose operon in Escherichia coli. Proc Natl Acad Sci USA 85: 5444–5448
Igo MM, Losick R (1986) Regulation of a promoter that is utilized by minor forms of RNA polymerase holoenzyme in Bacillus subtilis. J Mol Biol 191: 615–624
Martin K, Huo L, Schleifer RF (1986) The DNA loop model for ara repression: AraC protein occupies the proposed loop sites in vivo and repression-negative mutations lie in these same sites. Proc Natl Acad Sci USA 83: 3654–3658
Ptashne M (1992) The genetic switch. Cell Press and Blackwell Scientific Publications, Cambridge
Cara JH, Schleif RF (1993) Variation of half-site organization and DNA looping by AraC protein. EMBO J 12: 35–44
Coleman, Bown GS, Stormonth DA (1975) A model for the regulation of bacterial extracellular enzyme and toxin biosynthesis. J Theor Biol 52:143–148
Janzon L, Löfdahl S, Arvidson S (1989) Identification and nucleotide sequence of the delta-lysin gene, hld, adjacent to the accessory gene regulator (agr) of Staphylococcus aureus. Mol Gen Genet 70: 337–349
Janzon L, Arvidson S (1990) The role of the d-lysin gene (hld) in the regulation of virulence genes by the accessory gene regulator (agr) in Staphylococcus aureus. EMBO J 9:1391–1399
Ji G, Beavis RC, Novick RP (1995) Cell density control of Staphylococcal virulence mediated by an octapeptide pheromone. Proc Natl Acad Sci USA 92:12055–12059
Sanders DA, Koshland DE Jr (1988) Receptor interactions through phosphorylation and methylation pathways in bacterial chemotaxis. Proc Natl Acad Sci USA 85: 8425–8429
Sanders DA, Gillece-Castro BL, Stock AM, Burlington AL, Koshland DE Jr (1989) Identification of the site of phosphorylation of the chemotaxis response regulator protein. J Biol Chem 264: 21770–21778
Kupke T, Gotz F (1996) post-translational modifications of lantibiotics. Antonie van Leeuwenhoek 69:139–150
Ingram LC (1970) A ribosomal mechanism for synthesis of peptides related to nisin. Biochim Biophys Acta 224: 263–265
Kellner J, Jung G, Josten M, Kaletta C, Entian K-D, Sahl H-G (1989) Pep5, a new lantibiotic: structure elucidation and amino acid sequence of the propeptide. Angew Chem Int Ed Engl 28: 616–619
van Kamp M, Horstink LM, van den Hooven HW, Konings RNH, Hilbers CW, Frey A, Sahl H-G, Metzger JW, van de Ven FJM (1995) Sequence analysis by NMR spectroscopy of the peptide lantibiotic epilancin K7 from Staphylococcus epidermidis K7. Eur J Biochem 227: 757–771
Schnell N, Entian K-D, Götz F, Hörner T, Kellner R, Jung G (1989) Structural gene isolation and prepeptide sequence of gallidermin, a new lanthionine containing antibiotic. FEMS Microbiol Lett 58: 263–268
Kupke T, Stevanovic S, Sahl H-G, Götz F (1992) Purification and characterization of EpiD, a flavoprotein involved in the biosynthesis of the lantibiotic epidermin. J Bacteriol 174: 5354–5361
Augustin J, Rosenstein R, Wieland B, Schneider U, Schnell N, Engelke G, Entian K-D, Gotz F (1992) Genetic analysis of epidermin biosynthetic genes and epidermin-negative mutants of Staphylococcus epidermidis. Eur J Biochem 204:1149–1154
Bishop L, Agbayani R, Ambudkar SV, Maloney PC, Ames GF-L (1989) Reconstitution of a bacterial periplasmic permease in proteoliposomes and demonstration of ATP hydrolysis concomitant with transport. Proc Natl Acad Sci USA 86: 6953–6957
Mimmack ML, Gallangher MP, Pearce SR, Hyde SC, Booth IR, Higgins CF (1989) Energy coupling to periplasmic binding protein-dependent transport in vivo. Proc Natl Acad Sci USA 86: 8257–8261
Chang YF, Young R, Struck DK (1991) The Actinobacillus pleuropneumonia haemolysin determinant: unlinked appCA and appBD loci flanked by pseudogenes. J Bacteriol 173: 5151–5158
Juranka P, Zhang F, Kulpa J, Endicott J, Blight M, Holland IB, Ling V (1992) Characterization of the haemolysin transporter, HlyB, using an epitope insertion. J Biol Chem 267: 3764–3770
Guthmiller JM, Kolodrubetz D, Cagle MP, Kraig E (1990) Sequence of the lktB gene from Actinobacillus actinomycetecomitans. Nucl Acids Res 18: 5291–5293
Glaser P, Sakamoto H, Bellalou J, Ullmann A, Danchin A (1988) Secretion of cyclolysin, the calmodulin-sensitive adenylate cyclase-haemolysin bifunctional protein of Bordetella pertussis. EMBO J 7: 3997–4004
Felmlee T, Pellett S, Lee E-Y, Welch RA (1985) Escherichia coli haemolysin is released extracellularly without cleavage of a signal peptide. J Bacteriol 163: 88–93
Wandersmann C (1992) Secretion across the bacterial outer membrane. Trends Genet 8: 317–322
Ross KF, Ronson CW, Tagg JR (1993) Isolation and characterization of the lantibiotic salivaricin A and its structural gene salA from Streptococcus salivarius 20P3. Appl Environ Microbiol 59: 2014–2021
Dinh T, Paulsen IT, Saier MH (1994) A family of extracytoplasmic proteins that allow transport of large molecules across the outer membranes of Gram-negative bacteria. J Bacteriol 176: 3825–3831
Skvirsky RC, Shoba R, Xiaoyu S (1995) Topology analysis of the colicin V export protein CvaA in Escherichia coli. J Bacteriol 177: 6153–6159
Schulein R, Gentschev I, Mollenkopf H-J, Goebel W (1992) A topological model for the haemolysin translocator protein HlyD. Mol Gen Genet 234:155–163
Muriana PM, Klaenhammer TR (1991a) Purification and partial characterization lactacin F, a bacteriocin produced by Lactobacillus acidophilus 11088. Appl Environ Microbiol 57:114–121
Allison GE, Ahn C, Stiles ME, Klaenhammer TR (1995a) Utilization of the leucocin export system in Leuconostoc gelidum for production of a Lactobacillus bacteriocin. FEMS Microbiol 131: 87–93
Allison GE, Worobo RW, Stiles ME, Klaenhammer TR (1995b) Heterologous expression of the lactacin F peptides by Carnobacterium piscicola LV17. Appl Environ Microbiol 61:1371–1377
Chikindas ML, Venema K, Ledeboer AM, Venema G, Kok J (1995) Expression of lactococcin A and pediocin PA-1 in heterologous hosts. Lett Appl Microbiol 21:183–189
Sahl HG, Bierbaum G (1998) Lantibiotics: biosynthesis and biological activities modified peptides from Gram-positive bacteria. Annu Rev Microbiol 52, 41–79
Tomita H, Fujimoto S, Tanimoto K, Ike Y (1996) Cloning and genetic organization of the bacteriocin 31 determinant encoded on the Enterococcus faecalis pheromone-responsive conjugative plasmid pY117. J Bacteriol 178: 3585–3593
Cintas LM, Casaus P, Haverstein LS, Hernandez PE, Nes IF (1997) Biochemical and genetic characterization of enterocin P, a novel sec-dependent bacteriocin from Enterococcus faecium P13 with a broad antimicrobial spectrum. Appl Environ Microbiol 63: 4321–4330
Cintas LM, Casaus P, Holo H, Hernandez PE, Nes IF, Havarstein LS (1998) Enterocins L50 A and L50B, two novel bacteriocins from Enterococcus faecium L50, are related staphylococcal hemolysins. J Bacteriol 180:1988–1994
Piard J-C, Kuipers OP, Rollema HS, Desmazeaud MJ, De Vos MJ (1993) Structure, organization and expression of the lct gene for lacticin 481, a novel lantibiotic produced by Lactococcus lactis. In: La lacticine 481, une nouvelle bacteriocine de type lantibiotique produite par Lactococcus lactis: characterization biochemique et genetique, Piard J-C (PhD Thesis): Universite de Caen, France, pp 85–116
Author information
Authors and Affiliations
Rights and permissions
Copyright information
© 2000 Springer-Verlag Berlin Heidelberg
About this chapter
Cite this chapter
Sablon, E., Contreras, B., Vandamme, E. (2000). Antimicrobial Peptides of Lactic Acid Bacteria: Mode of Action, Genetics and Biosynthesis. In: New Products and New Areas of Bioprocess Engineering. Advances in Biochemical Engineering/Biotechnology, vol 68. Springer, Berlin, Heidelberg. https://doi.org/10.1007/3-540-45564-7_2
Download citation
DOI: https://doi.org/10.1007/3-540-45564-7_2
Received:
Published:
Publisher Name: Springer, Berlin, Heidelberg
Print ISBN: 978-3-540-67362-0
Online ISBN: 978-3-540-45564-6
eBook Packages: Springer Book Archive