Abstract
The genetic code allows most amino acids a choice of optimal and nonoptimal codons. We report that synonymous codon choice is tuned to promote interaction of nascent polypeptides with the signal recognition particle (SRP), which assists in protein translocation across membranes. Cotranslational recognition by the SRP in vivo is enhanced when mRNAs contain nonoptimal codon clusters 35–40 codons downstream of the SRP-binding site, the distance that spans the ribosomal polypeptide exit tunnel. A local translation slowdown upon ribosomal exit of SRP-binding elements in mRNAs containing these nonoptimal codon clusters is supported experimentally by ribosome profiling analyses in yeast. Modulation of local elongation rates through codon choice appears to kinetically enhance recognition by ribosome-associated factors. We propose that cotranslational regulation of nascent-chain fate may be a general constraint shaping codon usage in the genome.






Similar content being viewed by others
References
Kim, Y.E., Hipp, M.S., Bracher, A., Hayer-Hartl, M. & Hartl, F.U. Molecular chaperone functions in protein folding and proteostasis. Annu. Rev. Biochem. 82, 323–355 (2013).
Spencer, P.S., Siller, E., Anderson, J.F. & Barral, J.M. Silent substitutions predictably alter translation elongation rates and protein folding efficiencies. J. Mol. Biol. 422, 328–335 (2012).
Xu, Y. et al. Non-optimal codon usage is a mechanism to achieve circadian clock conditionality. Nature 495, 116–120 (2013).
Dana, A. & Tuller, T. The effect of tRNA levels on decoding times of mRNA codons. Nucleic Acids Res. 42, 9171–9181 (2014).
Keenan, R.J., Freymann, D.M., Stroud, R.M. & Walter, P. The signal recognition particle. Annu. Rev. Biochem. 70, 755–775 (2001).
Akopian, D., Shen, K., Zhang, X. & Shan, S.-o. Signal recognition particle: an essential protein-targeting machine. Annu. Rev. Biochem. 82, 693–721 (2013).
Powers, E.T., Morimoto, R.I., Dillin, A., Kelly, J.W. & Balch, W.E. Biological and chemical approaches to diseases of proteostasis deficiency. Annu. Rev. Biochem. 78, 959–991 (2009).
Landry, S.J. & Gierasch, L.M. Recognition of nascent polypeptides for targeting and folding. Trends Biochem. Sci. 16, 159–163 (1991).
Halic, M. et al. Structure of the signal recognition particle interacting with the elongation-arrested ribosome. Nature 427, 808–814 (2004).
Noriega, T.R. et al. Signal recognition particle-ribosome binding is sensitive to nascent chain length. J. Biol. Chem. 289, 19294–19305 (2014).
Ast, T., Cohen, G. & Schuldiner, M. A network of cytosolic factors targets SRP-independent proteins to the endoplasmic reticulum. Cell 152, 1134–1145 (2013).
Rapoport, T.A. Protein translocation across the eukaryotic endoplasmic reticulum and bacterial plasma membranes. Nature 450, 663–669 (2007).
von Heijne, G. Signal sequences: the limits of variation. J. Mol. Biol. 184, 99–105 (1985).
Janda, C.Y. et al. Recognition of a signal peptide by the signal recognition particle. Nature 465, 507–510 (2010).
Hegde, R.S. & Kang, S.-W. The concept of translocational regulation. J. Cell Biol. 182, 225–232 (2008).
Zheng, N. & Gierasch, L.M. Signal sequences: the same yet different. Cell 86, 849–852 (1996).
Kaiser, C.A., Preuss, D., Grisafi, P. & Botstein, D. Many random sequences functionally replace the secretion signal sequence of yeast invertase. Science 235, 312–317 (1987).
Kraut-Cohen, J. & Gerst, J.E. Addressing mRNAs to the ER: cis sequences act up. Trends Biochem. Sci. 35, 459–469 (2010).
Flanagan, J.J. et al. Signal recognition particle binds to ribosome-bound signal sequences with fluorescence-detected subnanomolar affinity that does not diminish as the nascent chain lengthens. J. Biol. Chem. 278, 18628–18637 (2003).
del Alamo, M. et al. Defining the specificity of cotranslationally acting chaperones by systematic analysis of mRNAs associated with ribosome-nascent chain complexes. PLoS Biol. 9, e1001100 (2011).
Fedyukina, D.V. & Cavagnero, S. Protein folding at the exit tunnel. Annu. Rev. Biophys. 40, 337–359 (2011).
Komar, A.A. A pause for thought along the co-translational folding pathway. Trends Biochem. Sci. 34, 16–24 (2009).
Zhang, G. & Ignatova, Z. Folding at the birth of the nascent chain: coordinating translation with co-translational folding. Curr. Opin. Struct. Biol. 21, 25–31 (2011).
O'Brien, E.P., Vendruscolo, M. & Dobson, C.M. Prediction of variable translation rate effects on cotranslational protein folding. Nat. Commun. 3, 868 (2012).
Pechmann, S. & Frydman, J. Evolutionary conservation of codon optimality reveals hidden signatures of cotranslational folding. Nat. Struct. Mol. Biol. 20, 237–243 (2013).
Gingold, H. & Pilpel, Y. Determinants of translation efficiency and accuracy. Mol. Syst. Biol. 7, 481 (2011).
Tuller, T. et al. An evolutionarily conserved mechanism for controlling the efficiency of protein translation. Cell 141, 344–354 (2010).
Ingolia, N.T., Ghaemmaghami, S., Newman, J.R.S. & Weissman, J.S. Genome-wide analysis in vivo of translation with nucleotide resolution using ribosome profiling. Science 324, 218–223 (2009).
Zinshteyn, B. & Gilbert, W.V. Loss of a conserved tRNA anticodon modification perturbs cellular signaling. PLoS Genet. 9, e1003675 (2013).
Berndt, U., Oellerer, S., Zhang, Y., Johnson, A.E. & Rospert, S. A signal-anchor sequence stimulates signal recognition particle binding to ribosomes from inside the exit tunnel. Proc. Natl. Acad. Sci. USA 106, 1398–1403 (2009).
Charneski, C.A. & Hurst, L.D. Positively charged residues are the major determinants of ribosomal velocity. PLoS Biol. 11, e1001508 (2013).
Dalley, J.A., Selkirk, A. & Pool, M.R. Access to ribosomal protein Rpl25p by the signal recognition particle is required for efficient cotranslational translocation. Mol. Biol. Cell 19, 2876–2884 (2008).
Zhang, D. & Shan, S.-o. Translation elongation regulates substrate selection by the signal recognition particle. J. Biol. Chem. 287, 7652–7660 (2012).
Mason, N., Ciufo, L.F. & Brown, J.D. Elongation arrest is a physiologically important function of signal recognition particle. EMBO J. 19, 4164–4174 (2000).
Doerfel, L.K. et al. EF-P is essential for rapid synthesis of proteins containing consecutive proline residues. Science 339, 85–88 (2013).
Yang, J.R., Chen, X. & Zhang, J. Codon-by-codon modulation of translational speed and accuracy via mRNA folding. PLoS Biol. 12, e1001910 (2014).
Sherman, M.Y. & Qian, S.-B. Less is more: improving proteostasis by translation slow down. Trends Biochem. Sci. 38, 585–591 (2013).
Kramer, G., Boehringer, D., Ban, N. & Bukau, B. The ribosome as a platform for co-translational processing, folding and targeting of newly synthesized proteins. Nat. Struct. Mol. Biol. 16, 589–597 (2009).
Fluman, N., Navon, S., Bibi, E. & Pilpel, Y. mRNA-programmed translation pauses in the targeting of E. coli membrane proteins. Elife 3, e03440 (2014).
El Yacoubi, B., Bailly, M. & De Crécy-Lagard, V. Biosynthesis and function of posttranscriptional modifications of transfer RNAs. Annu. Rev. Genet. 46, 69–95 (2012).
Tusher, V.G., Tibshirani, R. & Chu, G. Significance analysis of microarrays applied to the ionizing radiation response. Proc. Natl. Acad. Sci. USA 98, 5116–5121 (2001).
Willmund, F. et al. The cotranslational function of ribosome-associated Hsp70 in eukaryotic protein homeostasis. Cell 152, 196–209 (2013).
Krogh, A., Larsson, B., Von Heijne, G. & Sonnhammer, E.L. Predicting transmembrane protein topology with a hidden Markov model: application to complete genomes. J. Mol. Biol. 305, 567–580 (2001).
Nielsen, H., Engelbrecht, J., Brunak, S. & Von Heijne, G. Identification of prokaryotic and eukaryotic signal peptides and prediction of their cleavage sites. Protein Eng. 10, 1–6 (1997).
Käll, L., Krogh, A. & Sonnhammer, E.L. A combined transmembrane topology and signal peptide prediction method. J. Mol. Biol. 338, 1027–1036 (2004).
Holstege, F.C. et al. Dissecting the regulatory circuitry of a eukaryotic genome. Cell 95, 717–728 (1998).
Pechmann, S., Levy, E.D., Tartaglia, G.G. & Vendruscolo, M. Physicochemical principles that regulate the competition between functional and dysfunctional association of proteins. Proc. Natl. Acad. Sci. USA 106, 10159–10164 (2009).
D'haeseleer, P. What are DNA sequence motifs? Nat. Biotechnol. 24, 423–425 (2006).
Papadopoulos, J.S. & Agarwala, R. COBALT: constraint-based alignment tool for multiple protein sequences. Bioinformatics 23, 1073–1079 (2007).
dos Reis, M., Savva, R. & Wernisch, L. Solving the riddle of codon usage preferences: a test for translational selection. Nucleic Acids Res. 32, 5036–5044 (2004).
Qian, W., Yang, J.R., Pearson, N.M., Maclean, C. & Zhang, J. Balanced codon usage optimizes eukaryotic translational efficiency. PLoS Genet. 8, e1002603 (2012).
Li, G.W., Oh, E. & Weissman, J.S. The anti-Shine–Dalgarno sequence drives translational pausing and codon choice in bacteria. Nature 484, 538–541 (2012).
Zhang, F., Saha, S., Shabalina, S.A. & Kashina, A. Differential arginylation of actin isoforms is regulated by coding sequence–dependent degradation. Science 329, 1534–1537 (2010).
Zhang, G., Hubalewska, M. & Ignatova, Z. Transient ribosomal attenuation coordinates protein synthesis and co-translational folding. Nat. Struct. Mol. Biol. 16, 274–280 (2009).
Kimchi-Sarfaty, C. et al. A “silent” polymorphism in the MDR1 gene changes substrate specificity. Science 315, 525–528 (2007).
Fedyunin, I. et al. tRNA concentration fine tunes protein solubility. FEBS Lett. 586, 3336–3340 (2012).
Coleman, J.R. et al. Virus attenuation by genome-scale changes in codon pair bias. science 320, 1784–1787 (2008).
Acknowledgements
We thank members of the Frydman laboratory for helpful discussions and K. Dalton for comments on the manuscript. We gratefully acknowledge support from a European Molecular Biology Organization Long-Term Fellowship (ALTF 1334-2010) to S.P., US National Institutes of Health (NIH) grant GM108325 to J.C. and NIH grant GM56433 and Human Frontier Science Program Grant RGP0025/2012 to J.F.
Author information
Authors and Affiliations
Contributions
S.P. and J.F. conceived the research project. S.P. performed all computational analyses. J.C. performed the translocation experiment. S.P. and J.F. wrote the manuscript; all authors discussed the results and commented on the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Integrated supplementary information
Supplementary Figure 1 Classification of cotranslational SRP substrates.
a, Schematic overview of the classes of cotranslational SRP substrates analyzed in this study. b, Distributions of the length of the hydrophobic region in the SS for SRP-SE and SRP-E proteins. For SRP-NC proteins, we show the lengths of the first stretch of at least 5 hydrophobic amino acids. c, Comparison of the SRP pull-down to the translatome highlights enrichment in SRP interaction relative to the level of translation. Strongly enriched SS-proteins (SRP-SE) are shown in orange. d, Overview of the number of SRP substrates in each class.
Supplementary Figure 2 Hydrophobicity or putative sequence motifs alone cannot explain SRP enrichment.
a, Histogram of the length of the hydrophobic region in SS and TM segments. b, A compound score is defined as the product of the maximum hydrophobicity and the length of the hydrophobic region in SS or TM segments. Shown is the histogram of the compound score for all cotranslational SRP substrates. c, The compound score is highly correlated with the length of the hydrophobic region as its strongest determinant. d, Representative median hydrophobicity profiles of the SS of SRP-SE (n=29) and SRP-E (n=78) substrates, and longest N-terminal hydrophobic stretch for SRP-NC (n=109) substrates. e, Distributions of the most hydrophobic stretch of length 8, possibly the optimal SRP binding site, within the SS of SRP-E and SRP-SE proteins indicate no significant global differences in hydrophobicity. f, The most hydrophobic region of length 9–12 in the SS or N-terminal hydrophobic stretches is identified, and the distributions for SRP-SE, SRP-E and SRP-NC proteins compared. Independent of the length of the region considered, SRP-SE SS are not significantly more hydrophobic than SRP-E SS. g, Schematic overview of the quantification of possible sequence motifs in the hydrophobic regions of SS. In separate analyses, the sequences are aligned either at the start of the hydrophobic region, or as the best local alignment without gaps. From the sequence alignments, sequence logos, the alignment matrix and the information content (IC) are computed. h, Sequence logos of the hydrophobic binding sites in SRP-NC, SRP-E and SRP-SE substrates show no distinct sequence motifs for SRP-SE proteins that may explain their stronger enrichment. j, Distribution of the IC of the hydrophobic region in random subsets of SS-proteins. Because the IC for the SRP-SE proteins (solid line) falls very close to the expected mean of the random distribution (dotted line) instead of being clearly shifted to the right, we conclude that there is no pronounced sequence motif in the hydrophobic regions of SS that sets SRP-SE substrates apart and may explain their higher enrichment scores.
Supplementary Figure 3 Translational efficiency (TE) of translocated proteins.
a, Because secretory and trans-membrane proteins are translated in a spatially-organized manner at or near the ER membrane, we derived a scale of codon-specific TEs based on the availability of the cellular tRNA pool and the codon usage of the secretome. The 20% lowest efficiency codons were defined as “nonoptimal”. Nonoptimal codons that are at least 3-fold degenerate are indicated by “*”. b, The normalized TE scales that incorporate the codon usage of the full proteome (nTEP), and, alternatively only the codon usage of the secretome (nTES), differ as predicted: codons encoding hydrophobic amino acids become less efficient, and those encoded charged residues become more efficient. However, the differences, show by per-codon fold changes, are relatively small. c, The nTEP and nTES scales are very highly correlated and lead to identical definitions of nonoptimal codons. This analysis served to validate that our results are robust, irrespective of predicting the TE for the full proteome or only secreted and trans-membrane proteins.
Supplementary Figure 4 SRP recognition of SS proteins.
a, SRP-dependent translocation of the essential N-oligosaccharyl transferase (OST). OST subunits are some of the most enriched SRP substrates. b, Four of eight OST subunits, namely Ost1, Ost3, Swp1, and Wbp1 have a distinct structure of a SS followed by a long lumenal domain. The topology of the other four subunits, Ost2, Ost4, Ost5, and Stt3 are shown for comparison. c, The coding sequence of Invertase, the model substrate that could accommodate many random sequences as functional SS, is characterized by a very pronounced region of low TE (grey bar) ca. 45 codons downstream of the start of the hydrophobic region of the SS (yellow bar). d, Distribution of average predicted translational efficiencies 38-45 codons downstream of the start of the signal sequences for strongly-enriched (SRP-SE) and enriched (SRP-E) cotranslational SRP substrates compared to the average predicted translational efficiencies 38-45 codons downstream of N-terminal hydrophobic stretches in non-cognate SRP substrates (SRP-NC) and cytosolic proteins that do not bind to SRP (Control). e, mRNA expression levels (log scale) of SRP-SE, SRP-E, SRP-NC and Control genes. While TE generally correlates with expression, the strong dip in TE in SRP-SE substrates is not confounded by expression. SRP-NC substrates may interact non-canonically because they are translated generally more slowly linked to their lower levels of expression.
Supplementary Figure 5 Ribosome profiling analyses of SRP substrates.
a, Exemplary ribosome footprint density profiles for two replicas (top), and the resulting average profile (bottom). Plotted are the sequenced footprints per position. b, Relationship between average coverage and correlation between replicas in the analyzed ribosome profiling dataset. A minimum average coverage of ten sequencing reads per position eliminates many cases of very low reproducibility. c, Relative footprint density profiles for the OST subunits with SS. d, The median relative footprint density profile for the OST subunits reflects the consistent characteristics of the individual profiles.
Supplementary Figure 6 Analysis of REST sequences in TM proteins.
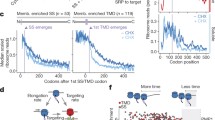
a, The average TE of individual TM-SE profiles (n=46) in the translational slowdown element downstream of the TM segments is significantly lower than in TM-nSE (n=165) substrates (Wilcoxon rank-sum test: p = 0.045). b, Distributions of the average footprint densities in the translational slowdown element is higher in TM-SE proteins (n=14) than in TM-nSE proteins (n=27), but the difference is not significant (Wilcoxon rank-sum test: p = 0.072). c, The translational slowdown element downstream of TM helices is more pronounced for more strongly enriched SRP substrates. Shown are medium TE profiles for all TM-proteins with dTM1–TM2 > 60 codons including those that do not significantly interact with SRP cotranslationally, only those that significantly interact with SRP cotranslationally, and those that are strongly enriched in SRP binding. d, The alpha-1,6-mannosyltransferase MNN11 has a single signal-anchor TM domain near the N-terminus that acts as SRP binding site. The region downstream of the TM, which is translated when the TM domain has emerged from the ribosome exit tunnel, is characterized by: e, locally low predicted translational efficiency, and f, high ribosome footprint density. Both the lower translational efficiency and the higher footprint density are observed ca. 55-60 codons downstream of the start of the TM, allowing the full TM domain to emerge from the ribosome exit tunnel. g, The distinct translational efficiency profile of MNN11 is evolutionarily conserved across yeasts, here shown for the closely related S. paradoxus, S. mikatae, S. bayanus, and D. hansenii. h, The average TE of TM segments themselves is comparable between TE-SE and TE-nSE proteins, but significantly lower than the average across the proteome. In agreement, the average footprint densities of the TM segments in TM-SE and TM-nSE substrates are significantly higher than for random regions of the same length across the proteome. Of note, the first TM segments of the TM-SE proteins are also on average characterized by higher footprint densities than the first TM segments in the TM-nSE proteins. j, Both predicted TE and experimental footprint densities of the SS themselves do not differ between SRP-SE and SRP-E proteins. Differences between distributions are tested by the Wilcoxon rank-sum test, and significance levels are indicated as *: p < 0.05; **: p < 0.01; ***: p < 0.001.
Supplementary Figure 7 Loss of REST element causes translocation defect.
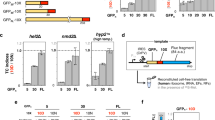
a, Translocation assays were performed as described in Dalley et al. (Mol Biol Cell, 2008). The plasmid pMP211 was kindly provided by Martin Pool (University of Manchester, UK). This pRS314-derived vector encodes an open reading frame, under control of the PHO5 promoter, comprising the first 82 residues of S. cerevisiae Pho8p fused to the amino terminus of the complete S. cerevisiae Ura3p. Proper secretion localizes Ura3p to the ER lumen where it cannot access its substrate orotidine-5′-phosphate (OMP) to produce uridine monophosphate (UMP). Secretion deficiencies accumulate functional Ura3p in the cytosol conferring uracil prototrophy. b, Synonymous mutations improving translational efficiency within the REST region of Pho8p were introduced, converting the sequence 5′-ATATTCTTCGTGACGGATGGAATGGGACCT-3′ to 5′-ATCTTCTTCGTCACCGACGGCATGGGCCCG-3′. c, Plasmids were transformed into BY4741 and grown on synthetic defined agar with or without uracil. Colonies began to appear with the no-REST plasmid after two days and continued to accumulate over a week (shown at day 6), indicating Ura3 accumulation is stochastic. After 7 days, colonies began to appear with the REST plasmid. Plasmids were recovered from five colonies of varying size from each strain and sequenced. No mutations occurred within the open reading frames, promoter, or untranslated regions.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–7 (PDF 4995 kb)
Supplementary Table 1
Translational efficiency of translocated proteins (XLSX 18 kb)
Rights and permissions
About this article
Cite this article
Pechmann, S., Chartron, J. & Frydman, J. Local slowdown of translation by nonoptimal codons promotes nascent-chain recognition by SRP in vivo. Nat Struct Mol Biol 21, 1100–1105 (2014). https://doi.org/10.1038/nsmb.2919
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nsmb.2919
- Springer Nature America, Inc.
This article is cited by
-
Genome-wide sequencing identifies a thermal-tolerance related synonymous mutation in the mussel, Mytilisepta virgata
Communications Biology (2023)
-
Unconventional secretion of Magnaporthe oryzae effectors in rice cells is regulated by tRNA modification and codon usage control
Nature Microbiology (2023)
-
Oligodendrocyte differentiation alters tRNA modifications and codon optimality-mediated mRNA decay
Nature Communications (2022)
-
Ageing exacerbates ribosome pausing to disrupt cotranslational proteostasis
Nature (2022)
-
Codon optimization of the synthetic 3-ketosphinganine reductase (3KSR) protein for enhancing sphingolipid biosynthetic enzyme expression
Molecular & Cellular Toxicology (2021)