Abstract
Intracellular space is highly crowded with soluble and insoluble biomolecules that range from large polymers, such as proteins and nucleic acids, to small molecules, including metabolites and metal ions. It is therefore of interest to understand the effects of molecular crowding on the structure, stability, and function of biomolecules. Moreover, molecular crowding is observed not only intracellularly but also in the extracellular matrix and under the conditions used in in vitro biotechnological and nanotechnological processes. However, most biochemical studies of biomolecules are performed under dilute conditions. Recent studies have demonstrated critical effects of molecular crowding on nucleic acids. In the present study we discuss how molecular crowding affects the properties of G-quadruplexes as well as other non-canonical nucleic acid structures.
Graphical Abstract
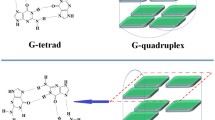
G-quadruplexes under molecular crowding conditions

Access provided by Autonomous University of Puebla. Download chapter PDF
Similar content being viewed by others
Keywords
1 Introduction
One of the ultimate goals of biochemical and biophysical studies is to reveal how biomolecules participate in fundamental biological processes in situ, i.e., inside living cells. Intracellular space is highly crowded with soluble and insoluble biomolecules that range from large polymers, such as proteins and nucleic acids, to small molecules, including metabolites and metal ions. Surprisingly, the total molecular concentration within cells reaches 400 mg mL−1 and biomolecules occupy up to 40% of the total intracellular space (Fig. 1) [1–4], creating conditions of molecular crowding. It is therefore of interest to understand the effects of molecular crowding on the structure, stability, and function of biomolecules. Molecular crowding has been demonstrated to affect the properties of biomolecules through effects on the thermodynamics and kinetics of macromolecular association and dissociation [5–7]. However, most biochemical studies are performed under non-physiological dilute conditions [8, 9], in contrast to other cellular environmental factors, such as temperature, pH, ion species and concentrations, and redox potential, which are adjusted during biochemical experiments to match physiological conditions. Since there are excellent reviews regarding molecular crowding effects on the structure and function of proteins and nucleic acids [1–6, 8–10], we discuss herein how molecular crowding affects the properties of G-quadruplexes and other non-canonical nucleic acid structures.
2 Molecular Crowding
Molecular crowding conditions are observed both intracellularly and in the extracellular matrix and also in in vitro biotechnological and nanotechnological processes. In this section we demonstrate how molecular crowding is critical in both in vivo and in vitro environments.
2.1 Molecular Crowding In Vivo
Molecular crowding occurs in both the nucleus and the cytoplasm where diagnostic target molecules exist and to where therapeutic molecules are targeted. Eukaryotic nuclei, in which DNA is present, are heterogeneous and contain a variety of subnuclear structures such as the nucleolus, splicing-factor compartments, Cajal bodies, promyelocytic leukemia bodies, replication factors, and transcription factors [11, 12]. In the nucleus, DNA is highly condensed and is packed with nuclear components to form chromosomes. DNA exists in a semi-ordered structure in these nuclear chromosomes, where it is wrapped around histones forming a composite material called chromatin (Fig. 2) [13, 14]. The nucleosome core consists of about 146 dsDNAs wrapped in left-handed superhelical turns around four identical pairs of proteins known as the histone octamer. DNA is then further packaged into metaphase chromosomes. Histone proteins regulate nucleosome assembly, DNA condensation, and DNA flexibility [15, 16]. Thus, the inside of the cell nucleus is crowded by molecules such as DNA and histones as well as by RNA, proteins, and small cosolutes.
Visualization of the cytoplasm, where RNA is present, by electron microscopy revealed the actin cytoskeleton, which was reconstructed without prior removal of membranes or extraction of soluble proteins [17, 18]. By using this technique, single macromolecular assemblies with distinct shapes, such as the proteasome and ribosomes, can be identified at a resolution of nanometers in an unperturbed cellular environment. Moreover, the cytoplasm of a living cell consists of a large number of additional molecules such as nucleic acids, proteins, carbohydrates, and lipids.
Therefore, the biomolecules occupy 20–40% of the total volume of the intracellular environment. The total concentration of intracellular biomolecules is estimated to be 300–400 mg mL−1, and the concentration is even higher in the mitochondrial matrix [1, 19–21]. The crowded nature of the cell interior can be visualized by comparison with lattice structures. In face-centered cubic lattices and simple cubic lattices, the atomic packing factors, which are the fractions of the volumes of crystal structures that are occupied by atoms, are 0.74 and 0.52, respectively (Fig. 3). The atomic packing factor of a diamond crystal is just 0.34, which is comparable with the fractional volume commonly occupied by biomolecules in living cells. Molecular crowding in living organisms involves high concentrations not only of macromolecules but also of small molecules. The concentration of such small molecules, which act as osmolytes, can reach the molar range. The most extreme example in mammals is the kidney medulla cells, which contain a urea concentration of up to 5.4 mol L−1 [22], which corresponds to 30% of the cell by mass. Thus, not only macromolecules but also small organic molecules are indispensable for maintaining homeostasis in all living organisms [23, 24].
2.2 Molecular Crowding In Vitro
It is obvious that molecular reactions that are designed to take place on a material surface in pharmaceutical and medicinal devices also proceed under conditions of high molecular crowding. Typical in vitro molecular crowding can be observed at liquid–solid interfaces, such as when a biomolecule is immobilized on a solid surface as occurs in analytical methods such as affinity chromatography, enzyme-linked immunosorbent assay (ELISA), and surface plasmon resonance (SPR). In such biosensing techniques, biomolecules are directly or indirectly attached to solid supports via covalent or non-covalent interactions (Fig. 4). When a DNA strand is directly immobilized, it is either surrounded by identical DNA strands or is on a self-assembled monolayer of an organic polymer that prevents non-specific adsorption of molecules onto the solid support. In the case of the direct immobilization of DNA strands onto gold nanoparticles, the density of the DNA strands was reported to be around 50 pmol cm−2 [25–27], resulting in around 3 × 1013 DNA strands cm−2. Since the diameter of DNA is 2 nm, the DNA strands cover around 90% of the surface, indicating that there is almost no available free surface after hybridization of the DNAs with their target complementary DNA strands. This demonstrates the molecular crowding conditions of immobilized biomolecules. It is therefore possible that the hybridization process that occurs at the solution–solid interface is different from the process occurring in a dilute solution, and that this difference is at least partly due to molecular crowding.
In nanodevices, the distinctive chemical conditions at the solution-solid interface further affect the properties of the whole system. A nanopore, whose diameter generally ranges from several to several ten of nanometers, can be created by a membrane protein or by a combination of lithography and ultrathin materials such as silicon and graphene [28–30]. At the nanoscale level, the chemical condition at the liquid–solid interface that is not observed in bulk solution dominates the characteristics of the whole solution. Since such nanodevices have been applied to the detection, separation, and purification of biomolecules [31], it is essential to investigate the biomolecular properties at the solution–solid interface where the surface and immobilized biomolecules create molecular crowding conditions.
2.3 Molecular Crowding Reagents
Because it is difficult to investigate quantitatively nucleic acid interactions in a living cell and on material surfaces, in vitro experimental systems that use a synthetic cosolute have been widely used to study biomolecular reactions under non-aqueous conditions. Such crowding cosolutes should meet the following criteria [10]: (1) they should be basically inert so that there is no chemical interaction between the target molecule and the crowding cosolute; (2) they should easily dissolve in water to stimulate molecular crowding; (3) in the case of large cosolutes, different polymer sizes should be available; and, (4) in the case of small cosolutes, different chemical properties should be available. Inert cosolutes in particular are convenient for the study of molecular environments. Thus neutral and highly water-soluble molecules are extremely useful cosolutes for quantitative studies. However, chemical interactions including electrostatic interactions between the target molecule and cosolutes should be taken into account when investigating intracellular biomolecular function [32, 33]. Poly(ethylene glycol) (PEG) and polysaccharides are often used to mimic molecular crowding conditions since they are inert to most biomolecules. These polymers can be dissolved in water to a relatively high concentration, and different polymer sizes are commercially available. Other small cosolutes that are useful for mimicking molecular crowding conditions are alcohols, glycols, amino acids, acetonitrile, trimethylamine N-oxide, and betaines [34–36]. Moreover, proteins such as chymotrypsin, albumin, hemoglobin, and lysozyme, as well as synthetic polymers such as poly(vinylpyrrolidone)s, are also utilized as crowding cosolutes to mimic better the chemical conditions inside living cells [37].
3 Molecular Crowding Effects on Non-canonical Structures of Nucleic Acids
The critical role of the nucleic acids, DNA and RNA, is to store and process the genetic information that includes all of the information that is necessary for life. In 1953, Watson and Crick demonstrated that DNA forms a double helix via Watson–Crick base pairs. Half a century after the discovery of the double helix, the Human Genome Project (HGP) identified the almost 3.2 billion base pairs in the entire human genome. The HGP also showed that repetitive DNA sequences, including dinucleotide and trinucleotide repeats, as well as telomeric and centromeric sequences, are widely distributed in the human genome. Most of this repetitive DNA can potentially fold into noncanonical structures. From this perspective, clarifying not only the primary structure (sequence) of nucleic acids but also thermodynamic and kinetic analyses of the higher-order structures and structure-function relationships of nucleic acids are essential for understanding the reactions that underlie biological processes and diseases. Thus, molecular crowding effects on the non-canonical structures of nucleic acids are of interest for a broad range of research fields. In this section, we will introduce the effects of molecular crowding on some of the non-canonical structures of nucleic acids.
3.1 DNA Triplexes
Spink and Chaires as well as Goobes and Minsky demonstrated that the Hoogsteen base pairs in a DNA triplex are generally stabilized by molecular crowding, whereas Watson–Crick base pairs are destabilized [38, 39]. These studies also suggested that the thermodynamics of DNA duplexes and triplexes are regulated by DNA hydration. Minsky et al. further estimated the thermodynamic parameters of the triplex formation of T18·A20·T20 and found that stabilization of the DNA triplex by PEG was driven by a large negative enthalpy change that exceeded the unfavorable entropy change [40]. Based on the parameters measured under conditions of various salt concentrations and temperatures, they proposed that alterations in environmental conditions could be effectively compensated for by crowding cosolutes; this compensation provides a mechanism for adaptation to, and buffering against, unfavorable conditions such as unfavorable ionic strength or temperature [40].
3.2 DNA Junctions
Junctions in nucleic acid structures arise when three or more helices meet at a single point (Fig. 5a). These branched junctions are important intermediates in many biological functions and play important roles in many cellular processes [41]. Three-way junctions (TWJs) are the simplest type of junction and are used as a model system to gain insight into other complex and multi-branched junction structures [42]. TWJs create a unique electrostatic environment near the junction point due to the close proximity of opposing charges. The effects of molecular crowding on the formation of TWJs of DNA have been systematically studied [43]. TWJs, consisting of the junction point and three helical duplex arms, were destabilized by molecular crowding both in the absence and presence of 5 mM Mg2+. However, the difference in the number of water molecules that were taken up by the TWJ as a whole and by each helical arm demonstrated that water molecules were released from the junction point. This dehydration upon the junction point formation suggested the stabilization of the junction point under the molecular crowding conditions. In fact, molecular crowding induced the structural transition from a bimolecular duplex to the intramolecular TWJ, even in the absence of Mg2+ (Fig. 5b). This finding demonstrates that intracellular conditions, under which water activity decreases and hydration becomes less favorable, facilitate the formation of TWJ structures rather than bimolecular duplex structures.
3.3 RNA Structures and Functions
There have been several studies of molecular crowding effects on high-order RNA structures and their catalytic activity. It was reported that methanol stabilized a 58-nt RNA structure but not RNA secondary structure [44]. Addition of the osmolyte trimethylamine-oxide (TMAO) enhanced the efficiency of ribosome reconstitution by up to 100-fold, providing a substantially improved system for the in vitro analysis of mutant ribosomes [45]. These findings demonstrated that cosolutes play an important role in stabilizing high-order RNA structures and their interactions with other biomolecules. This effect may be due to the stabilizing effect of TMAO on RNA tertiary structure. Indeed, TMAO can counteract the denaturing effects of urea on tRNA tertiary structure [46]. In order to understand the basis for the effects of TMAO on RNA structures, Draper and coworkers quantified the TMAO-induced stabilization of RNAs. They suggested that the formation of RNA tertiary structure is accompanied by substantial dehydration of the phosphate groups, and that TMAO affects this process by reducing the energetic penalty associated with this dehydration [47]. These results were supported by molecular dynamic simulations of a 22-nt RNA hairpin in aqueous TMAO solution [48]. Further reports of molecular simulations indicated that molecular crowding enhances the thermodynamic stability of a pseudoknot in human telomerase RNA, leading to a structural switch from a hairpin to a pseudoknot of human telomerase RNA [49].
It was reported that the RNA cleavage activity of a hairpin ribozyme increases in the presence of PEG 400 or dextran 10000 [50, 51]. The RNA compaction accompanied by water release that occurs during the folding of the hairpin ribozyme into the active conformation was proposed to be responsible for the enhanced cleavage activity under molecular crowding conditions. It was also found that the formation of a tertiary RNA structure of the GAAA tetraloop-receptor RNA tertiary motif that was observed in the group I ribozyme domain was favored by molecular crowding induced by higher molecular weight PEG and dextran but was not induced by lower molecular weight sucrose and glycerol [52]. Small angle X-ray scattering experiments showed that PEG favors more compact RNA structures and folding at low Mg2+ concentrations because of the excluded volume effects of molecular crowding [53]. Although the presence of the cosolutes decreased the thermal stability of the ribozyme stem (duplex) regions, the structure as a whole was stabilized by molecular crowding. More importantly, it was found that molecular crowding induced by PEG decreased the metal ion concentration that was required for the catalytic activity of a hammerhead ribozyme, facilitated catalytic turnover, and activated a ribozyme that was in inactive form [54]. This ribozyme also showed efficient activity in a solution at physiological salt concentration only under osmotic pressure induced by added cosolutes. The fact that the ribozyme functioned efficiently under physiological conditions implies that ribozymes have the potential to work efficiently within a living cell. These results indicate that molecular crowding is critical not only to maintain RNA structures but also to maintain their functions in living cells where there exist many biomolecules that can destabilize RNA structures.
4 Molecular Crowding Effects on G-Quadruplex Conformations
G-quadruplexes have a unique polymorphic nature [55–59]. The polymorphism of G-quadruplex conformation is induced and regulated not only by nucleotide sequence but also by chemical conditions. Since a G-quadruplex is stabilized by cation coordination to the center of the G-quartet, the effects of cations on the conformation and thermal stability of G-quadruplexes have been studied [60–69]. Using NMR, Wang and Patel were the first to show that the human telomeric DNA sequence, AG3(TTAG3)3, folded to form an antiparallel G-quadruplex in the presence of Na+ [70]. Subsequently, Parkinson et al., using X-ray crystallography, reported the formation of a parallel G-quadruplex of AG3(TTAG3)3 in the presence of K+ [71]. There are now at least five types of intermolecular G-quadruplexes known for human telomeric DNA sequences that contain TTAGGG repeats. NMR and chemical modification studies have shown the formation of mixed G-quadruplexes [(3+1) G-quadruplexes or hybrid G-quadruplexes] that involve Form 1 and Form 2 [72–74]. These studies have demonstrated that chemical conditions as well as nucleotide sequences should be taken into account when determining native G-quadruplex conformations in living cells. In addition to the effects of cation species and their concentration, it is now widely accepted that molecular crowding critically affects the structural polymorphism of G-quadruplexes. In this section we will discuss how molecular crowding regulates the conformation and the thermal stability of G-quadruplexes.
4.1 G-Quadruplex Conformations Under Molecular Crowding Conditions
The effect of molecular crowding on G-quadruplex conformations was first studied using the Oxytricha nova telomeric DNA sequence dG4T4G4, in the presence of Na+ (Table 1). Molecular crowding conditions were created by use of both the neutral cosolutes PEG (MW = 300) and glycerol and the positively charged cosolutes putrescine, cadaverine, and spermidine [75]. It was found that molecular crowding induced by PEG induced a conformational transition from an antiparallel to a parallel conformation of the d(G4T4G4) and d(G4T4)3 G4 G-quadruplexes (Fig. 6a), whereas molecular crowding using the cationic cosolutes did not alter the antiparallel G-quadruplex conformation. These results suggested that the conformational switch of G-quadruplexes is regulated by molecular crowding.
In 2005, Chaires and coworkers reported molecular crowding effects on the G-quadruplex conformation of the human telomeric DNA sequence dA(G3TTA)4 in the presence of Na+ and K+ [76]. Circular dichroism and fluorescence quenching studies showed dramatic changes in the G-quadruplex conformation under molecular crowding conditions in the presence of K+ but not in the presence of Na+. These results, as well as the results obtained for the O. nova telomeric DNA, indicated that molecular crowding effects on G-quadruplex conformations are dependent on the nature of the coexisting cation and on the nucleotide sequence. To study the polymorphic nature of DNA G-quadruplexes under molecular crowding conditions, the effects of the telomeric DNA sequence on the intermolecular and intramolecular G-quadruplexes formed from Tetrahymena, human, and O. nova telomeric sequences were studied under dilute and molecular crowding conditions in the absence and presence of various cosolutes [77]. The intermolecular and intramolecular G-quadruplex structures formed from Tetrahymena telomeric DNA formed very long, well-ordered G-wires in the presence of cosolutes in Na+ solution. Although this structure is adopted by many parallel-stranded telomeric sequences, the intermolecular and intramolecular G-quadruplex structures formed from human telomeric sequences remained as compact antiparallel G-quadruplex structures under these conditions (Fig. 6b). These results demonstrate that a single G to A base replacement in the loops of a G-quadruplex leads to a dramatically different structure under molecular crowding conditions in the presence of Na+. Conversely, the parallel conformation of a G-quadruplex formed from human telomeric DNA was observed in a K+-containing solution under molecular crowding conditions [78]. More complex behavior of the human telomeric DNA under molecular crowding conditions has also been reported: in the presence of K+, molecular crowding using ethanol induced a conformational transition of human telomeric DNA from the antiparallel to the parallel conformation via a mixed conformation [80]. Such structural conversion of the human telomeric DNA by molecular crowding was also observed in the absence of a monovalent cation [79] and, furthermore, was observed for long telomeric sequences containing two G-quadruplex units [81]. Heddi and Phan used NMR to study how molecular crowding affects the human telomeric DNAs that had folded into the four different conformations under dilute conditions in the presence of K+ [82]. The NMR spectra clearly demonstrated that the four different G-quadruplex conformations were all converted to the parallel G-quadruplex conformation under molecular crowding conditions induced by PEG 200 in the presence of K+. Notably, this parallel conformation was almost identical to the parallel conformation observed by X-ray crystallography under dilute conditions with K+. Furthermore, the addition of other the neutral cosolutes PEG 8000, dimethyl sulfoxide, ethanol, and acetonitrile also induced a similar conformation as that induced by PEG 200. These NMR results indicate that molecular crowding simplifies the polymorphic nature of G-quadruplexes, depending on the nucleotide sequence of the G-quadruplex.
4.2 Molecular Crowding Induced by Different Cosolutes
Although molecular crowding generally favors the parallel conformation of a G-quadruplex, the above results indicated that the molecular crowding effects on G-quadruplex conformation are highly dependent, not only on the nucleotide sequence of the G-quadruplex and the nature of the co-existing cation species but also on the cosolute species. From this perspective, a vertebrate telomeric DNA G-quadruplex was studied under molecular crowding conditions within a Xenopus laevis egg extract as well as in the presence of PEG 200 and Ficoll70 [83]. The conformation of the vertebrate telomeric DNA in the X. laevis egg extract or in Ficoll was different from that observed in the presence of PEG. Based on these results, the authors stated that PEG should not be used to mimic molecular crowding conditions. However, the usage of PEG to induce molecular crowding conditions can provide important information for evaluation of quantitative parameters including thermodynamic parameters of G-quadruplex structure and the hydration state of G-quadruplexes, as will be discussed below. Moreover, Trent and coworkers, using acetonitrile as a non-hydrogen-bonding dehydrating agent, performed CD and NMR studies of the conformation of human telomeric DNA in K+ and Na+ solutions [85]. They showed that, although the CD spectra obtained using acetonitrile were similar to those obtained using PEG 400, the NMR spectra obtained using these reagents were not similar to each other, indicating that assignment of G-quadruplex conformations based only on CD spectra is not sufficient. Interestingly, the NMR spectrum obtained under the molecular crowding condition induced by bovine serum albumin was similar to that obtained under the dilute condition.
4.3 Duplex–Quadruplex Competitions
It has been considered that most telomeric DNAs fold into a duplex with Watson–Crick base pairing (Fig. 7a). However, it remains unclear whether duplex formation is dominant in the G-rich/C-rich regions since these G-rich and C-rich regions can individually form quadruplex structures [86]. Thus, molecular crowding effects on duplex–quadruplex competition are of interest in terms of predicting the native structure of such regions. It was demonstrated that a duplex formed with G- and C-rich DNAs under a dilute condition in the presence of Na+ was dissociated upon molecular crowding and that the G- and C-rich DNA sequences folded into individual quadruplexes (Fig. 7b) [87]. Tan and coworkers also reported that molecular crowding induced telomere G-quadruplex formation in a salt-deficient solution and in a K+-containing solution and that it enhanced the competition between G-quadruplex formation and duplex formation [88, 89]. Kinetic studies of G-quadruplex formation from a duplex with the complementary sequence showed that the formation rate of a G-quadruplex depends on the PEG and the K+ concentration [90]. A quantitative analysis showed that, when 30 nmol L−1 of G-rich and C-rich human telomeric DNAs were mixed together in a 100 mM K+ solution, the amount of the G-quadruplex formed was 17.6 and 23.4 nmol L−1 in the absence and presence of 10% ethylene glycol, respectively [91]. As shown using these short oligonucleotides, molecular crowding made it possible for a G-rich sequence to form a stable G-quadruplex in a long double-stranded DNA [92]. These results indicate that molecular crowding is essential for G-quadruplex formation in the G-rich/C-rich double stranded region. In such regions, the polymorphic nature of the G-quadruplex can be induced by molecular crowding in vivo. These results also illustrate the difficulty of correctly extrapolating the structures of not only telomeric DNA sequences but also of many other biomolecules in vivo based on their structures in vitro. Thus, molecular crowding as well as other cellular environmental factors [93–96] play critical roles in the structure of a biomolecule and analysis of their effects is useful for understanding biomolecular behaviors in vivo and for efficient drug and ligand targeting by biomolecules.
(a) Schematic illustration of a telomeric region composed of G-rich and C-rich strands. (b) Structures of the telomeric DNAs under dilute and molecular crowding conditions. Under dilute and molecular crowding conditions the 1:1 mixture of the G-rich and C-rich strands folds into a duplex and quadruplexes, respectively
5 Molecular Crowding Effects on the Thermal Stability of G-Quadruplexes
As discussed above, molecular crowding stabilizes non-canonical structures of nucleic acids that involve DNA triplexes, DNA junctions, and RNA tertiary structures, and strongly stabilizes non-canonical G-quadruplexes. Moreover, molecular crowding creates and regulates the conformational changes between different types of G-quadruplexes, and between a G-quadruplex and a duplex. In this section we will discuss how molecular crowding stabilizes G-quadruplex structures, resulting in dynamic conformational transitions.
5.1 Thermodynamics of G-Quadruplexes Under Molecular Crowding Conditions
Table 2 summarizes the thermodynamic parameters of G-quadruplex formations under dilute and molecular crowding conditions. It is widely accepted that molecular crowding significantly stabilizes G-quadruplexes. In order to show the differences in molecular crowding effects on a non-canonical G-quadruplex and a canonical duplex, the thermodynamic parameters of the intramolecular antiparallel G-quadruplex formed by a thrombin binding aptamer (TBA) and those of an intramolecular antiparallel hairpin-looped duplex were evaluated under molecular crowding conditions induced by various neutral cosolutes [84]. It was quantitatively demonstrated that the antiparallel G-quadruplex is stabilized under molecular crowding conditions at various concentrations of PEG 200 but that these conditions destabilize the duplex. The melting temperature (T m) for 5 μmol L−1 of the antiparallel G-quadruplex increased from 54.1 °C to 58.7 °C as the PEG 200 concentration was increased from 0 to 40 wt% in a K+-containing solution. In contrast to the G-quadruplex, the T m for 5 μmol L−1 of the duplex in the presence of K+ decreased from 66.4 °C to 54.3 °C as the PEG 200 concentration was increased from 0 to 40 wt%. When the PEG 200 concentration was increased from 0 to 40 wt%, the values of ΔH˚ (enthalpy change), TΔS˚ (entropy change), and ΔG˚25 (free energy change at 25 °C) of the antiparallel G-quadruplex decreased from −42.0 to −53.0 kcal mol−1, from −38.5 to −47.5 kcal mol−1, and from −3.5 to −5.5 kcal mol−1, respectively. The difference in ΔG˚25 induced by molecular crowding was −2.0 kcal mol−1. On the other hand, the values of ΔH˚, TΔS˚, and ΔG˚25 for formation of the duplex increased from −81.5 to −75.8 kcal mol−1, from −71.7 to −68.9 kcal mol−1, and from −9.8 to −6.9 kcal mol−1, respectively, with the same change in PEG 200 concentration. These changes indicated that promotion of the G-quadruplex formation by molecular crowding was enhanced by a favorable enthalpic contribution that exceeded an unfavorable entropic contribution. Conversely, the duplex destabilization was due to an unfavorable enthalpic contribution that exceeded a favorable entropic contribution. Such an enhancement in the thermodynamic stability of G-quadruplexes by molecular crowding has been reported for various G-rich nucleotide sequences. In particular, the G-quadruplex formed by the sequence dG3T3G3TG3T3G3 was stabilized 3.5 kcal mol−1 in ΔG˚25 by 30% ethylene glycol in a K+ solution, which corresponded to more than two orders of difference in the equilibrium constant [97]. Stabilization effects were also reported for the structures formed by the human telomeric DNA [79, 85] and by bcl-2 DNA [98]. It is noteworthy that the stabilization and destabilization of DNA structures do not depend on their structure but instead depend on the type of base pairs that are involved in their structure [99]. In addition, it was recently reported that the Hoogsteen base pairs in triplex and G-quadruplex DNA structures were stabilized not only by molecular crowding but also by a model peptide that mimicked histone H3, which is critical for the formation of a high-order complex called a nucleosome that involves a DNA strand and histone octamers (Fig. 2) [100]. Since nucleosomes become further organized to form chromatin inside the eukaryotic cell nucleus, and since chromatin structure is dynamic and controls gene expression, the stabilization of G-quadruplexes and triplexes by a histone-mimicking peptide implies roles for such non-canonical structures in transcription. The stabilization of G-quadruplexes by molecular crowding supports both the formation of G-quadruplexes and the biological roles of G-quadruplexes within living cells.
5.2 Hydration of G-Quadruplexes
Water molecules play critical roles in generating and maintaining the structure, stability, and function of biomolecules. It is well known that many proteins unfold in non-aqueous solution and lose their functions in the absence of water. Nucleotides have a large number of hydration sites. The 11–12 hydration sites of a nucleotide are occupied by water molecules that directly interact with the nucleotide and form a primary hydration shell in which 8–9 water molecules per nucleotide are bound to the primary hydration shell [101]. These water molecules play fundamental roles in maintaining the structure, stability, and function of nucleic acids.
The use of the osmotic stress technique that uses an osmolyte as a cosolute provides information regarding biomolecular interactions with water molecules. Such analysis has revealed differences in the numbers of water molecules bound, −Δn w, which are evaluated as follows: –Δn w = n w,folded − n w,unfolded, where n w,folded and n w,unfolded indicate the numbers of water molecules bound to the folded and unfolded states, respectively. In general, an osmolyte that induces molecular crowding lowers the activity of water, a w, and thus decreases the chemical potential of water. Therefore, cosolutes affect the equilibrium constant of biomolecular reactions that involve the association or dissociation of water molecules. Indeed, water molecules participate in most biochemical reactions. The dependency of water activity on the equilibrium constant K of a biomolecular reaction reflects the number of water molecules released upon the reaction Δn w as presented by the equation (∂log K/∂log a W) = −Δn w, if other interactions between the cosolute and the nucleic acid, and between the cosolute and the water molecule, are negligible [102]. This technique can be applied to any equilibrium and parameters to evaluate how many water molecules participate in the reactions.
The osmotic stress method has been applied to the evaluation of the hydration state of G-quadruplexes. UV melting studies of the TBA in a K+ solution at 0 and 40 wt% PEG 200 showed that molecular crowding with PEG 200 stabilizes the TBA G-quadruplex structure [84]. Thermodynamic parameters including the equilibrium constant K obs can be evaluated based on the melting curves. Plots of ln K obs vs. ln a w can be drawn based on osmotic pressure measurements. Both of these K obs have a linear relationship with ln a w, and the slope of this line corresponds to the number of water molecules released upon structure formation, showing that water molecules are released upon the formation of G-quadruplexes. In contrast, water molecules are taken up upon duplex formation. It was quantitatively calculated that 4.5 and 4.0 water molecules per nucleotide were released upon G-quadruplex formation. Conversely, in solutions of K+ and Na+, 3.4 and 3.5 water molecules per nucleotide were taken up upon duplex formation.
The numbers of water molecules released upon G-quadruplex formation have been reported for various sequences (Table 3). Marky and coworkers reported values of Δn w for DNA aptamers, NHE-III, and a human telomere [103] in a K+ solution. They calculated that 0.1–1.5 water molecules were released per nucleotide through G-quadruplex folding depending on the nucleotide sequence. It was proposed that these values result from (1) the release of structural water from the random coil state upon formation of the G-quadruplex, (2) the uptake of electrostricted water molecules by G-quadruplexes with a higher charge density, and (3) the release of electrostricted water from K+ upon binding to the G-quadruplex core (G-quartet). Notably, they further separately evaluated Δn w for the loop and G-quartet (G-quadruplex core) regions using various substitutions [104] and reported that Δn w = 13 for the G-quartet and Δn w = −6∼+8 for the remaining loop region. In addition, by comparison of the Δn w values of not only G-quadruplexes but also of duplexes and triplexes, it was proposed that the opposing effects of molecular crowding on canonical structures (duplexes) and on non-canonical structures (G-quadruplexes and triplexes) were due to different behaviors of water molecules binding to the DNA strands [99]. Trent and coworkers further attempted to assess experimentally the role of steric crowding (excluded volume) and hydration on the structure of a human telomeric G-quadruplex. Using BSA and acetonitrile to induce molecular crowding conditions, it was shown that the excluded volume effect on G-quadruplex conformation under molecular crowding conditions induced by acetonitrile is small and that hydration is the dominant factor for determination of the conformation and stability of a G-quadruplex [85]. Although the number of water molecules released upon G-quadruplex formation varied from 0.14 to 8.5 depending on the experimental conditions and procedure used as well as on the nucleotide sequence and the conformation of the G-quadruplex (Table 3), the dehydration of the G-quadruplex through structure formation is clearly observed, and this dehydration should stabilize the G-quadruplex under cell-mimicking conditions.
6 Conclusions and Perspectives: Making G-Quadruplexes More Canonical with Molecular Crowding
In the present review we have introduced and discussed how molecular crowding alters the conformation and thermodynamics of nucleic acids with structures ranging from a canonical duplex to a non-canonical G-quadruplex. Changes in nucleic acid structures and their stabilities affect various enzymes including nucleases, polymerases, telomerases, and helicases [78, 106–110]. Although the molecular crowding effects on enzyme functions are not yet sufficiently understood, it is now generally accepted that molecular crowding is one of the most important players in the regulation of various biological processes via stabilization or destabilization of nucleic acid structures and enhancement or suppression of nucleic acid functions. In particular, molecular crowding effects on G-quadruplexes should be taken into account when considering the biological functions of G-quadruplexes because of the polymorphic nature of their conformation, thermal stability, and dynamics. In fact, the properties of G-quadruplexes under molecular crowding conditions critically affect ligand and drug designs. Tan and coworkers studied molecular crowding effects on the functions of G-quadruplex ligands in terms of their effects on the affinity of ligand binding to G-quadruplexes and on the efficiency of these ligands for inhibition of telomerase activity [111]. They found that, under molecular crowding conditions, the ligands TMPyP4, BMVC, and Hoechst 33258 became significantly less effective, or lost the ability to stabilize the G-quadruplex and to inhibit telomerase activity. On the other hand, recent studies showed that TMpyP4 bound to G-quadruplex structures with higher affinity than to duplexes [112]. The binding of TMPyP4 to G-quadruplexes of human and O. nova telomeric DNAs under molecular crowding conditions has also been reported [113, 114]. Moreover, some G-quadruplex ligands that can bind to G-quadruplexes with high affinity and specificity under molecular crowding conditions have been found [115, 116]. It will therefore be necessary to determine how molecular crowding affects G-quadruplex ligand functions and how we can rationally design G-quadruplex ligands so that they function under the conditions within living cells.
Molecular crowding effects on biomolecules are also important for various in vitro technologies, including drug delivery systems [117], the dispersion of carbon nanomaterials [118, 119], nanoparticle assembly [120, 121], response enhancement of electrochemical DNA sensors [122], DNA strand exchange [123], and the manipulation of single DNA molecules [124]. These studies clearly demonstrate that molecular crowding is a useful chemical stimulus for controlling the properties of nucleic acids toward in vitro applications. Since molecular crowding destabilizes canonical duplexes but stabilizes non-canonical structures especially G-quadruplexes, it will be possible to make G-quadruplex more canonical structure of nucleic acids with molecular crowding both in vivo and in vitro.
References
Zimmerman SB, Trach SO (1991) Estimation of macromolecule concentrations and excluded volume effects for the cytoplasm of Escherichia coli. J Mol Biol 222:599–620
Parsegian VA, Rand RP, Rau DC (2000) Osmotic stress, crowding, preferential hydration, and binding: a comparison of perspectives. Proc Natl Acad Sci USA 97:3987–3992
Minton AP (2001) The influence of macromolecular crowding and macromolecular confinement on biochemical reactions in physiological media. J Biol Chem 276:10577–10580
Ellis RJ, Minton AP (2003) Join the crowd. Nature 425:27–28
Zimmerman SB, Minton AP (1993) Macromolecular crowding: biochemical, biophysical, and physiological consequences. Annu Rev Biophys Biomol Struct 22:27–65
Zhou H-X, Rivas G, Minton AP (2008) Macromolecular crowding and confinement: biochemical, biophysical, and potential physiological consequences. Annu Rev Biophys 37:375–397
Lukacs GL, Haggie P, Seksek O, Lechardeur D, Freedman N, Verkman AS (2000) Size-dependent DNA mobility in cytoplasm and nucleus. J Biol Chem 275:1625–1629
Ellis RJ (2001) Macromolecular crowding: obvious but underappreciated. Trends Biochem Sci 26:597–604
Ellis RJ (2001) Macromolecular crowding: an important but neglected aspect of the intracellular environment. Curr Opin Struct Biol 11:114–119
Miyoshi D, Sugimoto N (2008) Molecular crowding effects on structure and stability of DNA. Biochimie 90:1040–1051
Lewis JD, Tollervey C (2000) Like attracts like: getting RNA processing together in the nucleus. Science 288:1385–1389
O’Brien TP, Bult CJ, Cremer C, Grunze M, Knowles BB, Langowski J, McNally J, Pederson T, Politz JC, Pombo A, Schmahl G, Spatz JP, van Driel R (2003) Genome function and nuclear architecture: from gene expression to nanoscience. Genome Res 13:1029–1041
Luger K, Mäder AW, Richmond RK, Sargent DF, Richmond TJ (1997) Crystal structure of the nucleosome core particle at 2.8 A resolution. Nature 389:251–260
Richmond TJ, Davey CA (2003) The structure of DNA in the nucleosome core. Nature 423:145–150
Narlikar GJ, Fan H-Y, Kingston RE (2002) Cooperation between complexes that regulate chromatin structure and transcription. Cell 108:475–487
Bloomfield VA (1996) DNA condensation. Curr Opin Struct Biol 6:334–341
Beck M, Lucić V, Förster F, Baumeister W, Medalia O (2007) Snapshots of nuclear pore complexes in action captured by cryo-electron tomography. Nature 449:611–615
Medalia O, Weber I, Frangakis AS, Nicastro D, Gerisch G, Baumeister W (2002) Macromolecular architecture in eukaryotic cells visualized by cryoelectron tomography. Science 298:1209–1213
Srere PA (1981) Protein crystals as a model for mitochondrial matrix proteins. Trends Biochem Sci 6:4–7
Fulton AB (1982) How crowded is the cytoplasm? Cell 30:345–347
Goodsell DS (1991) Inside a living cell. Trends Biochem Sci 16:203–206
MacMillen RE, Lee AK (1967) Australian desert mice: independence of exogenous water. Science 158:383–385
Yancey PH, Clark ME, Hand SC, Bowlus RD, Somero GN (1982) Living with water stress: evolution of osmolyte systems. Science 217:1214–1222
Lang F, Busch GL, Ritter M, Völkl H, Waldegger S, Gulbins E, Häussinger D (1998) Functional significance of cell volume regulatory mechanisms. Physiol Rev 78:247–306
Akamatsu K, Kimura M, Shibata Y, Nakano S, Miyoshi D, Nawafune H, Sugimoto N (2006) A DNA duplex with extremely enhanced thermal stability based on controlled immobilization on gold nanoparticles. Nano Lett 6:491–495
Sato K, Hosokawa K, Maeda M (2003) Rapid aggregation of gold nanoparticles induced by non-cross-linking DNA hybridization. J Am Chem Soc 125:8102–8103
Demers LM, Mirkin CA, Mucic RC, Reynolds RA 3rd, Letsinger RL, Elghanian R, Viswanadham G (2000) A fluorescence-based method for determining the surface coverage and hybridization efficiency of thiol-capped oligonucleotides bound to gold thin films and nanoparticles. Anal Chem 72:5535–5541
Bayley H (2009) Membrane-protein structure: piercing insights. Nature 459:651–652
Garaj S, Hubbard W, Reina A, Kong J, Branton D, Golovchenko J (2010) Graphene as a sub-nanometer trans-electrode membrane. Nature 467:190–193
Storm AJ, Chen JH, Ling XS, Zandbergen HW, Dekker C (2003) Fabrication of solid-state nanopores with single-nanometre precision. Nat Mater 2:537–540
Stroeve P, Ileri N (2011) Biotechnical and other applications of nanoporous membranes. Trends Biotechnol 29:259–266
Capp MW, Pegram LM, Saecker RM, Kratz M, Riccardi D, Wendorff T, Cannon JG, Record MT Jr (2009) Interactions of the osmolyte glycine betaine with molecular surfaces in water: thermodynamics, structural interpretation, and prediction of m-values. Biochemistry 48:10372–10379
Knowles DB, LaCroix AS, Deines NF, Shkel I, Record MT Jr (2011) Separation of preferential interaction and excluded volume effects on DNA duplex and hairpin stability. Proc Natl Acad Sci USA 108:12699–12704
Hong J, Capp MW, Saecker RM, Record MT Jr (2005) Use of urea and glycine betaine to quantify coupled folding and probe the burial of DNA phosphates in lac repressor-lac operator binding. Biochemistry 44:16896–16911
Koumoto K, Ochiai H, Sugimoto N (2008) Structural effect of synthetic zwitterionic cosolutes on the stability of DNA duplexes. Tetrahedron 64:168–174
Uversky VN, Li J, Fink AL (2001) Trimethylamine-N-oxide-induced folding of alpha-synuclein. FEBS Lett 509:31–35
Miklos AC, Li C, Sharaf NG, Pielak GJ (2010) Volume exclusion and soft interaction effects on protein stability under crowded conditions. Biochemistry 49:6984–6991
Spink CH, Chaires JB (1995) Selective stabilization of triplex DNA by poly(ethylene glycols). J Am Chem Soc 117:12887–12888
Goobes R, Minsky A (2001) Thermodynamic aspects of triplex DNA formation in crowded environments. J Am Chem Soc 123:12692–12693
Goobes R, Kahana N, Cohen O, Minsky A (2003) Metabolic buffering exerted by macromolecular crowding on DNA-DNA interactions: origin and physiological significance. Biochemistry 42:2431–2440
Kitts PA, Nash HA (1987) Homology-dependent interactions in phage lambda site-specific recombination. Nature 329:346–348
Lu M, Guo Q, Marky LA, Seeman NC, Kallenbach NR (1992) Thermodynamics of DNA branching. J Mol Biol 223:781–789
Muhuri S, Mimura K, Miyoshi D, Sugimoto N (2009) Stabilization of three-way junctions of DNA under molecular crowding conditions. J Am Chem Soc 131:9268–9280
Shiman R, Draper DE (2000) Stabilization of RNA tertiary structure by monovalent cations. J Mol Biol 302:79–91
Semrad K, Green R (2002) Osmolytes stimulate the reconstitution of functional 50S ribosomes from in vitro transcripts of Escherichia coli 23S rRNA. RNA 8:401–411
Gluick TC, Yadav S (2003) Trimethylamine N-oxide stabilizes RNA tertiary structure and attenuates the denaturating effects of urea. J Am Chem Soc 125:4418–4419
Lambert D, Leipply D, Draper DE (2010) The osmolyte TMAO stabilizes native RNA tertiary structures in the absence of Mg2+: evidence for a large barrier to folding from phosphate dehydration. J Mol Biol 404:138–157
Pincus DL, Hyeon C, Thirumalai D (2008) Effects of trimethylamine N-oxide (TMAO) and crowding agents on the stability of RNA hairpins. J Am Chem Soc 130:7364–7372
Denesyuk NA, Thirumalai D (2011) Crowding promotes the switch from hairpin to pseudoknot conformation in human telomerase RNA. J Am Chem Soc 133:11858–11861
Tobe S, Heams T, Vergne J, Herve G, Maurel MC (2005) The catalytic mechanism of hairpin ribozyme studied by hydrostatic pressure. Nucleic Acids Res 33:2557–2564
Herve G, Tobe S, Heams T, Vergne J, Maurel MC (2006) Hydrostatic and osmotic pressure study of the hairpin ribozyme. Biochim Biophys Acta 1764:573–577
Downey CD, Crisman RL, Randolph TW, Pardi A (2007) Influence of hydrostatic pressure and cosolutes on RNA tertiary structure. J Am Chem Soc 129:9290–9291
Kilburn D, Roh JH, Guo L, Briber RM, Woodson SA (2010) Molecular crowding stabilizes folded RNA structure by the excluded volume effect. J Am Chem Soc 132:8690–8696
Nakano S, Karimata HT, Kitagawa Y, Sugimoto N (2009) Facilitation of RNA enzyme activity in the molecular crowding media of cosolutes. J Am Chem Soc 131:16881–16888
Keniry MA (2000) Quadruplex structures in nucleic acids. Biopolymers 56:123–146
Simonsson T (2001) G-quadruplex DNA structures – variations on a theme. Biol Chem 382:621–628
Qin Y, Hurley LH (2008) Structures, folding patterns, and functions of intramolecular DNA G-quadruplexes found in eukaryotic promoter regions. Biochimie 90:1149–1171
Dai J, Carver M, Yang D (2008) Polymorphism of human telomeric quadruplex structures. Biochimie 90:1172–1183
Lane AN, Chaires JB, Gray RD, Trent JO (2008) Stability and kinetics of G-quadruplex structures. Nucleic Acids Res 36:5482–5515
Chen F-M (1992) Sr2+ facilitates intermolecular G-quadruplex formation of telomeric sequences. Biochemistry 31:3769–3776
Wang Y, Patel DJ (1992) Guanine residues in d(T2AG3) and d(T2G4) form parallel-stranded potassium cation stabilized G-quadruplexes with anti glycosidic torsion angles in solution. Biochemistry 31:8112–8119
Schultze P, Hud NV, Smith FW, Feigon J (1999) The effect of sodium, potassium and ammonium ions on the conformation of the dimeric quadruplex formed by the Oxytricha nova telomere repeat oligonucleotide d(G4T4G4). Nucleic Acids Res 27:3018–3028
Kankia BI, Marky LA (2001) Folding of the thrombin aptamer into a G-quadruplex with Sr2+: stability, heat, and hydration. J Am Chem Soc 123:10799–10804
Miyoshi D, Nakao A, Sugimoto N (2003) Structural transition from antiparallel to parallel G-quadruplex of d(G4T4G4) induced by Ca2+. Nucleic Acids Res 31:1156–1163
Wu G, Wong A, Gan Z, Davis JT (2003) Direct detection of potassium cations bound to G-quadruplex structures by solid-state 39K NMR at 19.6 T. J Am Chem Soc 125:7182–7183
Sket P, Crnugelj M, Plavec J (2004) d(G3T4G4) forms unusual dimeric G-quadruplex structure with the same general fold in the presence of K+, Na+ or NH +4 ions. Bioorg Med Chem 12:5735–5744
Ida R, Wu G (2005) Solid-state 87Rb NMR signatures for rubidium cations bound to a G-quadruplex. Chem Commun 4294–4296
Gill ML, Strobel SA, Loria JP (2005) 205TI NMR methods for the characterization of monovalent cation binding to nucleic acids. J Am Chem Soc 127:16723–16732
Gray RD, Chaires JB (2008) Kinetics and mechanism of K+ and Na+-induced folding of models of human telomeric DNA into G-quadruplex structures. Nucleic Acids Res 36:4191–4203
Wang Y, Patel DJ (1993) Solution structure of the human telomeric repeat d[AG3(T2AG3)3] G-tetraplex. Structure 1:263–282
Parkinson GN, Lee MP, Neidle S (2002) Crystal structure of parallel quadruplexes from human telomeric DNA. Nature 417:876–880
Ambrus A, Chen D, Dai J, Bialis T, Jones RA, Yang D (2006) Human telomeric sequence forms a hybrid-type intramolecular G-quadruplex structure with mixed parallel/antiparallel strands in potassium solution. Nucleic Acids Res 34:2723–2735
Luu KN, Phan AT, Kuryavyi V, Lacroix L, Patel DJ (2006) Structure of the human telomere in K+ solution: an intramolecular (3+1) G-quadruplex scaffold. J Am Chem Soc 128:9963–9970
Xu Y, Noguchi Y, Sugiyama H (2006) The new models of the human telomere d[AGGG(TTAGGG)3] in K+ solution. Bioorg Med Chem 14:5584–5591
Miyoshi D, Nakao A, Sugimoto N (2002) Molecular crowding regulates the structural switch of the DNA G-quadruplex. Biochemistry 41:15017–15024
Li J, Correia JJ, Wang L, Trent JO, Chaires JB (2005) Not so crystal clear: the structure of the human telomere G-quadruplex in solution differs from that present in a crystal. Nucleic Acids Res 33:4649–4659
Miyoshi D, Karimata H, Sugimoto N (2005) Drastic effect of a single base difference between human and Tetrahymena telomere sequences on their structures under molecular crowding conditions. Angew Chem Int Ed 44:3740–3744
Xue Y, Kan ZY, Wang Q, Yao Y, Liu J, Hao YH, Tan Z (2007) Human telomeric DNA forms parallel-stranded intramolecular G-quadruplex in K+ solution under molecular crowding condition. J Am Chem Soc 129:11185–11191
Zhou J, Wei C, Jia G, Wang X, Tang Q, Feng Z, Li C (2008) The structural transition and compaction of human telomeric G-quadruplex induced by excluded volume effect under cation-deficient conditions. Biophys Chem 136:124–127
Renciuk D, Kejnovská I, Skoláková P, Bednárová K, Motlová J, Vorlícková M (2009) Arrangements of human telomere DNA quadruplex in physiologically relevant K+ solutions. Nucleic Acids Res 37:6625–6634
Xu L, Feng S, Zhou X (2011) Human telomeric G-quadruplexes undergo dynamic conversion in a molecular crowding environment. Chem Commun 47:3517–3519
Heddi B, Phan AT (2011) Structure of human telomeric DNA in crowded solution. J Am Chem Soc 133:9824–9833
Hänsel R, Löhr F, Foldynová-Trantírková S, Bamberg E, Trantírek L, Dötsch V (2011) The parallel G-quadruplex structure of vertebrate telomeric repeat sequences is not the preferred folding topology under physiological conditions. Nucleic Acids Res 39:5768–5775
Miyoshi D, Karimata H, Sugimoto N (2006) Hydration regulates thermodynamics of G-quadruplex formation under molecular crowding conditions. J Am Chem Soc 128:7957–7963
Miller MC, Buscaglia R, Chaires JB, Lane AN, Trent JO (2010) Hydration is a major determinant of the G-quadruplex stability and conformation of the human telomere 3′ Sequence of d(AG3(TTAG3)3). J Am Chem Soc 132:17105–17107
Phan AT, Mergny JL (2002) Human telomeric DNA: G-quadruplex, i-motif and Watson–Crick double helix. Nucleic Acids Res 30:4618–4625
Miyoshi D, Matsumura S, Nakano S, Sugimoto N (2004) Duplex dissociation of telomere DNAs induced by molecular crowding. J Am Chem Soc 126:165–169
Kan Z-Y, Yao Y, Wang P, Li X-H, Hao Y-H, Tan Z (2006) Molecular crowding induces telomere G-quadruplex formation under salt-deficient conditions and enhances its competition with duplex formation. Angew Chem Int Ed 45:1629–1632
Kan ZY, Lin Y, Wang F, Zhuang XY, Zhao Y, Pang DW, Hao YH, Tan Z (2007) G-quadruplex formation in human telomeric (TTAGGG)4 sequence with complementary strand in close vicinity under molecularly crowded condition. Nucleic Acids Res 35:3646–3653
Zhou J, Wei C, Jia G, Wang X, Feng Z, Li C (2009) Human telomeric G-quadruplex formed from duplex under near physiological conditions: spectroscopic evidence and kinetics. Biochimie 91:1104–1111
Kumar N, Maiti S (2005) The effect of osmolytes and small molecule on Quadruplex-WC duplex equilibrium: a fluorescence resonance energy transfer study. Nucleic Acids Res 33:6723–6732
Zheng KW, Chen Z, Hao YH, Tan Z (2010) Molecular crowding creates an essential environment for the formation of stable G-quadruplexes in long double-stranded DNA. Nucleic Acids Res 38:327–338
Li W, Miyoshi D, Nakano S, Sugimoto N (2003) Structural competition involving G-quadruplex DNA and its complement. Biochemistry 42:11736–11744
Miyoshi D, Matsumura S, Li W, Sugimoto N (2003) Structural polymorphism of telomeric DNA regulated by pH and divalent cation. Nucleosides Nucleotides Nucleic Acids 22:203–221
Kumar N, Maiti S (2004) Quadruplex to Watson–Crick duplex transition of the thrombin binding aptamer: a fluorescence resonance energy transfer study. Biochem Biophys Res Commun 319:759–767
Miyoshi D, Inoue M, Sugimoto N (2006) DNA logic gates based on structural polymorphism of telomere DNA molecules responding to chemical input signals. Angew Chem Int Ed 45:7716–7719
Arora A, Maiti S (2009) Stability and molecular recognition of quadruplexes with different loop length in the absence and presence of molecular crowding agents. J Phys Chem B 113:8784–8792
Zhang DH, Fujimoto T, Saxena S, Yu HQ, Miyoshi D, Sugimoto N (2010) Monomorphic RNA G-quadruplex and polymorphic DNA G-quadruplex structures responding to cellular environmental factors. Biochemistry 49:4554–4563
Miyoshi D, Nakamura K, Tateishi-Karimata H, Ohmichi T, Sugimoto N (2009) Hydration of Watson–Crick base pairs and dehydration of Hoogsteen base pairs inducing structural polymorphism under molecular crowding conditions. J Am Chem Soc 131:3522–3531
Pramanik S, Nakamura K, Usui K, Nakano S, Saxena S, Matsui J, Miyoshi D, Sugimoto N (2011) Thermodynamic stability of Hoogsteen and Watson–Crick base pairs in the presence of histone H3-mimicking peptide. Chem Commun 47:2790–2792
Bloomfield VA, Crothers DM, Tinoco I Jr (2000) Nucleic acids structures, properties, and functions. University Science Books, Sausalito, California
Foley PL, Wilson DB, Shuler ML (2010) Macromolecular crowding can account for RNase-sensitive constraint of bacterial nucleoid structure. Biochem Biophys Res Commun 395:42–47
Olsen CM, Gmeiner WH, Marky LA (2006) Unfolding of G-quadruplexes: energetic, and ion and water contributions of G-quartet stacking. J Phys Chem B 110:6962–6969
Olsen CM, Lee HT, Marky LA (2009) Unfolding thermodynamics of intramolecular G-quadruplexes: base sequence contributions of the loops. J Phys Chem B 113:2587–2595
Arora A, Maiti S (2009) Differential biophysical behavior of human telomeric RNA and DNA quadruplex. J Phys Chem B 113:10515–10520
Zimmerman SB, Trach SO (1988) Macromolecular crowding extends the range of conditions under which DNA polymerase is functional. Biochim Biophys Acta 949:297–304
Jarvis TC, Ring DM, Daube SS, von Hippel PH (1990) “Macromolecular crowding”: thermodynamic consequences for protein–protein interactions within the T4 DNA replication complex. J Biol Chem 265:15160–15167
Sasaki Y, Miyoshi D, Sugimoto N (2006) Effect of molecular crowding on DNA polymerase activity. Biotechnol J 1:440–446
Sasaki Y, Miyoshi D, Sugimoto N (2007) Regulation of DNA nucleases by molecular crowding. Nucleic Acids Res 35:4086–4093
Yu HQ, Zhang DH, Gu XB, Miyoshi D, Sugimoto N (2008) Regulation of telomerase activity by the thermodynamic stability of a DNA x RNA hybrid. Angew Chem Int Ed 47:9034–9038
Chen Z, Zheng KW, Hao YH, Tan Z (2009) Reduced or diminished stabilization of the telomere G-quadruplex and inhibition of telomerase by small chemical ligands under molecular crowding condition. J Am Chem Soc 131:10430–10438
Martino L, Pagano B, Fotticchia I, Neidle S, Giancola C (2009) Shedding light on the interaction between TMPyP4 and human telomeric quadruplexes. J Phys Chem B 113:14779–14786
Wei C, Wang J, Zhang M (2010) Spectroscopic study on the binding of porphyrins to (G4T4G4)4 parallel G-quadruplex. Biophys Chem 148:51–55
Wei C, Jia G, Zhou J, Han G, Li C (2009) Evidence for the binding mode of porphyrins to G-quadruplex DNA. Phys Chem Chem Phys 11:4025–4032
Li W, Zhang M, Zhang JL, Li HQ, Zhang XC, Sun Q, Qiu CM (2006) Interactions of daidzin with intramolecular G-quadruplex. FEBS Lett 580:4905–4910
Petraccone L, Fotticchia I, Cummaro A, Pagano B, Ginnari-Satriani L, Haider S, Randazzo A, Novellino E, Neidle S, Giancola C (2011) The triazatruxene derivative azatrux binds to the parallel form of the human telomeric G-quadruplex under molecular crowding conditions: biophysical and molecular modeling studies. Biochimie 93:1318–1327
Dominak LM, Omiatek DM, Gundermann EL, Heien ML, Keating CD (2010) Polymeric crowding agents improve passive biomacromolecule encapsulation in lipid vesicles. Langmuir 26:13195–13200
Zhao C, Ren J, Qu X (2008) Single-walled carbon nanotubes binding to human telomeric i-motif DNA under molecular-crowding conditions: more water molecules released. Chemistry 14:5435–5439
Khripin CY, Arnold-Medabalimi N, Zheng M (2011) Molecular-crowding-induced clustering of DNA-wrapped carbon nanotubes for facile length fractionation. ACS Nano 5:8258–8266
Goodrich GP, Helfrich MR, Overberg JJ, Keating CD (2004) Effect of macromolecular crowding on DNA:Au nanoparticle bioconjugate assembly. Langmuir 20:10246–10251
Zaki A, Dave N, Liu J (2012) Amplifying the macromolecular crowding effect using nanoparticles. J Am Chem Soc 134:35–38
Ricci F, Lai RY, Heeger AJ, Plaxco KW, Sumner JJ (2007) Effect of molecular crowding on the response of an electrochemical DNA sensor. Langmuir 23:6827–6834
Feng B, Frykholm K, Nordén B, Westerlund F (2010) DNA strand exchange catalyzed by molecular crowding in PEG solutions. Chem Commun 46:8231–8233
Zhang C, Shao PG, van Kan JA, van der Maarel JR (2009) Macromolecular crowding induced elongation and compaction of single DNA molecules confined in a nanochannel. Proc Natl Acad Sci USA 106:16651–16666
Acknowledgments
This work was supported in part by Grants-in-Aid for Scientific Research, Scientific Research on Innovative Areas “Nanomedicine Molecular Science” (No. 2306), the “Strategic Research Foundation at Private Universities” (2009–2014), and the “Academic Frontier” project (2004–2009) from the Ministry of Education, Culture, Sports, Science and Technology, Japan, and the Hirao Taro Foundation of the Konan University Association for Academic Research. TF is a research fellow of Japan Society for the promotion of science.
Author information
Authors and Affiliations
Corresponding authors
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2012 Springer-Verlag Berlin Heidelberg
About this chapter
Cite this chapter
Miyoshi, D., Fujimoto, T., Sugimoto, N. (2012). Molecular Crowding and Hydration Regulating of G-Quadruplex Formation. In: Chaires, J., Graves, D. (eds) Quadruplex Nucleic Acids. Topics in Current Chemistry, vol 330. Springer, Berlin, Heidelberg. https://doi.org/10.1007/128_2012_335
Download citation
DOI: https://doi.org/10.1007/128_2012_335
Published:
Publisher Name: Springer, Berlin, Heidelberg
Print ISBN: 978-3-642-34742-9
Online ISBN: 978-3-642-34743-6
eBook Packages: Chemistry and Materials ScienceChemistry and Material Science (R0)