Abstract
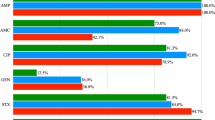
The susceptibility toward antibiotics was determined by disc and MIC methods in Lactobacillus and, for comparison, in Escherichia coli strains isolated from cloacal swabs of broiler chickens derived from various farms in Slovakia. The occurrence of acquired tetracycline resistance in E. coli and lactobacilli isolated from the same sample was similar. The presence of tet(M), tet(S), tet(L) and ermB genes was demonstrated in lactobacilli while the tet(M) gene was not detected in E. coli.
Article PDF
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Abbreviations
- GIT:
-
gastrointestinal tract
- MIC:
-
minimum inhibitory concentration
- MICxG:
-
geometric mean of MIC
References
Ammor M.S., Flórez A.B., van Hoek A.H.A.M., de los Reyes-Gavilán C.G., Aarts H.J.M., Margolles A., Mayo B.: Molecular characterization of intrinsic and acquired antibiotic resistance in lactic acid bacteria and bifidobacteria. J.Mol.Microbiol. Biotechnol.14, 6–15 (2008).
Bernardeau M., Guguen M., Vernoux J.P.: Beneficial lactobacilli in food and feed: long-term use, biodiversity and proposals for specific and realistic safety assessments. FEMS Microbiol.Rev.30, 487–513 (2006).
Bryan A., Shapir N., Sadowsky M.J.: Frequency and distribution of tetracycline resistance genes in genetically diverse, nonselected, and nonclinical Escherichia coli strains isolated from diverse human and animal sources. Appl.Environ.Microbiol.70, 2503–2507 (2004).
CLSI (Clinical Laboratory Standards Institute): Performance standards for antimicrobial disk and dilution. Susceptibility tests for bacteria isolated from animals; approved standard — third edition. CLSI Document M13-A328(8), 1–99 (2008); ISBN 1-56238-659-X.
Danielsen M., Wind A.: Susceptibility of Lactobacillus spp. to antimicrobial agents. Internat.J.Food Microbiol.82, 1–11 (2003).
Delgado S., O’sullivan E., Fitzgerald G., Mayo B.: Subtractive screening for probiotic properties of Lactobacillus species from the human gastrointestinal tract in the search for new probiotics. J.Food Sci.72, M310–M315 (2007).
Demankova D., Kmet V.: Occurrence of tetracycline resistance gene tet(M) in lactobacilli, pp. 3–5 in D. Toth, J. Brindza (Eds): Biological Safety and Agrofoods. (ISBN 978-80-552-0078-1) Slovak Agricultural University, Nitra (Slovakia) 2008.
Dubernet S., Desmasures N., Gueguen M.: A PCR-based method for identification of lactobacilli at the genus level. FEMS Microbiol. Lett.214, 271–275 (2002).
FEEDAP Panel: Opinion of the scientific panel on additives and products or substances used in animal feed on the updating of the criteria used in the assessment of bacteria for resistance to antibiotics of human or veterinary importance. EFSA J.223, 1–12 (2005).
Florez A.B, Ladero V., Alvarez-Martin P., Ammor M.S., Alvarez M.A., Mayo M.: Acquired macrolide resistance in the human intestinal strain Lactobacillus rhamnosus E41 associated with a transition mutation in 23S rRNA genes. Internat.J.Antimicrob. Agents30, 341–344 (2007).
Gattringer R., Niks M., Ostertag R., Schwarz K., Medvedovic H., Graninger W., Georgopoulos A.: Evaluation of Miditech automated colorimetric MIC reading for antimicrobial susceptibility testing. J.Antimicrob.Chemother.49, 651–659 (2002).
Kim H.B, Jang H.C, Nam H.J.: In vitro activities of 28 antimicrobial agents against Staphylococcus aureus isolates from tertiary-care hospitals in Korea: a nationwide survey. Antimicrob.Agents Chemother.48, 1124–1127 (2004).
Lauková A., Marciňáková M., Strompfová V., Ouwehand A.C.: Probiotic potential of enterococci isolated from canine feed. Folia Microbiol.53, 84–88 (2008).
Martineau F., Picard F.J., Grenier L., Roy P.H., Ouellette M., Bergeron M.G.: Multiplex PCR assays for the detection of clinically relevant antibiotic resistance genes in staphylococci isolated from patients infected after cardiac surgery. The ESPRIT Trial. J.Antimicrob.Chemother.46, 527–534 (2000).
Ng L.K., Martin I., Alfa M., Mulvey M: Multiplex PCR for the detection of tetracycline resistant genes. Mol.Cell.Probes15, 209–215 (2001).
Rosander A., Connolly E., Roos S.: Removal of antibiotic resistance gene-carrying plasmids from Lactobacillus reuteri ATCC 55730 and characterization of the resulting daughter strain, L. reuteri DSM 17938. Appl.Environ.Microbiol.74, 6032–6040 (2008).
Tenorio C., Zarazaga M., Martinez C., Torres C.: Bifunctional enzyme 6′-N-aminoglycoside acetyltransferase-2″-O-aminoglycoside phosphotransferase in Lactobacillus and Pediococcus isolates of animal origin. J.Clin.Microbiol.39, 824–825 (2001).
Vakulenko S.B., Donabedian S.M., Voskresenskiy A.M., Zervos M.J., Lerner S.A., Chow J.W.: Multiplex PCR for detection of aminoglycoside resistance genes in enterococci. Antimicrob.Agents Chemother.47, 1423–1426 (2003).
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Kmeť, V., Piatnicová, E. Antibiotic resistance in commensal intestinal microflora. Folia Microbiol 55, 332–335 (2010). https://doi.org/10.1007/s12223-010-0052-3
Received:
Revised:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12223-010-0052-3