Abstract
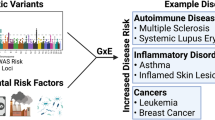
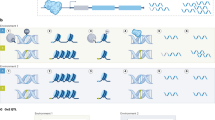
Gene–environment interactions (G × E), the interplay of genetic variation with environmental factors, have a pivotal impact on human complex traits and diseases. Statistically, G × E can be assessed by determining the deviation from expectation of predictive models based solely on the phenotypic effects of genetics or environmental exposures. Despite the unprecedented, widespread and diverse use of G × E analytical frameworks, heterogeneity in their application and reporting hinders their applicability in public health. In this Review, we discuss study design considerations as well as G × E analytical frameworks to assess polygenic liability dependent on the environment, to identify specific genetic variants exhibiting G × E, and to characterize environmental context for these dynamics. We conclude with recommendations to address the most common challenges and pitfalls in the conceptualization, methodology and reporting of G × E studies, as well as future directions.




Similar content being viewed by others
References
Brandes, N., Weissbrod, O. & Linial, M. Open problems in human trait genetics. Genome Biol. 23, 131 (2022). This review comprehensively discusses the major issues and challenges moving forward in human genetics.
Vicente, C. T., Revez, J. A. & Ferreira, M. A. R. Lessons from ten years of genome-wide association studies of asthma. Clin. Transl. Immunol. 6, e165 (2017).
Tsuo, K. et al. Multi-ancestry meta-analysis of asthma identifies novel associations and highlights the value of increased power and diversity. Cell Genomics 2, 100212 (2022).
Trivedi, M. & Denton, E. Asthma in children and adults — what are the differences and what can they tell us about asthma? Front. Pediatr. 7, 256 (2019).
Sandoval-Motta, S., Aldana, M., Martínez-Romero, E. & Frank, A. The human microbiome and the missing heritability problem. Front. Genet. 8, 80 (2017).
Manning, A. K. et al. A genome-wide approach accounting for body mass index identifies genetic variants influencing fasting glycemic traits and insulin resistance. Nat. Genet. 44, 659–669 (2012).
Pingault, J.-B. et al. Using genetic data to strengthen causal inference in observational research. Nat. Rev. Genet. 19, 566–580 (2018).
Ritz, B. R. et al. Lessons learned from past gene–environment interaction successes. Am. J. Epidemiol. 186, 778–786 (2017).
McAllister, K. et al. Current challenges and new opportunities for gene–environment interaction studies of complex diseases. Am. J. Epidemiol. 186, 753–761 (2017).
Thomas, D. Gene–environment-wide association studies: emerging approaches. Nat. Rev. Genet. 11, 259–272 (2010).
Virolainen, S. J., VonHandorf, A., Viel, K. C. M. F., Weirauch, M. T. & Kottyan, L. C. Gene–environment interactions and their impact on human health. Genes. Immun. 24, 1–11 (2023).
Wu, H., Eckhardt, C. M. & Baccarelli, A. A. Molecular mechanisms of environmental exposures and human disease. Nat. Rev. Genet. 24, 332–344 (2023). This review highlights the impact of environmental factors on molecular mechanisms and the challenges in exposome approaches.
Breton, C. V. et al. Exploring the evidence for epigenetic regulation of environmental influences on child health across generations. Commun. Biol. 4, 769 (2021).
Marderstein, A. R. et al. Leveraging phenotypic variability to identify genetic interactions in human phenotypes. Am. J. Hum. Genet. 108, 49–67 (2021).
Westerman, K. E. et al. Variance-quantitative trait loci enable systematic discovery of gene–environment interactions for cardiometabolic serum biomarkers. Nat. Commun. 13, 3993 (2022).
Shi, G. Genome-wide variance quantitative trait locus analysis suggests small interaction effects in blood pressure traits. Sci. Rep. 12, 12649 (2022).
Winkler, T. W. et al. Quality control and conduct of genome-wide association meta-analyses. Nat. Protoc. 9, 1192–1212 (2014).
Ives, C. et al. Linking complex disease and exposure data — insights from an environmental and occupational health study. J. Expo. Sci. Environ. Epidemiol. 33, 12–16 (2023).
Knapp, E. A. et al. The Environmental Influences on Child Health Outcomes (ECHO)-wide cohort. Am. J. Epidemiol. https://doi.org/10.1093/aje/kwad071 (2023).
Hamilton, C. M. et al. The PhenX Toolkit: get the most from your measures. Am. J. Epidemiol. 174, 253–260 (2011).
White, I. R., Royston, P. & Wood, A. M. Multiple imputation using chained equations: issues and guidance for practice. Stat. Med. 30, 377–399 (2011).
Austin, P. C., White, I. R., Lee, D. S. & van Buuren, S. Missing data in clinical research: a tutorial on multiple imputation. Can. J. Cardiol. 37, 1322–1331 (2021).
Hormozdiari, F. et al. Imputing phenotypes for genome-wide association studies. Am. J. Hum. Genet. 99, 89–103 (2016).
Dahl, A. et al. A multiple-phenotype imputation method for genetic studies. Nat. Genet. 48, 466–472 (2016).
Xu, H. et al. Lifestyle risk score: handling missingness of individual lifestyle components in meta-analysis of gene-by-lifestyle interactions. Eur. J. Hum. Genet. 29, 839–850 (2021).
Moore, C. M., Jacobson, S. A. & Fingerlin, T. E. Power and sample size calculations for genetic association studies in the presence of genetic model misspecification. Hum. Hered. 84, 256–271 (2019).
Kooperberg, C. & Hsu, L. powerGWASinteraction: power calculations for G × E and G × G interactions for GWAS. R package version 1.1.3 https://CRAN.R-project.org/package=powerGWASinteraction (2015).
Gauderman, W. J. Sample size requirements for matched case–control studies of gene–environment interaction. Stat. Med. 21, 35–50 (2002).
Gauderman, W. J. Candidate gene association analysis for a quantitative trait, using parent–offspring trios. Genet. Epidemiol. 25, 327–338 (2003).
Gjerdevik, M. et al. Haplin power analysis: a software module for power and sample size calculations in genetic association analyses of family triads and unrelated controls. BMC Bioinforma. 20, 165 (2019).
Gaye, A., Burton, T. W. Y. & Burton, P. R. ESPRESSO: taking into account assessment errors on outcome and exposures in power analysis for association studies. Bioinformatics 31, 2691–2696 (2015).
McCaw, Z. R., Lane, J. M., Saxena, R., Redline, S. & Lin, X. Operating characteristics of the rank-based inverse normal transformation for quantitative trait analysis in genome-wide association studies. Biometrics 76, 1262–1272 (2020).
Ueki, M., Fujii, M. & Tamiya, G.for Alzheimer’s Disease Neuroimaging Initiative and the Alzheimer’s Disease Metabolomics Consortium Quick assessment for systematic test statistic inflation/deflation due to null model misspecifications in genome-wide environment interaction studies. PLoS ONE 14, e0219825 (2019).
Shi, G. & Nehorai, A. Robustness of meta-analyses in finding gene × environment interactions. PLoS ONE 12, e0171446 (2017).
Rao, D. C. et al. Multiancestry study of gene–lifestyle interactions for cardiovascular traits in 610 475 individuals from 124 cohorts: design and rationale. Circ. Cardiovasc. Genet. 10, e001649 (2017).
Young, A. I. et al. Relatedness disequilibrium regression estimates heritability without environmental bias. Nat. Genet. 50, 1304–1310 (2018).
Bernabeu, E. et al. Sex differences in genetic architecture in the UK Biobank. Nat. Genet. 53, 1283–1289 (2021).
Wang, H. et al. Genotype-by-environment interactions inferred from genetic effects on phenotypic variability in the UK Biobank. Sci. Adv. 5, eaaw3538 (2019).
Laville, V. et al. Gene–lifestyle interactions in the genomics of human complex traits. Eur. J. Hum. Genet. 30, 730–739 (2022).
Kerin, M. & Marchini, J. Inferring gene-by-environment interactions with a Bayesian whole-genome regression model. Am. J. Hum. Genet. 107, 698–713 (2020).
Kerin, M. & Marchini, J. A non-linear regression method for estimation of gene–environment heritability. Bioinformatics 36, 5632–5639 (2021).
Dahl, A. et al. A robust method uncovers significant context-specific heritability in diverse complex traits. Am. J. Hum. Genet. 106, 71–91 (2020).
Sulc, J. et al. Quantification of the overall contribution of gene–environment interaction for obesity-related traits. Nat. Commun. 11, 1385 (2020).
Moore, R. et al. A linear mixed-model approach to study multivariate gene–environment interactions. Nat. Genet. 51, 180–186 (2019).
Robinson, M. R. et al. Genotype–covariate interaction effects and the heritability of adult body mass index. Nat. Genet. 49, 1174–1181 (2017).
Ni, G. et al. Genotype–covariate correlation and interaction disentangled by a whole-genome multivariate reaction norm model. Nat. Commun. 10, 2239 (2019).
Bulik-Sullivan, B. K. et al. LD score regression distinguishes confounding from polygenicity in genome-wide association studies. Nat. Genet. 47, 291–295 (2015).
Shin, J. & Lee, S. H. GxEsum: a novel approach to estimate the phenotypic variance explained by genome-wide G × E interaction based on GWAS summary statistics for biobank-scale data. Genome Biol. 22, 183 (2021).
Murcray, C. E., Lewinger, J. P. & Gauderman, W. J. Gene–environment interaction in genome-wide association studies. Am. J. Epidemiol. 169, 219–226 (2008).
Dudbridge, F. & Fletcher, O. Gene–environment dependence creates spurious gene–environment interaction. Am. J. Hum. Genet. 95, 301–307 (2014).
Miao, J. et al. Reimagining gene–environment interaction analysis for human complex traits. Preprint at bioRxiv https://doi.org/10.1101/2022.12.11.519973 (2022).
Warrier, V. et al. Gene–environment correlations and causal effects of childhood maltreatment on physical and mental health: a genetically informed approach. Lancet Psychiatry 8, 373–386 (2021).
Wang, Z., Shi, W., Carroll, R. J. & Chatterjee, N. Joint modeling of gene–environment correlations and interactions using polygenic risk scores in case–control studies. Preprint at bioRxiv https://doi.org/10.1101/2023.02.14.528572 (2023).
Spiller, W., Hartwig, F. P., Sanderson, E., Davey Smith, G. & Bowden, J. Interaction-based Mendelian randomization with measured and unmeasured gene-by-covariate interactions. PLoS ONE 17, e0271933 (2022).
Kim, Y., Balbona, J. V. & Keller, M. C. Bias and precision of parameter estimates from models using polygenic scores to estimate environmental and genetic parental influences. Behav. Genet. 51, 279–288 (2021).
Tchetgen Tchetgen, E., Sun, B. & Walter, S. The GENIUS approach to robust Mendelian randomization inference. Stat. Sci. 36, 443–464 (2021).
Liu, Z., Ye, T., Sun, B., Schooling, M. & Tchetgen, E. T. Mendelian randomization mixed-scale treatment effect robust identification and estimation for causal inference. Biometrics 79, 2208–2219 (2022).
Balbona, J. V., Kim, Y. & Keller, M. C. Estimation of parental effects using polygenic scores. Behav. Genet. 51, 264–278 (2021). This article describes a model to determine parental effects on child traits using polygenic risk scores while accounting for assortative mating.
Spiller, W., Slichter, D., Bowden, J. & Davey Smith, G. Detecting and correcting for bias in Mendelian randomization analyses using gene-by-environment interactions. Int. J. Epidemiol. 48, 702–712 (2019).
Karageorgiou, V., Tyrrell, J., Mckinley, T. J. & Bowden, J. Weak and pleiotropy robust sex-stratified Mendelian randomization in the one sample and two sample settings. Genet. Epidemiol. 47, 135–151 (2023).
Meisner, A., Kundu, P. & Chatterjee, N. Case-only analysis of gene–environment interactions using polygenic risk scores. Am. J. Epidemiol. 188, 2013–2020 (2019).
Tang, Y., You, D., Yi, H., Yang, S. & Zhao, Y. IPRS: leveraging gene–environment interaction to reconstruct polygenic risk score. Front. Genet. 13, 801397 (2022).
Arnau-Soler, A. et al. Genome-wide by environment interaction studies of depressive symptoms and psychosocial stress in UK Biobank and Generation Scotland. Transl. Psychiatry 9, 14 (2019).
Werme, J., van der Sluis, S., Posthuma, D. & de Leeuw, C. A. Genome-wide gene–environment interactions in neuroticism: an exploratory study across 25 environments. Transl. Psychiatry 11, 180 (2021).
Lau, M., Kress, S., Schikowski, T. & Schwender, H. Efficient gene–environment interaction testing through bootstrap aggregating. Sci. Rep. 13, 937 (2023).
Kraft, P., Yen, Y.-C., Stram, D. O., Morrison, J. & Gauderman, W. J. Exploiting gene–environment interaction to detect genetic associations. Hum. Hered. 63, 111–119 (2007).
Willer, C. J., Li, Y. & Abecasis, G. R. METAL: fast and efficient meta-analysis of genomewide association scans. Bioinformatics 26, 2190–2191 (2010).
Manning, A. K. et al. Meta-analysis of gene–environment interaction: joint estimation of SNP and SNP × environment regression coefficients. Genet. Epidemiol. 35, 11–18 (2011).
Aschard, H., Hancock, D. B., London, S. J. & Kraft, P. Genome-wide meta-analysis of joint tests for genetic and gene–environment interaction effects. Hum. Hered. 70, 292–300 (2010).
Randall, J. C. et al. Sex-stratified genome-wide association studies including 270,000 individuals show sexual dimorphism in genetic loci for anthropometric traits. PLoS Genet 9, e1003500 (2013).
Laville, V. et al. Deriving stratified effects from joint models investigating gene–environment interactions. BMC Bioinforma. 21, 251 (2020).
Sung, Y. J. et al. An empirical comparison of joint and stratified frameworks for studying G × E interactions: systolic blood pressure and smoking in the CHARGE Gene–Lifestyle Interactions Working Group. Genet. Epidemiol. 40, 404–415 (2016).
Winkler, T. W. et al. The influence of age and sex on genetic associations with adult body size and shape: a large-scale genome-wide interaction study. PLoS Genet. 11, e1005378 (2015).
Jin, Q. & Shi, G. Meta-analysis of joint test of SNP and SNP–environment interaction with heterogeneity. Hum. Hered. 86, 1–9 (2021).
Jin, Q. & Shi, G. Meta-analysis of SNP–environment interaction with heterogeneity. Hum. Hered. 84, 117–126 (2019).
Yu, Y. et al. Subset-based analysis using gene–environment interactions for discovery of genetic associations across multiple studies or phenotypes. Hum. Hered. 83, 283–314 (2018).
Wu, S., Xu, Y., Zhang, Q. & Ma, S. Gene–environment interaction analysis via deep learning. Genet. Epidemiol. 47, 261–286 (2023).
Gauderman, W. J. et al. Update on the state of the science for analytical methods for gene–environment interactions. Am. J. Epidemiol. 186, 762–770 (2017).
Mukherjee, B. & Chatterjee, N. Exploiting gene–environment independence for analysis of case–control studies: an empirical Bayes-type shrinkage estimator to trade-off between bias and efficiency. Biometrics 64, 685–694 (2008).
Lin, W.-Y., Huang, C.-C., Liu, Y.-L., Tsai, S.-J. & Kuo, P.-H. Polygenic approaches to detect gene–environment interactions when external information is unavailable. Brief. Bioinform. 20, 2236–2252 (2019).
Gauderman, W. J., Zhang, P., Morrison, J. L. & Lewinger, J. P. Finding novel genes by testing G × E interactions in a genome-wide association study. Genet. Epidemiol. 37, 603–613 (2013).
Hsu, L. et al. Powerful cocktail methods for detecting genome-wide gene–environment interaction. Genet. Epidemiol. 36, 183–194 (2012).
Majumdar, A. et al. A two-step approach to testing overall effect of gene–environment interaction for multiple phenotypes. Bioinformatics 36, 5640–5648 (2021).
Davis, A. P. et al. Comparative Toxicogenomics Database (CTD): update 2023. Nucleic Acids Res. 51, D1257–D1262 (2023).
Freshour, S. L. et al. Integration of the Drug–Gene Interaction Database (DGIdb 4.0) with open crowdsource efforts. Nucleic Acids Res. 49, D1144–D1151 (2021).
Miao, J. & Lu, Q. in Handbook of Statistical Bioinformatics (eds. Lu, H. H.-S., Schölkopf, B., Wells, M. T. & Zhao, H.) 257–270 (Springer, 2022).
Lyon, M. S., Millard, L. A. C., Smith, G. D., Gaunt, T. R. & Tilling, K. Hypothesis-free detection of gene-interaction effects on biomarker concentration in UK Biobank using variance prioritisation. Preprint at medRxiv https://doi.org/10.1101/2022.01.05.21268406 (2022).
Kim, J. et al. Joint analysis of multiple interaction parameters in genetic association studies. Genetics 211, 483–494 (2019).
Chi, J. T. et al. SEAGLE: a scalable exact algorithm for large-scale set-based gene–environment interaction tests in Biobank data. Front. Genet. 12, 710055 (2021).
Lin, X., Lee, S., Christiani, D. C. & Lin, X. Test for interactions between a genetic marker set and environment in generalized linear models. Biostatistics 14, 667–681 (2013).
Zhao, G., Marceau, R., Zhang, D. & Tzeng, J.-Y. Assessing gene–environment interactions for common and rare variants with binary traits using gene-trait similarity regression. Genetics 199, 695–710 (2015).
Tzeng, J.-Y. et al. Studying gene and gene–environment effects of uncommon and common variants on continuous traits: a marker-set approach using gene-trait similarity regression. Am. J. Hum. Genet. 89, 277–288 (2011).
Zhao, N., Zhang, H., Clark, J. J., Maity, A. & Wu, M. C. Composite kernel machine regression based on likelihood ratio test for joint testing of genetic and gene–environment interaction effect. Biometrics 75, 625–637 (2019).
Price, A. L. et al. Pooled association tests for rare variants in exon-resequencing studies. Am. J. Hum. Genet. 86, 832–838 (2010).
Wu, M. C. et al. Rare-variant association testing for sequencing data with the sequence kernel association test. Am. J. Hum. Genet. 89, 82–93 (2011).
Li, B. & Leal, S. M. Methods for detecting associations with rare variants for common diseases: application to analysis of sequence data. Am. J. Hum. Genet. 83, 311–321 (2008).
Jiao, S. et al. SBERIA: set-based gene–environment interaction test for rare and common variants in complex diseases. Genet. Epidemiol. 37, 452–464 (2013).
Zhao, Z., Zhang, J., Sha, Q. & Hao, H. Testing gene–environment interactions for rare and/or common variants in sequencing association studies. PLoS ONE 15, e0229217 (2020).
Zemlianskaia, N., Gauderman, W. J. & Lewinger, J. P. A scalable hierarchical lasso for gene–environment interactions. J. Comput. Graph. Stat. 31, 1091–1103 (2022).
Wang, X. et al. Efficient gene–environment interaction tests for large biobank-scale sequencing studies. Genet. Epidemiol. 44, 908–923 (2020).
Su, Y.-R., Di, C.-Z., Hsu, L. & Genetics and Epidemiology of Colorectal Cancer Consortium A unified powerful set-based test for sequencing data analysis of G × E interactions. Biostatistics 18, 119–131 (2017).
Chen, H., Meigs, J. B. & Dupuis, J. Incorporating gene–environment interaction in testing for association with rare genetic variants. Hum. Hered. 78, 81–90 (2014).
Lim, E., Chen, H., Dupuis, J. & Liu, C.-T. A unified method for rare variant analysis of gene–environment interactions. Stat. Med. 39, 801–813 (2020).
Lin, X. et al. Test for rare variants by environment interactions in sequencing association studies. Biometrics 72, 156–164 (2016).
Zhang, J. et al. Test gene–environment interactions for multiple traits in sequencing association studies. Hum. Hered. 84, 170–196 (2019).
He, Z. et al. Set-based tests for the gene–environment interaction in longitudinal studies. J. Am. Stat. Assoc. 112, 966–978 (2017).
Johnsen, P. V., Riemer-Sørensen, S., DeWan, A. T., Cahill, M. E. & Langaas, M. A new method for exploring gene–gene and gene–environment interactions in GWAS with tree ensemble methods and SHAP values. BMC Bioinforma. 22, 230 (2021).
Lee, S., Teslovich, T. M., Boehnke, M. & Lin, X. General framework for meta-analysis of rare variants in sequencing association studies. Am. J. Hum. Genet. 93, 42–53 (2013).
Wang, J. et al. A meta-analysis approach with filtering for identifying gene-level gene–environment interactions. Genet. Epidemiol. 42, 434–446 (2018).
Jin, X. & Shi, G. Variance-component-based meta-analysis of gene–environment interactions for rare variants. G3 Bethesda 11, jkab203 (2021).
Wang, X. et al. Genomic summary statistics and meta-analysis for set-based gene–environment interaction tests in large-scale sequencing studies. Preprint at medRxiv https://doi.org/10.1101/2022.05.08.22274819 (2022).
Bi, W. et al. A fast and accurate method for genome-wide scale phenome-wide G × E analysis and its application to UK Biobank. Am. J. Hum. Genet. 105, 1182–1192 (2019).
Zhong, W., Chhibber, A., Luo, L., Mehrotra, D. V. & Shen, J. A fast and powerful linear mixed model approach for genotype–environment interaction tests in large-scale GWAS. Brief. Bioinform. 24, bbac547 (2023).
Jin, X. & Shi, G. Kernel-based gene–environment interaction tests for rare variants with multiple quantitative phenotypes. PLoS ONE 17, e0275929 (2022).
Hecker, J. et al. A robust and adaptive framework for interaction testing in quantitative traits between multiple genetic loci and exposure variables. PLoS Genet. 18, e1010464 (2022).
Westerman, K. E. et al. GEM: scalable and flexible gene–environment interaction analysis in millions of samples. Bioinformatics 37, 3514–3520 (2021).
Osazuwa-Peters, O. L. et al. Identifying blood pressure loci whose effects are modulated by multiple lifestyle exposures. Genet. Epidemiol. 44, 629–641 (2020).
Zhou, Z., Ku, H.-C., Manning, S. E., Zhang, M. & Xing, C. A varying coefficient model to jointly test genetic and gene–environment interaction effects. Behav. Genet. https://doi.org/10.1007/s10519-022-10131-w (2023).
Mulligan, C. J. et al. Novel G × E effects and resilience: a case:control longitudinal study of psychosocial stress with war-affected youth. PLoS ONE 17, e0266509 (2022).
Zhang, W. et al. Detecting gene–environment interaction for maternal exposures using case–parent trios ascertained through a case with non-syndromic orofacial cleft. Front. Cell Dev. Biol. 9, 621018 (2021).
Dizier, M.-H. et al. Interactive effect between ATPase-related genes and early-life tobacco smoke exposure on bronchial hyper-responsiveness detected in asthma-ascertained families. Thorax 74, 254–260 (2019).
Pingault, J.-B. et al. Genetic nurture versus genetic transmission of risk for ADHD traits in the Norwegian Mother, Father and Child Cohort Study. Mol. Psychiatry https://doi.org/10.1038/s41380-022-01863-6 (2022).
Demange, P. A. et al. Estimating effects of parents’ cognitive and non-cognitive skills on offspring education using polygenic scores. Nat. Commun. 13, 4801 (2022).
Dunn, E. C. et al. Research review: gene–environment interaction research in youth depression — a systematic review with recommendations for future research. J. Child. Psychol. Psychiatry 52, 1223–1238 (2011).
Zhang, H. et al. A new method for multiancestry polygenic prediction improves performance across diverse populations. Nat. Genet. 55, 1757–1768 (2023).
All of Us Research Program Investigators et al. The ‘All of Us’ Research Program. N. Engl. J. Med. 381, 668–676 (2019).
Kachuri, L. et al. Gene expression in African Americans, Puerto Ricans and Mexican Americans reveals ancestry-specific patterns of genetic architecture. Nat. Genet. 55, 952–963 (2023).
Li, B. et al. Incorporating local ancestry improves identification of ancestry-associated methylation signatures and meQTLs in African Americans. Commun. Biol. 5, 401 (2022).
Czamara, D. et al. Integrated analysis of environmental and genetic influences on cord blood DNA methylation in new-borns. Nat. Commun. 10, 2548 (2019).
Majarian, T. D. et al. Multi-omics insights into the biological mechanisms underlying statistical gene-by-lifestyle interactions with smoking and alcohol consumption. Front. Genet. 13, 954713 (2022). This study elucidates how epigenomics, transcriptomics and G × E summary statistics can be combined to provide molecular evidence for G × E statistical interactions.
Findley, A. S. et al. Functional dynamic genetic effects on gene regulation are specific to particular cell types and environmental conditions. eLife 10, e67077 (2021).
Nagar, S. D., Nápoles, A. M., Jordan, I. K. & Mariño-Ramírez, L. Socioeconomic deprivation and genetic ancestry interact to modify type 2 diabetes ethnic disparities in the United Kingdom. EClinicalMedicine 37, 100960 (2021).
Park, D. S. et al. An ancestry-based approach for detecting interactions. Genet. Epidemiol. 42, 49–63 (2018).
Abdellaoui, A., Yengo, L., Verweij, K. J. H. & Visscher, P. M. 15 years of GWAS discovery: realizing the promise. Am. J. Hum. Genet. 110, 179–194 (2023).
Peterson, R. E. et al. Genome-wide association studies in ancestrally diverse populations: opportunities, methods, pitfalls, and recommendations. Cell 179, 589–603 (2019).
Buniello, A. et al. The NHGRI-EBI GWAS Catalog of published genome-wide association studies, targeted arrays and summary statistics 2019. Nucleic Acids Res. 47, D1005–D1012 (2019).
Lambert, S. A. et al. The Polygenic Score Catalog as an open database for reproducibility and systematic evaluation. Nat. Genet. 53, 420–425 (2021).
Wilkinson, M. D. et al. The FAIR guiding principles for scientific data management and stewardship. Sci. Data 3, 160018 (2016).
Reales, G. & Wallace, C. Sharing GWAS summary statistics results in more citations. Commun. Biol. 6, 116 (2023).
Little, J. et al. STrengthening the REporting of Genetic Association Studies (STREGA): an extension of the STROBE statement. PLoS Med. 6, e22 (2009).
Wand, H. et al. Improving reporting standards for polygenic scores in risk prediction studies. Nature 591, 211–219 (2021).
Khan, A. T. et al. Recommendations on the use and reporting of race, ethnicity, and ancestry in genetic research: experiences from the NHLBI TOPMed program. Cell Genomics 2, 100155 (2022).
Committee on the Use of Race, Ethnicity, and Ancestry as Population Descriptors in Genomics Research et al. Using Population Descriptors in Genetics and Genomics Research: A New Framework for an Evolving Field 26902 (National Academies Press, 2023).
Wijsman, E. M. Family-based approaches: design, imputation, analysis, and beyond. BMC Genet. 17, 9 (2016).
Truong, V. Q. et al. Quality control procedures for genome‐wide association studies. Curr. Protoc. 2, e603 (2022).
Hayhurst, J. et al. A community driven GWAS summary statistics standard. Preprint at bioRxiv https://doi.org/10.1101/2022.07.15.500230 (2022).
Sollis, E. et al. The NHGRI-EBI GWAS Catalog: knowledgebase and deposition resource. Nucleic Acids Res. 51, D977–D985 (2023).
Ottman, R. Gene–environment interaction: definitions and study designs. Prev. Med. 25, 764–770 (1996).
Wright, A. F., Carothers, A. D. & Campbell, H. Gene–environment interactions — the BioBank UK study. Pharmacogenomics J. 2, 75–82 (2002).
Olvera Alvarez, H. A., Appleton, A. A., Fuller, C. H., Belcourt, A. & Kubzansky, L. D. An integrated socio-environmental model of health and well-being: a conceptual framework exploring the joint contribution of environmental and social exposures to health and disease over the life span. Curr. Environ. Health Rep. 5, 233–243 (2018).
Acknowledgements
The authors’ work was funded by the Combining advances in Genomics and Environmental science to accelerate Actionable Research and practice in ASD (GEARS) Network (R01ES034554). E.H.-L. and G.L.-W. received support from the National Institutes of Health (NIH) National Human Genome Research Institute (NHGRI) (R35HG011944-02). No funding agencies had a role in the study design, data collection and analysis, decision to publish or preparation of the manuscript. The views expressed are those of the authors and not necessarily those of any funder.
Author information
Authors and Affiliations
Contributions
E.H.-L., C.L.-A. and G.L.W. researched the literature. E.H.-L. and G.L.W. wrote the article. All authors contributed to discussions of the content and reviewed and/or edited the manuscript.
Corresponding author
Peer review
Peer review information
Nature Reviews Genetics thanks Seunggeun Lee, and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Competing interests
The authors declare no competing interests.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Related links
Comparative Toxicogenomics Database: https://ctdbase.org/
Drug–Gene Interaction Database: https://www.dgidb.org/
GWAS Catalog: https://www.ebi.ac.uk/gwas/
PGS Catalog: https://www.pgscatalog.org/
PhenX Toolkit: https://www.phenxtoolkit.org
Supplementary information
Glossary
- Biological or functional interaction
-
The scenario whereby the exposures physically interact to produce an outcome.
- Broad-sense heritability
-
(H2). The proportion of phenotypic variation attributed to genetic variation, including additive, dominance, epistasis and gene–environment interactions (G × E).
- G × E correlation
-
(rGE). A situation when there are genetic differences in exposure to specific environments. rGEcan be classified as passive, active or evocative according to its source.
- G × E heritability
-
(h2GE). The proportion of the phenotypic variability attributable to G × E.
- Genome-wide interaction studies
-
(GWIS). Analytical approaches to investigate the modifying effect of an exposure variable on a genetic association.
- Heteroscedasticity
-
A statistical phenomenon where the variability of the residuals (errors) in a regression model varies across different levels of the predictor variables.
- Imputation
-
The statistical process of predicting missing genetic data based on observed data, typically using reference panels of known genetic variants.
- Instrumental variables
-
Variables associated with an exposure that is not associated with the outcome through any other pathway.
- Mechanistic interactions
-
Situations when there are individuals for whom the outcome would occur if both of two exposures were present but not if one or both of the exposures were absent.
- Mendelian randomization
-
(MR). An analytical method that uses ‘instrumental variables’ for the predictor to determine the causal influences of a predictor on an outcome. MR is based on three key assumptions: instrumental variables are associated with the exposure; instrumental variables are not associated with any confounder; and instrumental variables do not exert effects on the phenotype not mediated by the exposure (also known as ‘horizontal pleiotropy’).
- Multiple testing
-
Repeated statistical testing of hypotheses within a single study, increasing the likelihood of false positives. Adjustments such as Bonferroni correction help control the family-wise error rate.
- Polygenic score
-
(PGS). A single value that aggregates the effect of variants across an individual’s germ-line genome to quantify the genetic predisposition to a trait. Usually calculated as a weighted sum of several trait-associated alleles (dosage), where the per-allele effect sizes (βi) are extracted from the genome-wide association study (GWAS) summary statistics of independent studies.
- Set-based analyses
-
Analytical methods that aggregate G × E signals within a gene or genomic region.
- Statistical interactions
-
Situations when the effect of a risk factor on an outcome varies across strata of another factor. An interaction on an additive scale occurs when the combined effect of the risk factors differs with regards to the sum of the individual effects, whereas an interaction on a multiplicative scales occurs when the combined effect of the risk factors differs with regards to the product of the individual effects.
- Variance quantitative trait locus
-
(vQTL). Genetic variants associated with the variance of a continuous trait.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Herrera-Luis, E., Benke, K., Volk, H. et al. Gene–environment interactions in human health. Nat Rev Genet (2024). https://doi.org/10.1038/s41576-024-00731-z
Accepted:
Published:
DOI: https://doi.org/10.1038/s41576-024-00731-z
- Springer Nature Limited
This article is cited by
-
The interplay between male fertility, mental health and sexual function
Nature Reviews Urology (2024)
-
CD147 TagSNP is associated with the vulnerability to lung cancer in the Chinese population: a case–control study
Discover Oncology (2024)