Abstract
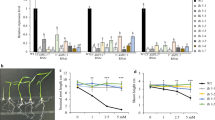
NH4+ is an important nitrogen resource for rice plants in paddy soil. Therefore, it is likely that NH4+-triggered plant growth interacts with phytohormone-mediated developmental mechanisms. Our previous transcriptomic analysis revealed that many genes involved in auxin signaling and efflux are sensitive to NH4+. In the current study, we found that NH4+ treatment causes a delayed gravity response in rice roots. To further elucidate the interlocking relationship between NH4+ and auxin signaling during root development, we utilized mutants and overexpressors of a key NH4+ signaling transcription factor INDETERMINATE DOMAIN 10 (IDD10), encoding a transcription factor that regulates the expression of NH4+ uptake and N-assimilation genes. We obtained several lines of evidence that auxin affects NH4+-mediated gene expression and root development in rice plants via IDD10. First, the gravity response was delayed in idd10 roots and accelerated in IDD10 overexpressor (IDD10 OX) roots in the absence and (especially) presence of NH4+. Second, idd10 plants showed strong root coiling only in the presence of NH4+. However, treatment of 1-N-naphthylphthalamic acid (NPA), a polar auxin transport inhibitor suppressed the NH4+-specific root phenotype of idd10. Third, the expression of NH4+-responsive auxin-related genes was affected in idd10 and IDD10 overexpressors. Finally, IDD10 expression was induced by IAA and suppressed by NPA. These findings suggest that the gene expression patterns and phenotypes triggered by NH4+ are influenced by the actions of auxin during root development, pointing to a regulatory circuit between NH4+ and auxin signaling that functions in root development in rice.
Article PDF
Similar content being viewed by others
Avoid common mistakes on your manuscript.
References
Abiko T, Obara M, Ushioda A, Hayakawa T, Hodges M, Yamaya T (2005) Localization of NAD-isocitrate dehydrogenase and glutamate dehydrogenase in rice roots: candidates for providing carbon skeletons to NADH-glutamate synthase. Plant Cell Physiol 46: 1724−1734
Bi YM, Wang RL, Zhu T, Rothstein SJ (2007) Global transcription profiling reveals differential responses to chronic nitrogen stress and putative nitrogen regulatory components in Arabidopsis. BMC Genomics 8: 281
Britto DT, Kronzucker HJ (2002) NH4+ toxicity in higher plants: a critical review. J Plant Physiol 159: 567−584
Cai H, Lu Y, Xie W, Zhu T, Lian X (2012) Transcriptome response to nitrogen starvation in rice. J Bioscience 37: 731−747
Cao Y, Glass AD, Crawford NM (1993) Ammonium inhibition of Arabidopsis root growth can be reversed by potassium and by auxin resistance mutations aux1, axr1, and axr2. Plant Physiol 102: 983–989
Chandran AK, Priatama RA, Kumar V, Xuan Y, Je BI, Kim CM, Jung KH, Han CD (2016) Genome-wide transcriptome analysis of expression in rice seedling roots in response to supplemental nitrogen. J Plant Physiol 200: 62−75
Gifford ML, Dean A, Gutierrez RA, Coruzzi GM, Birnbaum KD (2008) Cell-specific nitrogen responses mediate developmental plasticity. P Natl Acad Sci USA 105: 803−808
Hirano T, Satoh Y, Ohki A, Takada R, Arai T, Michiyama H (2008) Inhibition of ammonium assimilation restores elongation of seminal rice roots repressed by high levels of exogenous ammonium. Physiol Plantarum 134: 183−190
Krouk G, Lacombe B, Bielach A, Perrine-Walker F, Malinska K, Mounier E, Hoyerova K, Tillard P, Leon S, Ljung K, Zazimalova E, Benkova E, Nacry P, Gojon A (2010) Nitrate-regulated auxin transport by NRT1.1 defines a mechanism for nutrient sensing in plants. Dev Cell 18: 927−937
Li B, Li Q, Su Y, Chen H, Xiong L, Mi G, Kronzucker HJ, Shi W (2011) Shoot-supplied ammonium targets the root auxin influx carrier AUX1 and inhibits lateral root emergence in Arabidopsis. Plant Cell Environ 34: 933−946
Li B, Li Q, Xiong L, Kronzucker HJ, Kramer U, Shi W (2012) Arabidopsis plastid AMOS1/EGY1 integrates abscisic acid signaling to regulate global gene expression response to ammonium stress. Plant Physiol 160: 2040−2051
Li G, Li B, Dong G, Feng X, Kronzucker HJ, Shi W (2013) Ammoniuminduced shoot ethylene production is associated with the inhibition of lateral root formation in Arabidopsis. J Exp Bot 64: 1413−1425
Lian X, Wang S, Zhang J, Feng Q, Zhang L, Fan D, Li X, Yuan D, Han B, Zhang Q (2006) Expression profiles of 10,422 genes at early stage of low nitrogen stress in rice assayed using a cDNA microarray. Plant Mol Biol 60: 617−631
Mao C, Wang S, Jia Q, Wu P (2006) OsEIL1, a rice homolog of the Arabidopsis EIN3 regulates the ethylene response as a positive component. Plant Mol Biol 61: 141−152
Patterson K, Cakmak T, Cooper A, Lager I, Rasmusson AG, Escobar MA (2010) Distinct signalling pathways and transcriptome response signatures differentiate ammonium-and nitrate-supplied plants. Plant Cell Environ 33: 1486−1501
Scheible WR, Morcuende R, Czechowski T, Fritz C, Osuna D, Palacios-Rojas N, Schindelasch D, Thimm O, Udvardi MK, Stitt M (2004) Genome-wide reprogramming of primary and secondary metabolism, protein synthesis, cellular growth processes, and the regulatory infrastructure of Arabidopsis in response to nitrogen. Plant Physiol 136: 2483−2499
Shimizu H, Tanabata T, Xie X, Inagaki N, Takano M, Shinomura T, Yamamoto KT (2009) Phytochrome-mediated growth inhibition of seminal roots in rice seedlings. Physiol Plantarum 137: 289−297
Wang R, Guegler K, LaBrie ST, Crawford NM (2000) Genomic analysis of a nutrient response in Arabidopsis reveals diverse expression patterns and novel metabolic and potential regulatory genes induced by nitrate. Plant Cell 12: 1491−1509
Wang R, Okamoto M, Xing X, Crawford NM (2003) Microarray analysis of the nitrate response in Arabidopsis roots and shoots reveals over 1,000 rapidly responding genes and new linkages to glucose, trehalose-6-phosphate, iron, and sulfate metabolism. Plant Physiol 132: 556−567
Xuan YH, Priatama RA, Huang J, Je BI, Liu JM, Park SJ, Piao HL, Son DY, Lee JJ, Park SH, Jung KH, Kim TH, Han CD (2013) Indeterminate domain 10 regulates ammonium-mediated gene expression in rice roots. New Phytol 197: 791−804
Xuan YH, Duan FY, Je BI, Kim CM, Li TY, Liu JM, Park SJ, Cho JH, Kim TH, von Wirén N, Han CD (2016) Related to ABI3/VP1-Like 1 (RAVL1) regulates brassinosteroid-mediated activation of AMT1;2 in rice (Oryza sativa). J Exp Bot 68: 727−737
Yang SY, Hao DL, Song ZZ, Yang GZ, Wang L, Su YH (2015a) RNA-Seq analysis of differentially expressed genes in rice under varied nitrogen supplies. Gene 555: 305−317
Yang W, Yoon J, Choi H, Fan Y, Chen R, An G (2015b) Transcriptome analysis of nitrogen-starvation-responsive genes in rice. BMC Plant Biol 15: 31
Zou N, Li B, Chen H, Su Y, Kronzucker HJ, Xiong L, Baluska F, Shi W (2013) GSA-1/ARG1 protects root gravitropism in Arabidopsis under ammonium stress. New Phytol 200: 97−111
Author information
Authors and Affiliations
Corresponding authors
Electronic supplementary material
Rights and permissions
About this article
Cite this article
Xuan, Y.H., Kumar, V., Zhu, X.F. et al. IDD10 is Involved in the Interaction between NH4+ and Auxin Signaling in Rice Roots. J. Plant Biol. 61, 72–79 (2018). https://doi.org/10.1007/s12374-017-0423-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12374-017-0423-2