Abstract
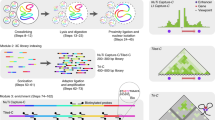
With the advent of massively parallel sequencing, considerable work has gone into adapting chromosome conformation capture (3C) techniques to study chromosomal architecture at a genome-wide scale. We recently demonstrated that the inactive murine X chromosome adopts a bipartite structure using a novel 3C protocol, termed in situ DNase Hi-C. Like traditional Hi-C protocols, in situ DNase Hi-C requires that chromatin be chemically cross-linked, digested, end-repaired, and proximity-ligated with a biotinylated bridge adaptor. The resulting ligation products are optionally sheared, affinity-purified via streptavidin bead immobilization, and subjected to traditional next-generation library preparation for Illumina paired-end sequencing. Importantly, in situ DNase Hi-C obviates the dependence on a restriction enzyme to digest chromatin, instead relying on the endonuclease DNase I. Libraries generated by in situ DNase Hi-C have a higher effective resolution than traditional Hi-C libraries, which makes them valuable in cases in which high sequencing depth is allowed for, or when hybrid capture technologies are expected to be used. The protocol described here, which involves ∼4 d of bench work, is optimized for the study of mammalian cells, but it can be broadly applicable to any cell or tissue of interest, given experimental parameter optimization.




Similar content being viewed by others
Accession codes
References
Cremer, T. & Cremer, C. Chromosome territories, nuclear architecture and gene regulation in mammalian cells. Nat. Rev. Genet. 2, 292–301 (2001).
Fraser, P. & Bickmore, W. Nuclear organization of the genome and the potential for gene regulation. Nature 447, 413–417 (2007).
Rieder, C.L. & Khodjakov, A. Mitosis through the microscope: advances in seeing inside live dividing cells. Science 300, 91–96 (2003).
Barr, M.L. & Bertram, E.G. A morphological distinction between neurones of the male and female, and the behaviour of the nucleolar satellite during accelerated nucleoprotein synthesis. Nature 163, 676 (1949).
Hochstrasser, M. & Sedat, J.W. Three-dimensional organization of Drosophila melanogaster interphase nuclei. I. Tissue-specific aspects of polytene nuclear architecture. J. Cell Biol. 104, 1455–1470 (1987).
Manuelidis, L. Individual interphase chromosome domains revealed by in situ hybridization. Hum. Genet. 71, 288–293 (1985).
Pinkel, D., Straume, T. & Gray, J.W. Cytogenetic analysis using quantitative, high-sensitivity, fluorescence hybridization. Proc. Natl. Acad. Sci. USA 83, 2934–2938 (1986).
Schardin, M., Cremer, T., Hager, H.D. & Lang, M. Specific staining of human chromosomes in Chinese hamster x man hybrid cell lines demonstrates interphase chromosome territories. Hum. Genet. 71, 281–287 (1985).
Lawrence, J.B., Singer, R.H. & Marselle, L.M. Highly localized tracks of specific transcripts within interphase nuclei visualized by in situ hybridization. Cell 57, 493–502 (1989).
Zirbel, R.M., Mathieu, U.R., Kurz, A., Cremer, T. & Lichter, P. Evidence for a nuclear compartment of transcription and splicing located at chromosome domain boundaries. Chromosome Res. 1, 93–106 (1993).
van Steensel, B. & Henikoff, S. Identification of in vivo DNA targets of chromatin proteins using tethered dam methyltransferase. Nat. Biotechnol. 18, 424–428 (2000).
Vogel, M.J., Peric-Hupkes, D. & van Steensel, B. Detection of in vivo protein-DNA interactions using DamID in mammalian cells. Nat. Protoc. 2, 1467–1478 (2007).
Orian, A. et al. Genomic binding by the Drosophila Myc, Max, Mad/Mnt transcription factor network. Genes Dev. 17, 1101–1114 (2003).
de Wit, E., Greil, F. & van Steensel, B. Genome-wide HP1 binding in Drosophila: developmental plasticity and genomic targeting signals. Genome Res. 15, 1265–1273 (2005).
Izzo, A. et al. The genomic landscape of the somatic linker histone subtypes H1.1 to H1.5 in human cells. Cell Rep. 3, 2142–2154 (2013).
Guelen, L. et al. Domain organization of human chromosomes revealed by mapping of nuclear lamina interactions. Nature 453, 948–951 (2008).
Dekker, J., Rippe, K., Dekker, M. & Kleckner, N. Capturing chromosome conformation. Science 295, 1306–1311 (2002).
Mukherjee, S., Erickson, H. & Bastia, D. Enhancer-origin interaction in plasmid R6K involves a DNA loop mediated by initiator protein. Cell 52, 375–383 (1988).
Cullen, K.E. et al. Interaction between transcription regulatory regions of prolactin chromatin. Science 261, 203–206 (1993).
de Wit, E. et al. Nuclear organization of active and inactive chromatin domains uncovered by chromosome conformation capture-on-chip (4C). Nat. Genet. 38, 1348–1354 (2006).
Dostie, J. et al. Chromosome conformation capture carbon copy (5C): A massively parallel solution for mapping interactions between genomic elements. Genome Res. 16, 1299–1309 (2006).
Lieberman-Aiden, E. et al. Comprehensive mapping of long-range interactions reveals folding principles of the human genome. Science 326, 289–293 (2009).
Duan, Z. et al. A three-dimensional model of the yeast genome. Nature 465, 363–367 (2010).
Kalhor, R., Tjong, H., Jayathilaka, N., Alber, F. & Chen, L. Genome architectures revealed by tethered chromosome conformation capture and population-based modeling. Nat. Biotechnol. 30, 90–98 (2012).
Sexton, T. et al. Three-dimensional folding and functional organization principles of the Drosophila genome. Cell 148, 458–472 (2012).
Zhang, Y. et al. Spatial organization of the mouse genome and its role in recurrent chromosomal translocations. Cell 148, 908–921 (2012).
Le, T.B.K. et al. High-resolution mapping of the spatial organization of a bacterial chromosome. Science 342, 731–734 (2013).
Mizuguchi, T. et al. Cohesin-dependent globules and heterochromatin shape 3D genome architecture in S. pombe. Nature 516, 432–435 (2014).
Dixon, J.R. et al. Chromatin architecture reorganization during stem cell differentiation. Nature 518, 331–336 (2015).
Naumova, N. et al. Organization of the mitotic chromosome. Science 342, 948–953 (2013).
Ay, F. et al. Three-dimensional modeling of the P. falciparum genome during the erythrocytic cycle reveals a strong connection between genome architecture and gene expression. Genome Res. 24, 974–988 (2014).
Splinter, E. et al. The inactive X chromosome adopts a unique three-dimensional conformation that is dependent on Xist RNA. Genes Dev. 25, 1371–1383 (2011).
Nora, E.P. et al. Spatial partitioning of the regulatory landscape of the X-inactivation centre. Nature 485, 381–385 (2012).
Giorgetti, L. et al. Predictive polymer modeling reveals coupled fluctuations in chromosome conformation and transcription. Cell 157, 950–963 (2014).
Minajigi, A. et al. Chromosomes. A comprehensive Xist interactome reveals cohesin repulsion and an RNA-directed chromosome conformation. Science 349, aab2276– (2015).
Dixon, J.R. et al. Topological domains in mammalian genomes identified by analysis of chromatin interactions. Nature 485, 376–380 (2012).
Hsieh, T.-H.S. et al. Mapping nucleosome resolution chromosome folding in yeast by Micro-C. Cell 162, 108–119 (2015).
ENCODE Project Consortium. An integrated encyclopedia of DNA elements in the human genome. Nature 489, 57–74 (2012).
Sabo, P.J. et al. Integrative analysis of 111 reference human epigenomes. Nature 518, 317–330 (2015).
Levine, M., Cattoglio, C. & Tjian, R. Looping back to leap forward: transcription enters a new era. Cell 157, 13–25 (2014).
Griffith, J., Hochschild, A. & Ptashne, M. DNA loops induced by cooperative binding of lambda repressor. Nature 322, 750–752 (1986).
Müller, H.-P., Sogo, J. & Schaffner, W. An enhancer stimulates transcription in trans when attached to the promoter via a protein bridge. Cell 58, 767–777 (1989).
Rao, S.S.P. et al. A 3D map of the human genome at kilobase resolution reveals principles of chromatin looping. Cell 159, 1665–1680 (2014).
Nagano, T. et al. Single-cell Hi-C reveals cell-to-cell variability in chromosome structure. Nature 502, 59–64 (2013).
Mifsud, B. et al. Mapping long-range promoter contacts in human cells with high-resolution capture Hi-C. Nat Genet. 47, 598–606 (2015).
Schoenfelder, S. et al. The pluripotent regulatory circuitry connecting promoters to their long-range interacting elements. Genome Res. 25, 582–597 (2015).
Sahlén, P. et al. Genome-wide mapping of promoter-anchored interactions with close to single-enhancer resolution. Genome Biol. 16, 156 (2015).
Ma, W. et al. Fine-scale chromatin interaction maps reveal the cis-regulatory landscape of human lincRNA genes. Nat Methods 12, 71–78 (2015).
Gavrilov, A.A. et al. Disclosure of a structural milieu for the proximity ligation reveals the elusive nature of an active chromatin hub. Nucleic Acids Res. 41, 3563–3575 (2013).
Nagano, T. et al. Comparison of Hi-C results using in-solution versus in-nucleus ligation. Genome Biol. 16, 175 (2015).
Deng, X. et al. Bipartite structure of the inactive mouse X chromosome. Genome Biol. 16, 152 (2015).
He, H.H. et al. Refined DNase-seq protocol and data analysis reveals intrinsic bias in transcription factor footprint identification. Nat. Methods 11, 73–78 (2014).
Imakaev, M. et al. Iterative correction of Hi-C data reveals hallmarks of chromosome organization. Nat. Methods 9, 999–1003 (2012).
de Wit, E. et al. The pluripotent genome in three dimensions is shaped around pluripotency factors. Nature 501, 227–231 (2013).
Acknowledgements
We thank S. Kasinathan and members of the Shendure laboratory for helpful comments on the manuscript. This work was funded by a National Institutes of Health (NIH) Director's Pioneer Award (1DP1HG007811Q14 to J.S.) and an NIH National Human Genome Research Institute (NHGRI) Genome Training Grant (5T32HG000035 to V.R.).
Author information
Authors and Affiliations
Contributions
V.R. and Z.D. developed the protocol. V.R., D.A.C., R.J.H., R.Q., and Z.D. performed experiments and optimized the protocol. W.M. and W.S.N. devised the processing pipeline for in situ DNase Hi-C data. X.D., C.A.B., C.M.D., W.S.N., J.S., and Z.D. supervised the research. V.R., J.S., and Z.D. wrote the manuscript, with input from all authors.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Integrated supplementary information
Supplementary Figure 1 Relative Abundances of ligation types in 3 biological replicate GM12878 libraries, vs. a Patski library
In situ DNase Hi-C reads demonstrate some cell-type specificity for the relative breakdown in long-range intrachromosomal read-pairs with respect to interchromosomal and short-range read-pairs, but these differences are reproducible, as shown when comparing three biological replicate libraries derived from the immortalized lymphoblastoid GM12878 cell line.
Supplementary information
Supplementary Figure 1
Relative abundance of ligation types in three biological replicate GM12878 libraries versus a Patski library (PDF 234 kb)
Rights and permissions
About this article
Cite this article
Ramani, V., Cusanovich, D., Hause, R. et al. Mapping 3D genome architecture through in situ DNase Hi-C. Nat Protoc 11, 2104–2121 (2016). https://doi.org/10.1038/nprot.2016.126
Published:
Issue Date:
DOI: https://doi.org/10.1038/nprot.2016.126
- Springer Nature Limited
This article is cited by
-
Orchestrating chromosome conformation capture analysis with Bioconductor
Nature Communications (2024)
-
Computational methods for analysing multiscale 3D genome organization
Nature Reviews Genetics (2024)
-
3D genomics and its applications in precision medicine
Cellular & Molecular Biology Letters (2023)
-
Predicting scale-dependent chromatin polymer properties from systematic coarse-graining
Nature Communications (2023)
-
Visualization and data exploration of chromosome conformation capture data using Voronoi diagrams with v3c-viz
Scientific Reports (2023)