Abstract
Recent advances have expanded our ability to conduct a comprehensive genetic evaluation for dilated cardiomyopathy (DCM). By evaluating recent literature, this review aims to bring the reader up-to-date on the genetic evaluation of DCM. Updated guidelines have been published. Mutations in BAG3, including a large deletion, were identified in 2 % of DCM. Truncating mutations in TTN were reported in 25 % of DCM. Two new genes have been reported with autosomal recessive DCM. These studies illustrate the role of improved technologies while raising the possibility of a complex genetic model for DCM. The inclusion of TTN has led to an increased genetic testing detection rate of 40 %. While our ability to identify disease-causing variants has increased, so has the identification of variants of unknown significance. A genetic evaluation for DCM must therefore address this complexity.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Dilated cardiomyopathy (DCM) is one of the most common forms of cardiomyopathy with a prevalence likely exceeding 1/500 [1]. The diagnosis is based on increased left ventricular end diastolic diameter and systolic dysfunction. DCM can be present as an isolated finding or as part of a syndrome (Table 1). This review focuses on the genetic basis of nonsyndromic, isolated DCM. While most commonly evaluated by echocardiography, left ventricular enlargement (LVE) and systolic dysfunction may also be identified by left ventricular angiography or cardiac magnetic resonance imaging. LVE by echocardiography is deemed to be present when the diastolic left ventricular diameter exceeds the 95th percentile of a population-based height and gender-based approach [4]. Left ventricular dysfunction is defined as an ejection fraction of less than 50 % [5] (or more stringently, 45 % [6] or a fractional shortening of less than 25 %).
Other structural cardiac findings, such as right ventricular (RV) enlargement [7], fatty infiltration [8], and left ventricular noncompaction (LVNC) [9, 10] may also be present. DCM with RV involvement, however, must be differentiated from arrhythmogenic right ventricular cardiomyopathy (ARVC), a diagnosis involving right ventricular involvement, fibrofatty infiltration, prominent arrhythmia, and sudden death. LV involvement in ARVC, once thought to be part of the late phase of the condition [11], is also observed in ARVC with early, predominant LV involvement [12]. To account for this wide phenotypic spectrum, the more inclusive term “arrhythmogenic cardiomyopathy” has been proposed [13].
When symptomatic, DCM presents late in the disease course with heart failure (HF), arrhythmia, and conduction system disease (CSD; first, second, or third degree heart block) with or without sudden death, or with thromboembolism [14]. Because DCM is commonly asymptomatic until late in its disease course [15], it may only be identified serendipitously or as part of cardiovascular screening of at-risk family members. Onset usually occurs by age 60 [1], although it has been observed during the fetal stage [16], in infants, children [17•], and the elderly [8]. A subset of DCM, peripartum cardiomyopathy or pregnancy associated cardiomyopathy (PPCM/PACM), has been associated with pregnancy [18], although a causal role for pregnancy has yet to be demonstrated in humans. Features of LVNC have been reported in PPCM, and a familial basis has been suggested based on the transmission of LVNC from a mother with both PPCM and LVNC to her newborn child [19]. A true association between PPCM and LVNC needs to be confirmed with larger studies.
Management includes pharmacologic therapy with ACE inhibitors and beta blockers, and consideration of intracardiac defibrillator/pacemaker (ICD/PCM) for those with a history of or at risk for lethal arrhythmia. Advanced therapies, including left ventricular assist device and heart transplantation, are indicated for patients with severe or intractable disease. Pregnancy is contraindicated in women with DCM or who have been diagnosed with PPCM/PACM.
Etiology
A wide variety of etiologies underlie DCM. In the United States, ischemic disease from coronary artery disease (CAD) is the most common cause. Nonischemic disease follows, which encompasses a wide range of causes, including cardiotoxic drug exposure, endocrine disease, congenital heart defects, infections (viral, bacterial, parasitic, and fungal forms), and genetically-mediated forms. The term idiopathic dilated cardiomyopathy (IDC), coined before evidence of a genetic basis became available, refers to DCM in which all identifiable causes (excluding genetic) have been excluded. It is now known that a portion of IDC also includes genetic forms. For simplicity, and for the remainder of this review, the term DCM will be used for cases with known genetic cause (eg, LMNA-DCM, following a recent nomenclature recommendation)[1] and those in which ischemic, cardiotoxic, and infectious causes have been ruled out (also known as IDC).
Using cardiovascular screening, familial disease (familial dilated cardiomyopathy, FDC) is identified in up to 48 % of IDC [20–22]. In the clinical cardiovascular literature, pedigrees with more than 1 individual with IDC (multiplex pedigrees) are denoted as FDC while all other nonfamilial cases (simplex pedigrees) preserve their IDC categorization. Besides the observable difference in number of affected subjects when familial and simplex pedigrees are compared, the clinical presentation of DCM cases is the same, despite a familial or apparently sporadic nature [23].
Genetics
Mutations in over 30 genes of diverse ontology have been reported in DCM, with more convincing evidence supporting causation in familial cases [15]. Mutations in genes encoding proteins of the desmosome, traditionally associated with ARVC, are also present in DCM [24, 25, 26•], perhaps explaining the mixed ARVC/DCM phenotypes. Most mutations lead to an autosomal dominant pattern of inheritance, however, a minority are associated with recessive [27], X-linked [28], or maternal mitochondrial [29] forms, among which extra-cardiac features and early onset may be more frequent. Autosomal recessive phenotypes, which are more frequent in consanguineous families, have been reported with mutations in TNNI3, and more recently, with GATAD1 (nonsyndromic DCM; Table 2) [30•] and DOLK (predominantly nonsyndromic DCM) [3•] mutations. Penetrance may be incomplete (the proportion of mutation-positive individuals who show the phenotype) and disease expression (the degree of severity among known affected, mutation-positive individuals) is variable. Various disease mechanisms are involved (depending on the mutation involved); however, myocardial injury is consistently the end-result [1]. At the population level, > 200 mutant alleles have been reported in the now >30 known genes [33], all leading to essentially the same clinical picture (allelic heterogeneity).
Missense, frameshift, nonsense, and small insertion/deletions mutations have been reported. The role of larger deletions in DCM was demonstrated in a gene discovery study that for the first time identified mutations in BAG3 (BCL2-associated athanogene 3) as causative of DCM [31•] (Table 2). The study identified a whole exon deletion in BAG3 in a large FDC pedigree and frameshift, nonsense and missense mutations in 2 % of DCM, including familial and simplex cases [31•]. BAG3, a 535 amino acid protein coded by 4 exons, is also implicated in severe dominant childhood muscular dystrophy [34]. The gene product works as a co-chaperone of heat shock proteins. Its binding to proteins related to DCM pathophysiology, such as Bcl-2 (a regulator of apoptosis), is thought to be disrupted in BAG3-DCM [31]. Larger deletions have been observed in EYA4 [35] and LMNA [36], however, a multiplex ligation probe amplification screening of LMNA in 58 probands failed to identify copy number variation [37]. The prevalence of copy number variation in DCM thus remains unknown until more comprehensive studies are performed.
Truncating mutations in TTN (Table 2) have also been identified in approximately 20 % of cases [32•], thus, increasing the proportion of known genetic cause in DCM to 40 %. The clinical presentation of TTN-DCM does not appear to be different than that of other forms of genetic DCM. TTN is a key sarcomeric component that encodes the giant protein titin. It comprises 283 kilobases containing 363 exons coding for 38,138 amino acids [38]. TTN undergoes dramatic alternative splicing to produce many skeletal muscle and heart isoforms [39], and is regulated by RBM20, another gene associated with approximately 2 % of DCM [40–42]. Cardiac-relevant isoforms include N2B, N2BA, and novex-3 [32•]. TTN had been reported with DCM in early studies [43, 44], but the recent study is significant for using next generation sequencing technology to comprehensively evaluate TTN in a larger number of DCM probands (n = 203) with both simplex and familial disease. The study, which suggested a causative role for TTN truncating mutations, led to the immediate addition of this gene to clinical genetic testing panels. However, because the recent study also identified TTN truncating mutations in 3 % of their 249 controls [32•], and because mutations are frequently novel, the complexity of adjudicating causality of genetic DCM variants has escalated to unprecedented levels. Furthermore, the Herman et al. study did not evaluate missense mutations, which are very common in most individuals [45].
Further complicating molecular interpretation is the finding of multiple mutations in DCM. Examples include FDC pedigrees with mutations in LMNA and RBM20 [42], MYH7 and LDB3, and PSEN2, and SCN5A [46]. Multiple mutations in simplex pedigrees have also been observed, including probands with mutations in MYBPC3 and TNNC1, TNNT2 and TPM1, TNNC1 and MYH7, and a fourth one with TCAP, LMNA, and MYH6 mutations [47]. Triple mutations were also seen in a proband with familial disease and 2 mutations in MYH6 and 1 in TNNT2 [47]. Compound heterozygosity was identified in a proband with familial disease and 2 different mutations in MYBPC3 [47]. A homozygous TNNT2 mutation was also observed in a simplex, consanguineous pedigree [46, 48]. It is difficult to sort out the relevance of each individual finding, as many of these mutations were novel while others were previously reported with DCM or other cardiovascular phenotypes. They were also rare in the general population and changed a key amino acid, all usual criteria to establish pathogenicity [15]. Therefore, the role of multiple mutations in DCM may be relevant [49], but has not been formally studied. A possible link between HCM and DCM, however, has been proposed based on multiple mutation findings [50].
Genetic Evaluation
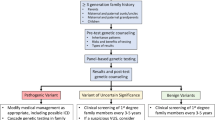
In 2009, guidelines for the genetic evaluation of cardiomyopathy from the Heart Failure Society of America, including DCM, recommended family history, periodic cardiovascular screening of at-risk family members (by echocardiogram, electrocardiogram, and physical exam), and consideration of genetic testing with counseling for individuals with DCM, and, when applicable, their family members [51]. In families in which the disease-causing mutation is known, targeted testing should be offered to at-risk family members. Those who test positive for the family mutation should undergo periodic screening on a yearly basis, while those who test negative for the family mutation may be discharged [51, 52]. The family members of probands in which genetic testing fails to identify a mutation or in which genetic testing has not been performed should undergo screening every 3–5 years [51].
More recent guidelines from the Heart Rhythm Society in conjunction with the European Heart Rhythm Association that included cardiomyopathy were published in 2011, which recommended LMNA and SCN5A genetic testing for individuals with DCM and significant CSD or premature, unexpected sudden cardiac death and targeted testing for at-risk family members when a mutation is identified in a proband [53•]. These guidelines also considered it helpful to offer genetic testing to individuals with FDC for diagnosis confirmation, identification of at-risk family members (including cases when previously undetected syndromic disease may be present), and for family planning. The guidelines also distinguished the clinical value of genetic testing for probands with and without CSD. For the former, genetic testing was deemed to have a greater impact, as genetic information can be useful not only for diagnostic purposes, but also for prognostic and perhaps therapeutic value, especially when mutations in LMNA and DES are present [53•]. For these individuals, prophylactic ICD/PCM may be considered.
Both guidelines, published before the TTN report [32•], do not fully address the genetic complexity of DCM and provide little insight to help navigate complex genetic testing reports in order to make results clinically useful. One of these issues is the likely finding of a variant of unknown significance (VUS). To illustrate this concept, before the recent TTN report, clinical laboratories were quoting a detection rate of approximately 20 %–40 %. However, an analysis of clinical genetic results concluded that disease causing mutations were present in 17.4 % whereas 10.6 % had a VUS [54•]. Many TTN variants, including truncating ones (also identified in 3 % of controls), fall into this category. That a VUS is a disappointing finding, and one that should be handled with care, is not a new concept; the novelty lies in the increased volume of VUSs that reports may now include, especially because all variants in the enormous gene TTN, including missense variants, are reported. Moreover, variants thought to be pathogenic may be reclassified as benign and vice-versa. These issues warrant careful discussion with patients during pre-test informed consent.
Genetic testing for DCM involves large gene panels. For DCM with more prominent than usual right ventricular (RV) involvement, or DCM with a prominent arrhythmogenic phenotype, multicardiomyopathy genetic testing panels including desmosomal genes can help to clarify diagnosis. All inclusive panels evaluating cardiomyopathy, channelopathy, and structural heart disease in a single test are also available, with some now including >50 genes. Careful review of the genes included in these panels is key to better address the evaluation needs of a patient. For example, some panels include mitochondrial genes (that may be more relevant in pediatric or multisystem involvement cases) while others do not. In addition, because gene panels are continually updated in response to gene discoveries or improved technology, a previously done panel may rapidly become outdated. Also, due to the possibility of multiple mutations, a panel may be offered to more than 1 affected family member (or even repeated, with an expanded number of genes, in a patient with a known gene mutation). Similarly, for families with mutations in genes with incomplete evidence for pathogenicity, such as TTN, alerting genotype- and phenotype-negative family members about signs and symptoms of DCM, and suggesting follow-up genetic evaluation later in life may be wise (until fully interpretable whole exome or whole genome testing becomes the standard approach to clinical genetic evaluations). The decision to repeat genetic testing panels or recommend additional clinical cardiovascular care should be exercised with caution, by expert cardiovascular genetics practitioners.
Until these issues are more appropriately addressed on the research arena with comprehensive sequencing of a much larger number of probands (and possibly with functional studies of key genes), clinical cardiovascular screening remains a reliable, yet more expensive, tool to evaluate all at-risk family members. The logical follow-up question would be how willing or able are patients to undergo clinical screening? This question was recently addressed in a retrospective chart review of 57 patients (11 DCM, 46 HCM) that reported an uptake of 57 % and a 25 % detection of familial disease [55•]. In this study, uptake of screening (proportion of at-risk first and second degree relatives who underwent echocardiogram and electrocardiogram) was more likely among first degree relatives of probands, relatives of probands with a known mutation, and in families with multiple affected living relatives. Among relatives of probands with a known mutation, the uptake of cardiac surveillance was higher (59 %) than the uptake of genetic testing (39 %). The authors speculated that the lower uptake of genetic testing may result from, among others factors, perception of high cost and fear of genetic discrimination [55•]. Although the results of this study should be viewed as preliminary due to the small sample size, especially for DCM, they provide a glimpse into the practical aspects of genetic evaluation guidelines for DCM.
To address these issues, providers may periodically re-contact laboratories offering clinical genetic testing for DCM, as cost may no longer be an issue for some patients due to updated customer service policies. Patients should also be informed about the benefits and limitations of the Genetic Information Nondiscrimination Act (GINA) [56]. Discussing that identifying silent disease through cardiac screening may have insurance implications may also help putting things in perspective. Patients may also benefit from visualizing the risk of genetic discrimination presented against the risk of not having a clear diagnosis or not being able to prospectively identify at-risk family members. For the reasons outlined above, informed consent for genetic testing should not only include technical information, but also a discussion to provide anticipatory guidance regarding potential reactions to results, discrimination risks, family communication issues, and cost, among others. For example, before testing begins, patients should be encouraged to picture different scenarios, including positive, negative, or uninformative results, and the possibility of results reclassification. In the setting of a positive result, duty to warn issues should be proactively addressed, so that ethical dilemmas related to results disclosure among family members may be avoided. Genetic counselors are uniquely equipped to assist patients and their families to explore these issues, which have been summarized in the setting of cardiovascular disease [57•].
A genetic evaluation should always begin with a careful and comprehensive phenotype, which includes a detailed, skillfully obtained family history by an informed professional [58, 59]. Current data supports panel genetic testing for the >30 genes in individuals with idiopathic dilated cardiomyopathy, preferably those with FDC. Cardiovascular data should be sought to confirm the phenotype. Similarly, especially given the complexity of genetic testing results, counseling of at-risk family members requires both phenotype (from recent cardiovascular screening) and genotype information. Those with suspected syndromic disease (eg, birth defects, abnormal stature, dysmorphic features, learning disabilities, multisystem involvement) should be evaluated by a clinical geneticist to rule out syndromic disease, for which other forms of genetic testing and management are more appropriate.
Conclusions and Future Directions
The field has reached a critical juncture in which sequencing technology may soon, if it has not already, outpace our ability to synthesize genetic data into clinically useful information. Although exome and whole genome sequencing, along with biomarker and environmental data, are essential to help solve these issues, managing large volumes of information using a multidisciplinary bioinformatics, clinical, and molecular approach will be key. Furthermore, deeper genome expeditions warrant at least equally deeper phenotype assessments to fulfill the promise of genomics and, ultimately, personalized medicine.
References
Papers of particular interest, published recently, have been highlighted as: • Of importance
Hershberger RE, Morales A, Siegfried JD. Clinical and genetic issues in dilated cardiomyopathy: a review for genetics professionals. Gen Med. 2010;12:655–67.
Harakalova M, van Harssel JJ, Terhal PA, van Lieshout S, Duran K, Renkens I, et al. Dominant missense mutations in ABCC9 cause cantu syndrome. Nat Genet. 2012;44:793–6.
Lefeber DJ, de Brouwer AP, Morava E, Riemersma M, Schuurs-Hoeijmakers JH, Absmanner B, et al. Autosomal recessive dilated cardiomyopathy due to DOLK mutations results from abnormal dystroglycan O-mannosylation. PLoS Genet. 2011. DOLK mutations are reported as a cause of primarily non syndromic autosomal recessive DCM in 11 pediatric cases.
Vasan R, Larson MG, Levy D, Evans JC, Benjamin EJ. Distribution and categorization of echocardiographic measurements in relation to reference limits. The Framingham heart study: formulation of a height- and sex-specific classification and its prospective validation. Circulation. 1997;96:1863–73.
Burkett EL, Hershberger RE. Clinical and genetic issues in familial dilated cardiomyopathy. J Am Coll Cardiol. 2005;45:969–81.
Mestroni L, Maisch B, McKenna WJ, Schwartz K, Charron P, Rocco C, et al. Guidelines for the study of familial dilated cardiomyopathies. Eur Heart J. 1999;20:93–102.
Elliott P, Andersson B, Arbustini E, Bilinska Z, Cecchi F, Charron P, et al. Classification of the cardiomyopathies: a position statement from the European society of cardiology working group on myocardial and pericardial diseases. Eur Heart J. 2008;29:270–6.
Morales A, Pinto JR, Siegfried JD, Li D, Norton N, Hofmeyer M, et al. Late onset sporadic dilated cardiomyopathy caused by a cardiac troponin T mutation. Clin Trans Sci. 2010;3:219–26.
Vatta M, Mohapatra B, Jimenez S, Sanchez X, Faulkner G, Perles Z, et al. Mutations in Cypher/ZASP in patients with dilated cardiomyopathy and left ventricular noncompaction. J Am Coll Cardiol. 2003;42:2014–27.
Moller DV, Andersen PS, Hedley P, Ersboll MK, Bundgaard H, Moolman-Smook J, et al. The role of sarcomere gene mutations in patients with idiopathic dilated cardiomyopathy. Eur J Hum Genet. 2009;17:1241–9.
Sen-Chowdhry S, Lowe MD, Sporton SC, McKenna WJ. Arrhythmogenic right ventricular cardiomyopathy: clinical presentation, diagnosis, and management. Am J Med. 2004;117:685–95.
Sen-Chowdhry S, Syrris P, Prasad SK, Hughes SE, Merrifield R, Ward D, et al. Left-dominant arrhythmogenic cardiomyopathy: an under-recognized clinical entity. J Am Coll Cardiol. 2008;52:2175–87.
Rizzo S, Pilichou K, Thiene G, Basso C. The changing spectrum of arrhythmogenic (right ventricular) cardiomyopathy. Cell Tissue Res. 2012;348:319–23.
Maron BJ, Towbin JA, Thiene G, Antzelevitch C, Corrado D, Arnett D, et al. Contemporary definitions and classification of the cardiomyopathies: an American heart association scientific statement from the council on clinical cardiology, heart failure and transplantation committee; quality of care and outcomes research and functional genomics and translational biology interdisciplinary working groups; and council on epidemiology and prevention. Circulation. 2006;113:1807–16.
Hershberger RE, Siegfried JD. State of the art review. Update 2011: clinical and genetic issues in familial dilated cardiomyopathy. J Am Coll Cardiol. 2011;57:1641–9.
Sivasankaran S, Sharland GK, Simpson JM. Dilated cardiomyopathy presenting during fetal life. Cardiol Young. 2005;15:409–16.
Rampersaud E et al. Rare variant mutations identified in pediatric patients with dilated cardiomyopathy. Prog Pediatr Cardiol. 2011;31:39–47. Clinical and molecular genetic data from pediatric DCM cases were analyzed, showing that nonsynonymous rare variants, predominantly in sarcomeric genes, are causative of pediatric DCM.
Elkayam U. Clinical characteristics of peripartum cardiomyopathy in the United States: diagnosis, prognosis, and management. J Am Coll Cardiol. 2011;58:659–70.
Rajagopalan N, Attili AK, Bodiwala K, Bailey AL. Features of left ventricular noncompaction in peripartum cardiomyopathy: a case series. Int J Cardiol. 2012.
Michels M, Hoedemaekers YM, Kofflard MJ, Frohn-Mulder I, Dooijes D, Majoor-Krakauer D, et al. Familial screening and genetic counselling in hypertrophic cardiomyopathy: the Rotterdam experience. Neth Heart J. 2007;15:184–90.
Grunig E, Tasman JA, Kucherer H, Franz W, Kubler W, Katus HA. Frequency and phenotypes of familial dilated cardiomyopathy [see comments]. J Am Coll Cardiol. 1998;31:186–94.
Baig MK, Goldman JH, Caforio AP, Coonar AS, Keeling PJ, McKenna WJ. Familial dilated cardiomyopathy: cardiac abnormalities are common in asymptomatic relatives and may represent early disease. J Am Coll Cardiol. 1998;31:195–201.
Kushner JD, Nauman D, Burgess D, Ludwigsen S, Parks S, Pantely G, et al. Clinical characteristics of 304 kindreds evaluated for familial dilated cardiomyopathy. J Cardiac Failure. 2006;12:422–9.
Elliott P, O'Mahony C, Syrris P, Evans A, Rivera Sorensen C, Sheppard MN, et al. Prevalence of desmosomal protein gene mutations in patients with dilated cardiomyopathy. Circ Cardiovasc Genet. 2010;3:314–22.
Posch MG et al. A missense variant in desmoglein-2 predisposes to dilated cardiomyopathy. Mol Genet Metab. 2008;95:74–80.
Garcia-Pavia P, Syrris P, Salas C, Evans A, Mirelis JG, Cobo-Marcos M, et al. Desmosomal protein gene mutations in patients with idiopathic dilated cardiomyopathy undergoing cardiac transplantation: a clinicopathological study. Heart. 2011;97:1744. Mutations in desmosomal genes were identified in patients with advanced DCM, suggesting that the addition of these genes may improve the yield of genetic testing for DCM.
Murphy RT, Mogensen J, Shaw A, Kubo T, Hughes S, McKenna WJ. Novel mutation in cardiac troponin I in recessive idiopathic dilated cardiomyopathy. Lancet. 2004;363:371–2.
Towbin JA, Hejtmancik JF, Brink P, Gelb B, Zhu XM, Chamberlain JS, et al. X-linked dilated cardiomyopathy. Circulation. 1993;87:1854–65.
Zaragoza MV, Brandon MC, Diegoli M, Arbustini E, Wallace DC. Mitochondrial cardiomyopathies: how to identify candidate pathogenic mutations by mitochondrial DNA sequencing, MITOMASTER and phylogeny. Eur J Hum Genet. 2011;19:200–7.
Theis JL, Sharpe KM, Matsumoto ME, Chai HS, Nair AA, Theis JD, et al. Homozygosity mapping and exome sequencing reveal GATAD1 mutation in autosomal recessive dilated cardiomyopathy. Circ Cardiovasc Genet. 2011;4:585–94. Using exome sequencing and homozygosity mapping in a consanguineous family, a homozygous variant in a novel gene associated with autosomal recessive DCM was identified. The gene product, GATA zinc finger domain-containing protein 1, is relevant for gene expression.
Norton N, Li D, Reider MJ, Siegfried JD, Rampersaud E, Zuchner S, et al. Genome-wide studies of copy number variation and exome sequencing identify rare variants in BAG3 as a cause of dilated cardiomyopathy. Am J Hum Genet. 2011;88:273–82. Variants in a novel gene, including a large deletion, associated with autosomal dominant, non syndromic DCM are reported for the first time.
Herman DS, Lam L, Taylor MR, Wang L, Teekakirikul P, Christodoulou D, et al. Truncations of titin causing dilated cardiomyopathy. N Engl JMed. 2012;366:619–28. Comprehensive evaluation of TTN suggesting that truncating mutations cause 25% of DCM.
Norton N, Robertson PD, Rieder MJ, Zuchner S, Rampersaud E, Martin E, et al. Evaluating pathogenicity of rare variants from dilated cardiomyopathy in the Exome era. Circ Cardiovasc Genet. 2012;5:167–74.
Selcen D, Muntoni F, Burton BK, Pegoraro E, Sewry C, Bite AV, et al. Mutation in BAG3 causes severe dominant childhood muscular dystrophy. Ann Neurol. 2009;65:83–9.
Schonberger J, Wang L, Shin JT, Kim SD, Depreux FF, Zhu H, et al. Mutation in the transcriptional coactivator EYA4 causes dilated cardiomyopathy and sensorineural hearing loss. Nat Genet. 2005;37:418–22.
Gupta P, Bilinska ZT, Sylvius N, Boudreau E, Veinot JP, Labib S, et al. Genetic and ultrastructural studies in dilated cardiomyopathy patients: a large deletion in the lamin A/C gene is associated with cardiomyocyte nuclear envelope disruption. Basic Res Cardiol. 2010;105:365–77.
Norton N, Siegfried JD, Li D, Hershberger RE. Assessment of LMNA copy number variation in 58 probands with dilated cardiomyopathy. Clin Transl Sci. 2011;4:351–2.
Bang ML, Centner T, Fornoff F, Geach AJ, Gotthardt M, McNabb M, et al. The complete gene sequence of titin, expression of an unusual approximately 700-kDa titin isoform, and its interaction with obscurin identify a novel Z-line to I-band linking system. Circ Res. 2001;89:1065–72.
Guo W, Bharmal SJ, Esbona K, Greaser ML. Titin diversity—alternative splicing gone wild. J Biomed Biotechnol. 2010;2010:753675.
Guo W, Schafer S, Greaser ML, Radke MH, Liss M, Govindarajan T, et al. RBM20, a gene for hereditary cardiomyopathy, regulates titin splicing. Nat Med. 2012;18:766–73.
Brauch KM, Karst ML, Herron KJ, de Andrade M, Pellikka PA, Rodeheffer RJ, et al. Mutations in ribonucleic acid binding protein gene cause familial dilated cardiomyopathy. J Am Coll Cardiol. 2009;54:930–41.
Li D, Morales A, Gonzalez Quintana J, Norton N, Siegfried JD, Hofmeyer M, et al. Identification of novel mutations In RBM20 in patients with dilated cardiomyopathy. Clin Trans Sci. 2010;3:90–7.
Gerull B, Gramlich M, Atherton J, McNabb M, Trombitas K, Sasse-Klaassen S, et al. Mutations of TTN, encoding the giant muscle filament titin, cause familial dilated cardiomyopathy. Nat Genet. 2002;14:14.
Itoh-Satoh M, Hayashi T, Nishi H, Koga Y, Arimura T, Koyanagi T, et al. Titin mutations as the molecular basis for dilated cardiomyopathy. Biochem Biophys Res Commun. 2002;291:385–93.
Norton N, Li D, Rampersaud E, Morales A, Martin ER, Zuchner S, et al. Exome sequencing and genome-wide linkage analysis in 17 families demonstrates the complex contribution of TTN truncating variants to dilated cardiomyopathy. Circ Cardiovasc Genet. 2013;6:144–53.
Hershberger RE, Parks SB, Kushner JD, Li D, Ludwigsen S, Jakobs PM, et al. Coding sequence mutations identified in MYH7, TNNT2, SCN5A, CSRP3, LBD3, and TCAP from 313 patients with familial or idiopathic dilated cardiomyopathy. Clin Transl Sci. 2008;1:21–6.
Hershberger RE, Norton N, Morales A, Li D, Siegfried JD, Gonzalez-Quintana J. Coding sequence rare variants identified in MYBPC3, MYH6, TPM1, TNNC1, and TNNI3 from 312 patients with familial or idiopathic dilated cardiomyopathy. Circ Cardiovasc Genet. 2010;3:155–61.
Hershberger RE, Pinto JR, Parks SB, Kushner JD, Li D, Ludwigsen S, et al. Clinical and functional characterization of TNNT2 mutations identified in patients with dilated cardiomyopathy. Circ Cardiovasc Genet. 2009;2:306–13.
Hershberger RE. A glimpse into multigene rare variant genetics: triple mutations in hypertrophic cardiomyopathy. J Am Coll Cardiol. 2010;55:1454–5.
Kelly M, Semsarian C. Multiple mutations in genetic cardiovascular disease: a marker of disease severity? Circ Cardiovasc Genet. 2009;2:182–90.
Hershberger RE, Lindenfeld J, Mestroni L, Seidman CE, Taylor MR, Towbin JA. Genetic evaluation of cardiomyopathy—a heart failure society of America practice guideline. J Card Fail. 2009;15:83–97.
Hershberger RE, Cowan J, Morales A, Siegfried JD. Progress with genetic cardiomyopathies: screening, counseling, and testing in dilated, hypertrophic, and arrhythmogenic right ventricular dysplasia/cardiomyopathy. Circ Heart Fail. 2009;2:253–61.
Ackerman MJ, Priori SG, Willems S, Berul C, Brugada R, Calkins H, et al. HRS/EHRA expert consensus statement on the state of genetic testing for the channelopathies and cardiomyopathies. This document was developed as a partnership between the Heart Rhythm Society (HRS) and the European Heart Rhythm Association (EHRA). Hear Rhythm. 2011;8:1308–39. Recent clinical guidelines highlighting the value of genetic testing for DCM for diagnostic and prognostic purposes in some cases.
Lakdawala NK, Funke BH, Baxter S, Cirino AL, Roberts AE, Judge DP, et al. Genetic testing for dilated cardiomyopathy in clinical practice. J Card Fail. 2012;18:296–303. Examination of the role of genetic testing for DCM in clinical practice, including a quantification of variants of unknown significance.
Miller EM, Wang Y, Ware SM. Uptake of cardiac screening and genetic testing among hypertrophic and dilated cardiomyopathy families. J Genet Couns. 2012. Chart review of HCM and DCM cases exploring the implementation of cardiac screening and genetic testing guidelines.
Genetic information nondiscrimination act (GINA) of 2008. 2010 [cited 2010 December 17]. A brief history of GINA, with information for researchers and health care professionals. ]. Available from: http://www.genome.gov/24519851.
Aatre RD, Day SM. Psychological issues in genetic testing for inherited cardiovascular diseases. Circ Cardiovasc Genet. 2011;4:81–90. Discussion of psychosocial issues in cardiovascular genetic counseling, including proposed guidelines for counseling and testing in inherited cardiovascular disease.
Morales A, Cowan J, Dagua J, Hershberger RE. Family history: an essential tool for cardiovascular genetic medicine. Congest Heart Fail. 2008;14:37–45.
Nauman D, Morales A, Cowan J, Dagua J, Hershberger RE. The family history as a tool to identify patients at risk for dilated cardiomyopathy. Prog Cardiovasc Nurs. 2008;23:41–4.
Compliance with Ethics Guidelines
ᅟ
Conflict of Interest
Ana Morales declares that she has no conflict of interest. Ray E. Hershberger declares that he has no conflict of interest.
Human and Animal Rights and Informed Consent
This article does not contain any studies with human or animal subjects performed by any of the authors.
Author information
Authors and Affiliations
Corresponding author
Additional information
This article is part of the Topical Collection on Congestive Heart Failure
Rights and permissions
About this article
Cite this article
Morales, A., Hershberger, R.E. Genetic Evaluation of Dilated Cardiomyopathy. Curr Cardiol Rep 15, 375 (2013). https://doi.org/10.1007/s11886-013-0375-1
Published:
DOI: https://doi.org/10.1007/s11886-013-0375-1