Abstract
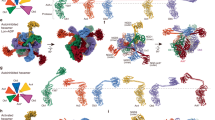
Lon proteases degrade defective or denature proteins as well as some folded proteins for the control of cellular protein quality. There are two types of Lon proteases, LonA and LonB. Each consists of two functional components: a protease component and an ATPase associated with various cellular activities (AAA+ module). Here, we report the 2.03 —resolution crystal structure of the isolated AAA+ module (iAAA+ module) of LonB from Thermococcus onnurineus NA1 (TonLonB). The iAAA+ module, having no bound nucleotide, adopts a conformation virtually identical to the ADP-bound conformation of AAA+ modules in the hexameric structure of TonLonB; this provides insights into the ATP-independent proteolytic activity observed in a LonB protease. Structural comparison of AAA+ modules between LonA and LonB revealed that the AAA+ modules of Lon proteases are separated into two distinct clades depending on their structural features. The AAA+ module of LonB belongs to the -H2 & Ins1 insert clade (HINS clade)- defined for the first time in this study, while the AAA+ module of LonA is a member of the HCLR clade.
Article PDF
Similar content being viewed by others
Avoid common mistakes on your manuscript.
References
Adams, P.D., Grosse-Kunstleve, R.W., Hung, L.W., Ioerger, T.R., McCoy, A.J., Moriarty, N.W., Read, R.J., Sacchettini, J.C., Sauter, N.K., and Terwilliger, T.C. 2002. PHENIX: building new software for automated crystallographic structure determination. Acta Crystallogr. D Biol. Crystallogr. 58, 1948–1954.
Ammelburg, M., Frickey, T., and Lupas, A.N. 2006. Classification of AAA+ proteins. J. Struct. Biol. 156, 2–11.
An, Y.J., Lee, C.R., Supangat, S., Lee, H.S., Lee, J.H., Kang, S.G., and Cha, S.S. 2010. Crystallization and preliminary X-ray crystallographic analysis of Lon from Thermococcus onnurineus NA1. Acta Crystallogr. Sect. F. Struct. Biol. Cryst. Commun. 66, 54–56.
Cha, S.S., An, Y.J., Lee, C.R., Lee, H.S., Kim, Y.G., Kim, S.J., Kwon, K.K., De Donatis, G.M., Lee, J.H., Maurizi, M.R., et al. 2010. Crystal structure of Lon protease: molecular architecture of gated entry to a sequestered degradation chamber. EMBO J. 29, 3520–3530.
Collaborative Computational Project, Number 4. 1994. The CCP4 suite: programs for protein crystallography. Acta Crystallogr. D Biol. Crystallogr. 50, 760–763.
Duman, R.E. and L-we, J. 2010. Crystal structures of Bacillus subtilis Lon protease. J. Mol. Biol. 401, 653–670.
Emsley, P. and Cowtan, K. 2004. Coot: model-building tools for molecular graphics. Acta Crystallogr. D Biol. Crystallogr. 60, 2126–2132.
Erzberger, J.P. and Berger, J.M. 2006. Evolutionary relationships and structural mechanisms of AAA+ proteins. Annu. Rev. Biophys. Biomol. Struct. 35, 93–114.
Fukui, T., Eguchi, T., Atomi, H., and Imanaka, T. 2002. A membrane-bound archaeal Lon protease displays ATP-independent proteolytic activity towards unfolded proteins and ATP-dependent activity for folded proteins. J. Bacteriol. 184, 3689–3698.
Goldberg, A.L. and St John, A.C. 1976. Intracellular protein degradation in mammalian and bacterial cells: Part 2. Annu. Rev. Biochem. 45, 747–803.
Gottesman, S. 1996. Proteases and their targets in Escherichia coli. Annu. Rev. Genet. 30, 465–506.
Gottesman, S. 2003. Proteolysis in bacterial regulatory circuits. Annu. Rev. Cell. Dev. Biol. 19, 565–587.
Hershko, A. and Ciechanover, A. 1992. The ubiquitin system for protein degradation. Annu. Rev. Biochem. 61, 761–807.
Iyer, L.M., Leipe, D.D., Koonin, E.V., and Aravind, L. 2004. Evolutionary history and higher order classification of AAA+ ATPases. J. Struct. Biol. 146, 11–31.
Li, J.K., Liao, J.H., Li, H., Kuo, C.I., Huang, K.F., Yang, L.W., Wu, S.H., and Chang, C.I. 2013. The N-terminal substrate-recognition domain of a LonC protease exhibits structural and functional similarity to cytosolic chaperones. Acta Crystallogr. D. Biol. Crystallogr. 69, 1789–1797.
Liao, J.H., Ihara, K., Kuo, C.I., Huang, K.F., Wakatsuki, S., Wu, S.H., and Chang, C.I. 2013. Structures of an ATP-independent Lon-like protease and its complexes with covalent inhibitors. Acta Crystallogr. D. Biol. Crystallogr. 69, 1395–1402.
Liao, J.H., Kuo, C.I., Huang, Y.Y., Lin, Y.C., Lin, Y.C., Yang, C.Y., Wu, W.L., Chang, W.H., Liaw, Y.C., Lin, L.H., et al. 2012. A Lon-like protease with no ATP-powered unfolding activity. PLoS One 7, e40226.
McClellan, A.J., Tam, S., Kaganovich, D., and Frydman, J. 2005. Protein quality control: chaperones culling corrupt conformations. Nat. Cell Biol. 7, 736–741.
Otwinowski, Z. and Minor, W. 1997. Processing of X-ray diffraction data collected in oscillation mode. Macromolecular Crystallography Pt A 276, 307–326.
Rotanova, T.V., Botos, I., Melnikov, E.E., Rasulova, F., Gustchina, A., Maurizi, M.R., and Wlodawer, A. 2006. Slicing a protease: structural features of the ATP-dependent Lon proteases gleaned from investigations of isolated domains. Protein Sci. 15, 1815–1828.
Sauer, R.T. and Baker, T.A. 2011. AAA+ proteases: ATP-fueled machines of protein destruction. Annu. Rev. Biochem. 80, 587–612.
Sauer, R.T., Bolon, D.N., Burton, B.M., Burton, R.E., Flynn, J.M., Grant, R.A., Hersch, G.L., Joshi, S.A., Kenniston, J.A., Levchenko, I., et al. 2004. Sculpting the proteome with AAA(+) proteases and disassembly machines. Cell 119, 9–18.
Talarico, L.A., Gil, M.A., Yomano, L.P., Ingram, L.O., and Maupin-Furlow, J.A. 2005. Construction and expression of an ethanol production operon in Gram-positive bacteria. Microbiology 151, 4023–4031.
Author information
Authors and Affiliations
Corresponding author
Additional information
These authors contributed equally to this work.
Supplemental material for this article may be found at http://www.springerlink.com/content/120956.
Electronic supplementary material
Rights and permissions
About this article
Cite this article
An, Y.J., Na, JH., Kim, MI. et al. Structural basis for the ATP-independent proteolytic activity of LonB proteases and reclassification of their AAA+ modules. J Microbiol. 53, 711–717 (2015). https://doi.org/10.1007/s12275-015-5417-5
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12275-015-5417-5