Abstract
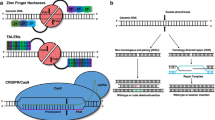
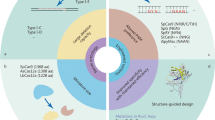
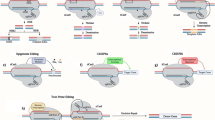
Many novel CRISPR-based genome-editing tools, with a wide variety of applications, have been developed in the past few years. The original CRISPR–Cas9 system was developed as a tool to alter genomic sequences in living organisms in a simple way. However, the functions of new CRISPR tools are not limited to conventional genome editing mediated by non-homologous end-joining or homology-directed repair but expand into gene-expression control, epigenome editing, single-nucleotide editing, RNA editing and live-cell imaging. Furthermore, genetic perturbation screening by multiplexing guide RNAs is gaining popularity as a method to identify causative genes and pathways in an unbiased manner. New CRISPR tools can also be applied to ex vivo or in vivo therapeutic genome editing for the treatment of conditions such as hyperlipidaemia. In this Review, we first provide an overview of the diverse new CRISPR tools that have been developed to date. Second, we summarize how these new CRISPR tools are being used to study biological processes and disease mechanisms in cardiovascular research and medicine. Finally, we discuss the prospect of therapeutic genome editing by CRISPR tools to cure genetic cardiovascular diseases.
Key points
-
New CRISPR-based tools with various functions provide an increasing number of options to study disease mechanisms and cure genetic diseases.
-
CRISPR screens to identify causative genes are becoming more common and can be combined with new CRISPR tools.
-
New CRISPR tools, particularly base editors, have potential for therapeutic genome editing.
-
In cardiovascular medicine, the focus of therapeutic genome editing is on the liver to reduce blood LDL-cholesterol levels.
-
The high efficiency and specificity of new CRISPR tools could enable therapeutic genome editing of inherited cardiac and vascular diseases.
-
Therapeutic genome editing requires further investigation of in vivo off-target effects and improved delivery methods.





Similar content being viewed by others
References
Marraffini, L. A. CRISPR-Cas immunity in prokaryotes. Nature 526, 55–61 (2015).
Hsu, P. D., Lander, E. S. & Zhang, F. Development and applications of CRISPR-Cas9 for genome engineering. Cell 157, 1262–1278 (2014).
Wright, A. V., Nunez, J. K. & Doudna, J. A. Biology and applications of CRISPR systems: harnessing nature’s toolbox for genome engineering. Cell 164, 29–44 (2016).
Dominguez, A. A., Lim, W. A. & Qi, L. S. Beyond editing: repurposing CRISPR-Cas9 for precision genome regulation and interrogation. Nat. Rev. Mol. Cell Biol. 17, 5–15 (2016).
Yeh, C. D., Richardson, C. D. & Corn, J. E. Advances in genome editing through control of DNA repair pathways. Nat. Cell Biol. 21, 1468–1478 (2019).
Cong, L. et al. Multiplex genome engineering using CRISPR/Cas systems. Science 339, 819–823 (2013).
Gasiunas, G., Barrangou, R., Horvath, P. & Siksnys, V. Cas9-crRNA ribonucleoprotein complex mediates specific DNA cleavage for adaptive immunity in bacteria. Proc. Natl Acad. Sci. USA 109, E2579–E2586 (2012).
Jinek, M. et al. A programmable dual-RNA-guided DNA endonuclease in adaptive bacterial immunity. Science 337, 816–821 (2012).
Qi, L. S. et al. Repurposing CRISPR as an RNA-guided platform for sequence-specific control of gene expression. Cell 152, 1173–1183 (2013).
Gilbert, L. A. et al. CRISPR-mediated modular RNA-guided regulation of transcription in eukaryotes. Cell 154, 442–451 (2013).
Xu, X. & Qi, L. S. A CRISPR-dCas toolbox for genetic engineering and synthetic biology. J. Mol. Biol. 431, 34–47 (2019).
Musunuru, K. et al. In vivo CRISPR base editing of PCSK9 durably lowers cholesterol in primates. Nature 593, 429–434 (2021).
Rothgangl, T. et al. In vivo adenine base editing of PCSK9 in macaques reduces LDL cholesterol levels. Nat. Biotechnol. 39, 949–957 (2021).
Branzei, D. & Foiani, M. Regulation of DNA repair throughout the cell cycle. Nat. Rev. Mol. Cell Biol. 9, 297–308 (2008).
Sfeir, A. & Symington, L. S. Microhomology-mediated end joining: a back-up survival mechanism or dedicated pathway? Trends Biochem. Sci. 40, 701–714 (2015).
Ishizu, T. et al. Targeted genome replacement via homology-directed repair in non-dividing cardiomyocytes. Sci. Rep. 7, 9363 (2017).
Kohama, Y. et al. Adeno-associated virus-mediated gene delivery promotes S-phase entry-independent precise targeted integration in cardiomyocytes. Sci. Rep. 10, 15348 (2020).
Zetsche, B. et al. Cpf1 is a single RNA-guided endonuclease of a class 2 CRISPR-Cas system. Cell 163, 759–771 (2015).
Anzalone, A. V., Koblan, L. W. & Liu, D. R. Genome editing with CRISPR-Cas nucleases, base editors, transposases and prime editors. Nat. Biotechnol. 38, 824–844 (2020).
Rees, H. A. & Liu, D. R. Base editing: precision chemistry on the genome and transcriptome of living cells. Nat. Rev. Genet. 19, 770–788 (2018).
Bikard, D. et al. Programmable repression and activation of bacterial gene expression using an engineered CRISPR-Cas system. Nucleic Acids Res. 41, 7429–7437 (2013).
Gilbert, L. A. et al. Genome-scale CRISPR-mediated control of gene repression and activation. Cell 159, 647–661 (2014).
Mandegar, M. A. et al. CRISPR interference efficiently induces specific and reversible gene silencing in human iPSCs. Cell Stem Cell 18, 541–553 (2016).
Limpitikul, W. B. et al. A precision medicine approach to the rescue of function on malignant calmodulinopathic long-QT syndrome. Circ. Res. 120, 39–48 (2017).
Chowdhury, T. A. et al. Temporal and spatial post-transcriptional regulation of zebrafish tie1 mRNA by long noncoding RNA during brain vascular assembly. Arterioscler. Thromb. Vasc. Biol. 38, 1562–1575 (2018).
Krause, M. D. et al. Genetic variant at coronary artery disease and ischemic stroke locus 1p32.2 regulates endothelial responses to hemodynamics. Proc. Natl Acad. Sci. USA 115, E11349–E11358 (2018).
Savage, A. M. et al. tmem33 is essential for VEGF-mediated endothelial calcium oscillations and angiogenesis. Nat. Commun. 10, 732 (2019).
Stolze, L. K. et al. Systems genetics in human endothelial cells identifies non-coding variants modifying enhancers, expression, and complex disease traits. Am. J. Hum. Genet. 106, 748–763 (2020).
Thakore, P. I. et al. RNA-guided transcriptional silencing in vivo with S. aureus CRISPR-Cas9 repressors. Nat. Commun. 9, 1674 (2018).
Maeder, M. L. et al. CRISPR RNA-guided activation of endogenous human genes. Nat. Methods 10, 977–979 (2013).
Perez-Pinera, P. et al. RNA-guided gene activation by CRISPR-Cas9-based transcription factors. Nat. Methods 10, 973–976 (2013).
Tanenbaum, M. E., Gilbert, L. A., Qi, L. S., Weissman, J. S. & Vale, R. D. A protein-tagging system for signal amplification in gene expression and fluorescence imaging. Cell 159, 635–646 (2014).
Chavez, A. et al. Highly efficient Cas9-mediated transcriptional programming. Nat. Methods 12, 326–328 (2015).
Konermann, S. et al. Genome-scale transcriptional activation by an engineered CRISPR-Cas9 complex. Nature 517, 583–588 (2015).
Dal-Pra, S., Hodgkinson, C. P. & Dzau, V. J. Induced cardiomyocyte maturation: cardiac transcription factors are necessary but not sufficient. PLoS ONE 14, e0223842 (2019).
Schoger, E. et al. CRISPR-mediated activation of endogenous gene expression in the postnatal heart. Circ. Res. 126, 6–24 (2020).
Sano, T., Ito, T., Ishigami, S., Bandaru, S. & Sano, S. Intrinsic activation of cardiosphere-derived cells enhances myocardial repair. J. Thorac. Cardiovasc. Surg. https://doi.org/10.1016/j.jtcvs.2020.05.040 (2020).
Suzuki, M. M. & Bird, A. DNA methylation landscapes: provocative insights from epigenomics. Nat. Rev. Genet. 9, 465–476 (2008).
Amabile, A. et al. Inheritable silencing of endogenous genes by hit-and-run targeted epigenetic editing. Cell 167, 219–232.e14 (2016).
Liu, X. S. et al. Editing DNA methylation in the mammalian genome. Cell 167, 233–247 (2016).
Vojta, A. et al. Repurposing the CRISPR-Cas9 system for targeted DNA methylation. Nucleic Acids Res. 44, 5615–5628 (2016).
Morita, S. et al. Targeted DNA demethylation in vivo using dCas9-peptide repeat and scFv-TET1 catalytic domain fusions. Nat. Biotechnol. 34, 1060–1065 (2016).
Xu, X. et al. A CRISPR-based approach for targeted DNA demethylation. Cell Discov. 2, 16009 (2016).
Nuñez, J. K. et al. Genome-wide programmable transcriptional memory by CRISPR-based epigenome editing. Cell 184, 2503–2519 (2021).
Nakamura, M., Ivec, A. E., Gao, Y. & Qi, L. S. Durable CRISPR-based epigenetic silencing. BioDesign Res. 2021, 9815820 (2021).
Bannister, A. J. & Kouzarides, T. Regulation of chromatin by histone modifications. Cell Res. 21, 381–395 (2011).
Kouzarides, T. Chromatin modifications and their function. Cell 128, 693–705 (2007).
Hilton, I. B. et al. Epigenome editing by a CRISPR-Cas9-based acetyltransferase activates genes from promoters and enhancers. Nat. Biotechnol. 33, 510–517 (2015).
Kearns, N. A. et al. Functional annotation of native enhancers with a Cas9-histone demethylase fusion. Nat. Methods 12, 401–403 (2015).
Cano-Rodriguez, D. et al. Writing of H3K4Me3 overcomes epigenetic silencing in a sustained but context-dependent manner. Nat. Commun. 7, 12284 (2016).
Kim, J. M. et al. Cooperation between SMYD3 and PC4 drives a distinct transcriptional program in cancer cells. Nucleic Acids Res. 43, 8868–8883 (2015).
Kwon, D. Y., Zhao, Y. T., Lamonica, J. M. & Zhou, Z. Locus-specific histone deacetylation using a synthetic CRISPR-Cas9-based HDAC. Nat. Commun. 8, 15315 (2017).
Wang, H. et al. CRISPR-mediated programmable 3D genome positioning and nuclear organization. Cell 175, 1405–1417 (2018).
Shin, Y. et al. Liquid nuclear condensates mechanically sense and restructure the genome. Cell 175, 1481–1491 (2018).
Gao, Y., Han, M., Shang, S., Wang, H. & Qi, L. S. Interrogation of the dynamic properties of higher-order heterochromatin using CRISPR-dCas9. Mol. Cell 81, 4287–4299 (2021).
Komor, A. C., Kim, Y. B., Packer, M. S., Zuris, J. A. & Liu, D. R. Programmable editing of a target base in genomic DNA without double-stranded DNA cleavage. Nature 533, 420–424 (2016).
Nishida, K. et al. Targeted nucleotide editing using hybrid prokaryotic and vertebrate adaptive immune systems. Science 353, aaf8729 (2016).
Cuella-Martin, R. et al. Functional interrogation of DNA damage response variants with base editing screens. Cell 184, 1081–1097 (2021).
Hanna, R. E. et al. Massively parallel assessment of human variants with base editor screens. Cell 184, 1064–1080 (2021).
Chadwick, A. C., Wang, X. & Musunuru, K. In vivo base editing of PCSK9 (proprotein convertase subtilisin/kexin type 9) as a therapeutic alternative to genome editing. Arterioscler. Thromb. Vasc. Biol. 37, 1741–1747 (2017).
Kim, K. et al. Highly efficient RNA-guided base editing in mouse embryos. Nat. Biotechnol. 35, 435–437 (2017).
Liu, Z. et al. Highly efficient RNA-guided base editing in rabbit. Nat. Commun. 9, 2717 (2018).
Ryu, S. M. et al. Adenine base editing in mouse embryos and an adult mouse model of Duchenne muscular dystrophy. Nat. Biotechnol. 36, 536–539 (2018).
Xu, L. et al. Efficient precise in vivo base editing in adult dystrophic mice. Nat. Commun. 12, 3719 (2021).
Chemello, F. et al. Precise correction of Duchenne muscular dystrophy exon deletion mutations by base and prime editing. Sci. Adv. 7, abg4910 (2021).
Levy, J. M. et al. Cytosine and adenine base editing of the brain, liver, retina, heart and skeletal muscle of mice via adeno-associated viruses. Nat. Biomed. Eng. 4, 97–110 (2020).
Koblan, L. W. et al. In vivo base editing rescues Hutchinson–Gilford progeria syndrome in mice. Nature 589, 608–614 (2021).
Ma, S. et al. Efficient correction of a hypertrophic cardiomyopathy mutation by ABEmax-NG. Circ. Res. 129, 895–908 (2021).
Bose, S. K. et al. In utero adenine base editing corrects multi-organ pathology in a lethal lysosomal storage disease. Nat. Commun. 12, 4291 (2021).
McGrath, E. et al. Targeting specificity of APOBEC-based cytosine base editor in human iPSCs determined by whole genome sequencing. Nat. Commun. 10, 5353 (2019).
Jin, S. et al. Cytosine, but not adenine, base editors induce genome-wide off-target mutations in rice. Science 364, 292–295 (2019).
Zuo, E. et al. Cytosine base editor generates substantial off-target single-nucleotide variants in mouse embryos. Science 364, 289–292 (2019).
Grunewald, J. et al. Transcriptome-wide off-target RNA editing induced by CRISPR-guided DNA base editors. Nature 569, 433–437 (2019).
Rees, H. A., Wilson, C., Doman, J. L. & Liu, D. R. Analysis and minimization of cellular RNA editing by DNA adenine base editors. Sci. Adv. 5, eaax5717 (2019).
Zhou, C. et al. Off-target RNA mutation induced by DNA base editing and its elimination by mutagenesis. Nature 571, 275–278 (2019).
Anzalone, A. V. et al. Search-and-replace genome editing without double-strand breaks or donor DNA. Nature 576, 149–157 (2019).
Gao, P. et al. Prime editing in mice reveals the essentiality of a single base in driving tissue-specific gene expression. Genome Biol. 22, 83 (2021).
Abudayyeh, O. O. et al. C2c2 is a single-component programmable RNA-guided RNA-targeting CRISPR effector. Science 353, aaf5573 (2016).
East-Seletsky, A. et al. Two distinct RNase activities of CRISPR-C2c2 enable guide-RNA processing and RNA detection. Nature 538, 270–273 (2016).
Abudayyeh, O. O. et al. RNA targeting with CRISPR-Cas13. Nature 550, 280–284 (2017).
Cox, D. B. T. et al. RNA editing with CRISPR-Cas13. Science 358, 1019–1027 (2017).
Abudayyeh, O. O. et al. A cytosine deaminase for programmable single-base RNA editing. Science 365, 382–386 (2019).
Chen, B. et al. Dynamic imaging of genomic loci in living human cells by an optimized CRISPR/Cas system. Cell 155, 1479–1491 (2013).
Anton, T., Bultmann, S., Leonhardt, H. & Markaki, Y. Visualization of specific DNA sequences in living mouse embryonic stem cells with a programmable fluorescent CRISPR/Cas system. Nucleus 5, 163–172 (2014).
Yang, L. Z. et al. Dynamic imaging of RNA in living cells by CRISPR-Cas13 systems. Mol. Cell 76, 981–997 (2019).
Wang, H. et al. CRISPR-mediated live imaging of genome editing and transcription. Science 365, 1301–1305 (2019).
Fujita, T., Yuno, M., Suzuki, Y., Sugano, S. & Fujii, H. Identification of physical interactions between genomic regions by enChIP-Seq. Genes Cell 22, 506–520 (2017).
Liu, X. et al. In situ capture of chromatin interactions by biotinylated dCas9. Cell 170, 1028–1043 (2017).
Kitagawa, A. et al. CRISPR-mediated single nucleotide polymorphism modeling in rats reveals insight into reduced cardiovascular risk associated with Mediterranean G6PD variant. Hypertension 76, 523–532 (2020).
Lambert, M. et al. Characterization of Kcnk3-mutated rat, a novel model of pulmonary hypertension. Circ. Res. 125, 678–695 (2019).
Waghulde, H. et al. Attenuation of microbiotal dysbiosis and hypertension in a CRISPR/Cas9 gene ablation rat model of GPER1. Hypertension 72, 1125–1132 (2018).
Zhao, Y. et al. Hyperlipidemia induces typical atherosclerosis development in Ldlr and Apoe deficient rats. Atherosclerosis 271, 26–35 (2018).
Chen, Y. et al. Functional disruption of the dystrophin gene in rhesus monkey using CRISPR/Cas9. Hum. Mol. Genet. 24, 3764–3774 (2015).
Niu, Y. et al. Generation of gene-modified cynomolgus monkey via Cas9/RNA-mediated gene targeting in one-cell embryos. Cell 156, 836–843 (2014).
Strong, A. & Musunuru, K. Genome editing in cardiovascular diseases. Nat. Rev. Cardiol. 14, 11–20 (2017).
Shi, Y., Inoue, H., Wu, J. C. & Yamanaka, S. Induced pluripotent stem cell technology: a decade of progress. Nat. Rev. Drug Discov. 16, 115–130 (2017).
Lee, J. et al. Activation of PDGF pathway links LMNA mutation to dilated cardiomyopathy. Nature 572, 335–340 (2019).
Takahashi, K. & Yamanaka, S. Induction of pluripotent stem cells from mouse embryonic and adult fibroblast cultures by defined factors. Cell 126, 663–676 (2006).
Chen, I. Y., Matsa, E. & Wu, J. C. Induced pluripotent stem cells: at the heart of cardiovascular precision medicine. Nat. Rev. Cardiol. 13, 333–349 (2016).
Sayed, N., Liu, C. & Wu, J. C. Translation of human-induced pluripotent stem cells: from clinical trial in a dish to precision medicine. J. Am. Coll. Cardiol. 67, 2161–2176 (2016).
Tapia, N. & Scholer, H. R. Molecular obstacles to clinical translation of iPSCs. Cell Stem Cell 19, 298–309 (2016).
Magdy, T. et al. RARG variant predictive of doxorubicin-induced cardiotoxicity identifies a cardioprotective therapy. Cell Stem Cell 28, 2076–2089 (2021).
Chang, Y. J. et al. CRISPR base editing in induced pluripotent stem cells. Methods Mol. Biol. 2045, 337–346 (2019).
Lee, S. et al. Single C-to-T substitution using engineered APOBEC3G-nCas9 base editors with minimum genome- and transcriptome-wide off-target effects. Sci. Adv. 6, eaba1773 (2020).
Nishiga, M., Qi, L. S. & Wu, J. C. CRISPRi/a screening with human iPSCs. Methods Mol. Biol. 2320, 261–281 (2021).
Joung, J. et al. Genome-scale CRISPR-Cas9 knockout and transcriptional activation screening. Nat. Protoc. 12, 828–863 (2017).
Joung, J. et al. Genome-scale activation screen identifies a lncRNA locus regulating a gene neighbourhood. Nature 548, 343–346 (2017).
Shalem, O. et al. Genome-scale CRISPR-Cas9 knockout screening in human cells. Science 343, 84–87 (2014).
Liu, S. J. et al. CRISPRi-based genome-scale identification of functional long noncoding RNA loci in human cells. Science 355, aah7111 (2017).
Liu, Y. et al. CRISPR activation screens systematically identify factors that drive neuronal fate and reprogramming. Cell Stem Cell 23, 758–771 (2018).
Adamson, B. et al. A multiplexed single-cell CRISPR screening platform enables systematic dissection of the unfolded protein response. Cell 167, 1867–1882 (2016).
Dixit, A. et al. Perturb-Seq: dissecting molecular circuits with scalable single-cell RNA profiling of pooled genetic screens. Cell 167, 1853–1866 (2016).
Replogle, J. M. et al. Combinatorial single-cell CRISPR screens by direct guide RNA capture and targeted sequencing. Nat. Biotechnol. 38, 954–961 (2020).
Datlinger, P. et al. Pooled CRISPR screening with single-cell transcriptome readout. Nat. Methods 14, 297–301 (2017).
Pierce, S. E., Granja, J. M. & Greenleaf, W. J. High-throughput single-cell chromatin accessibility CRISPR screens enable unbiased identification of regulatory networks in cancer. Nat. Commun. 12, 2969 (2021).
Mimitou, E. P. et al. Multiplexed detection of proteins, transcriptomes, clonotypes and CRISPR perturbations in single cells. Nat. Methods 16, 409–412 (2019).
Feldman, D. et al. Optical pooled screens in human cells. Cell 179, 787–799 (2019).
Wang, C., Lu, T., Emanuel, G., Babcock, H. P. & Zhuang, X. Imaging-based pooled CRISPR screening reveals regulators of lncRNA localization. Proc. Natl Acad. Sci. USA 116, 10842–10851 (2019).
Tian, R. et al. Genome-wide CRISPRi/a screens in human neurons link lysosomal failure to ferroptosis. Nat. Neurosci. 24, 1020–1034 (2021).
Xu, J. et al. Genome-wide CRISPR screen identifies ZIC2 as an essential gene that controls the cell fate of early mesodermal precursors to human heart progenitors. Stem Cell 38, 741–755 (2020).
Sapp, V. et al. Genome-wide CRISPR/Cas9 screening in human iPS derived cardiomyocytes uncovers novel mediators of doxorubicin cardiotoxicity. Sci. Rep. 11, 13866 (2021).
Parvez, S. et al. MIC-Drop: a platform for large-scale in vivo CRISPR screens. Science 373, 1146–1151 (2021).
VanDusen, N. J. et al. Massively parallel in vivo CRISPR screening identifies RNF20/40 as epigenetic regulators of cardiomyocyte maturation. Nat. Commun. 12, 4442 (2021).
Juang, J. J. et al. GSTM3 variant is a novel genetic modifier in Brugada syndrome, a disease with risk of sudden cardiac death. EBioMedicine 57, 102843 (2020).
VanDusen, N. J. et al. Author Correction: massively parallel in vivo CRISPR screening identifies RNF20/40 as epigenetic regulators of cardiomyocyte maturation. Nat. Commun. 12, 5105 (2021).
Cox, D. B., Platt, R. J. & Zhang, F. Therapeutic genome editing: prospects and challenges. Nat. Med. 21, 121–131 (2015).
Nishiga, M., Qi, L. S. & Wu, J. C. Therapeutic genome editing in cardiovascular diseases. Adv. Drug Deliv. Rev. 168, 147–157 (2021).
Abifadel, M. et al. Mutations in PCSK9 cause autosomal dominant hypercholesterolemia. Nat. Genet. 34, 154–156 (2003).
Cohen, J. C., Boerwinkle, E., Mosley, T. H. Jr. & Hobbs, H. H. Sequence variations in PCSK9, low LDL, and protection against coronary heart disease. N. Engl. J. Med. 354, 1264–1272 (2006).
Ding, Q. et al. Permanent alteration of PCSK9 with in vivo CRISPR-Cas9 genome editing. Circ. Res. 115, 488–492 (2014).
Ran, F. A. et al. In vivo genome editing using Staphylococcus aureus Cas9. Nature 520, 186–191 (2015).
Wang, X. et al. CRISPR-Cas9 targeting of PCSK9 in human hepatocytes in vivo — brief report. Arterioscler. Thromb. Vasc. Biol. 36, 783–786 (2016).
Gaudelli, N. M. et al. Directed evolution of adenine base editors with increased activity and therapeutic application. Nat. Biotechnol. 38, 892–900 (2020).
Gillmore, J. D. et al. CRISPR-Cas9 in vivo gene editing for transthyretin amyloidosis. N. Engl. J. Med. 385, 493–502 (2021).
Fairclough, R. J., Wood, M. J. & Davies, K. E. Therapy for Duchenne muscular dystrophy: renewed optimism from genetic approaches. Nat. Rev. Genet. 14, 373–378 (2013).
Iftikhar, M., Frey, J., Shohan, M. J., Malek, S. & Mousa, S. A. Current and emerging therapies for Duchenne muscular dystrophy and spinal muscular atrophy. Pharmacol. Ther. 220, 107719 (2021).
Long, C. et al. Prevention of muscular dystrophy in mice by CRISPR/Cas9-mediated editing of germline DNA. Science 345, 1184–1188 (2014).
Long, C. et al. Postnatal genome editing partially restores dystrophin expression in a mouse model of muscular dystrophy. Science 351, 400–403 (2016).
Nelson, C. E. et al. In vivo genome editing improves muscle function in a mouse model of Duchenne muscular dystrophy. Science 351, 403–407 (2016).
Tabebordbar, M. et al. In vivo gene editing in dystrophic mouse muscle and muscle stem cells. Science 351, 407–411 (2016).
Young, C. S. et al. A single CRISPR-Cas9 deletion strategy that targets the majority of DMD patients restores dystrophin function in hiPSC-derived muscle cells. Cell Stem Cell 18, 533–540 (2016).
Amoasii, L. et al. Gene editing restores dystrophin expression in a canine model of Duchenne muscular dystrophy. Science 362, 86–91 (2018).
Long, C. et al. Correction of diverse muscular dystrophy mutations in human engineered heart muscle by single-site genome editing. Sci. Adv. 4, eaap9004 (2018).
Nelson, C. E. et al. Long-term evaluation of AAV-CRISPR genome editing for Duchenne muscular dystrophy. Nat. Med. 25, 427–432 (2019).
Zeng, Y. et al. Correction of the Marfan syndrome pathogenic FBN1 mutation by base editing in human cells and heterozygous embryos. Mol. Ther. 26, 2631–2637 (2018).
Kotterman, M. A. & Schaffer, D. V. Engineering adeno-associated viruses for clinical gene therapy. Nat. Rev. Genet. 15, 445–451 (2014).
Hulot, J. S., Ishikawa, K. & Hajjar, R. J. Gene therapy for the treatment of heart failure: promise postponed. Eur. Heart J. 37, 1651–1658 (2016).
Zsebo, K. et al. Long-term effects of AAV1/SERCA2a gene transfer in patients with severe heart failure: analysis of recurrent cardiovascular events and mortality. Circ. Res. 114, 101–108 (2014).
Bohm, S. et al. A gene therapy for inherited blindness using dCas9-VPR-mediated transcriptional activation. Sci. Adv. 6, eaba5614 (2020).
Kim, E. et al. In vivo genome editing with a small Cas9 orthologue derived from Campylobacter jejuni. Nat. Commun. 8, 14500 (2017).
Amrani, N. et al. NmeCas9 is an intrinsically high-fidelity genome-editing platform. Genome Biol. 19, 214 (2018).
Ibraheim, R. et al. All-in-one adeno-associated virus delivery and genome editing by Neisseria meningitidis Cas9 in vivo. Genome Biol. 19, 137 (2018).
Edraki, A. et al. A compact, high-accuracy Cas9 with a dinucleotide PAM for in vivo genome editing. Mol. Cell 73, 714–726 (2019).
Liu, J. J. et al. CasX enzymes comprise a distinct family of RNA-guided genome editors. Nature 566, 218–223 (2019).
Kannan, S. et al. Compact RNA editors with small Cas13 proteins. Nat. Biotechnol. https://doi.org/10.1038/s41587-021-01030-2 (2021).
Xu, X. et al. Engineered miniature CRISPR-Cas system for mammalian genome regulation and editing. Mol. Cell 81, 4333–4345 (2021).
Charlesworth, C. T. et al. Identification of preexisting adaptive immunity to Cas9 proteins in humans. Nat. Med. 25, 249–254 (2019).
Mitchell, M. J. et al. Engineering precision nanoparticles for drug delivery. Nat. Rev. Drug Discov. 20, 101–124 (2021).
Acknowledgements
The authors are supported by the American Heart Association (AHA) 17MERIT33610009 (J.C.W.) and 19CDA34760019 (C.L.), Leducq Foundation 18CVD05 (J.C.W.), National Institutes of Health (NIH) R01 HL126527, R01 HL123968, R01 HL141371, R01 HL141851, R01 HL150693 (J.C.W.) and U01 DK127405 (L.S.Q.), National Science Foundation CAREER Award 2046650 (L.S.Q.), Li Ka Shing Foundation (L.S.Q.) and Tobacco-Related Disease Research Program (TRDRP) postdoctoral fellowship T31FT1758 (M.N.).
Author information
Authors and Affiliations
Contributions
M.N. and C.L. researched data for the article, and M.N., C.L. and J.C.W. discussed its content. M.N., C.L., L.S.Q. and J.C.W. wrote the manuscript, and all the authors reviewed and edited the manuscript before submission.
Corresponding authors
Ethics declarations
Competing interest
L.S.Q. is a co-founder of Epicrispr Biotechnologies and Refuge Biotechnologies, and J.C.W. is a co-founder of Greenstone Biosciences. The work presented here is completely independent of these companies. The other authors declare no competing interests.
Additional information
Peer review information
Nature Reviews Cardiology thanks Joseph Miano and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Related links
ClinVar database: https://www.ncbi.nlm.nih.gov/clinvar/
Supplementary information
Glossary
- Non-homologous end-joining
-
(NHEJ). An endogenous cellular mechanism to repair double-strand breaks (DSBs), in which the ends at a cut site are directly ligated to each other.
- Microhomology-mediated end-joining
-
(MMEJ). An endogenous cellular mechanism to repair DSBs, in which microhomologous sequences at both ends of a cut site are used to align the ends, resulting in the removal of the flanking region.
- Homology-directed repair
-
(HDR). An endogenous cellular mechanism to repair DSBs, in which the sequence at a cut site is replaced by a sequence specified in a donor template, typically via homologous recombination.
- Protospacer adjacent motif
-
(PAM). A short, specific DNA sequence (2–6 nucleotides) that follows the DNA sequence targeted by a CRISPR system and is required for a Cas nuclease to bind to the target region.
- Induced pluripotent stem cell
-
(iPSC). A type of stem cell that can be generated directly from somatic cells, such as fibroblasts or blood cells, by introducing specific sets of genes.
- Adeno-associated virus
-
(AAV). A small, non-pathogenic virus that can infect many types of human cell and is often used as a vector for the delivery of gene therapy.
- Doxorubicin
-
A chemotherapy drug that is effective for many different types of cancer, including breast cancer and leukaemias, with a well-known adverse effect of cardiac toxicity.
Rights and permissions
About this article
Cite this article
Nishiga, M., Liu, C., Qi, L.S. et al. The use of new CRISPR tools in cardiovascular research and medicine. Nat Rev Cardiol 19, 505–521 (2022). https://doi.org/10.1038/s41569-021-00669-3
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41569-021-00669-3
- Springer Nature Limited
This article is cited by
-
Consequences of ionizing radiation exposure to the cardiovascular system
Nature Reviews Cardiology (2024)
-
Precision Genetic Therapies: Balancing Risk and Benefit in Patients with Heart Failure
Current Cardiology Reports (2024)
-
Recent advances in CRISPR-based genome editing technology and its applications in cardiovascular research
Military Medical Research (2023)
-
Current and future perspectives of single-cell multi-omics technologies in cardiovascular research
Nature Cardiovascular Research (2023)
-
Base editing of CaMKIIδ shows promise as cardiac therapy
Nature Reviews Cardiology (2023)