Abstract
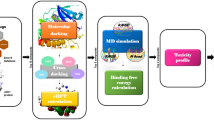
Breast cancer is the most common cause for women’s deaths worldwide. LMTK3 has been demonstrated as critical biomarker for ERα positive breast cancer. It regulates breast cancer by phosphorylating estrogen receptor. Association of LMTK3 in breast cancer is connected with disease free and poor overall survival. In this current computational study, virtual screening was accomplished on human LMTK3 using a large library of NCI database in Schrodinger. From the ligand library, the best compounds were selected and evaluated based on molecular docking using Glide module and their relative molecular dynamics using Desmond. Different parameters like binding energy and interactions like hydrogen bond and hydrophobic contacts have a significant impact on LMTK3 inhibition. Based on docking score, the best lead molecules were separated and analyzed for ADME properties using QikProp tool. Overall, our results confirmed the compounds NCI26194 had been screened from the NCI database, which has the potential to act as key drug molecule for ERα positive breast cancer. In conclusion, our computer aided technique on human LMTK3 has high perspective for the development of novel anticancer agent for breast cancer treatment.
Article PDF
Similar content being viewed by others
Avoid common mistakes on your manuscript.
References
D. R. Robinson, Y. M. Wu and S. F. Lin, Oncogene, 19, 5548 (2000).
G. Giamas, A. Filipovic, J. Jacob, W. Messier, H. Zhang, D. Yang, W. Zhang, B. A. Shifa, A. Photiou, C. Tralau-Stewart and L. Castellano, Nat. Med., 17, 715 (2011).
A. B. Johnson and B. W. O’Malley, Nat. Med., 17, 660 (2011).
Y. Xu, H. Zhang and G. Giamas, Onco Target, 5, 5192 (2014).
J. Harris, M. E. Lippman, U. Veronesi and W. Willet, N Engl. J. Med., 327, 319 (1992).
F. Labrie, C. Labrie, A. Bélanger, J. Simard, S. Gauthier, V. Luu-The, Y. Giguere, B. Candas, S. Luo and C. Martel, J. Steroid Biochem., 69, 51 (1999).
J. Stebbing, A. Filipovic and G. Giamas, Oncotarget, 2, 428 (2011).
R. C. Wu, J. Qin, P. Yi, J. Wong, S. Y. Tsai, M. J. Tsai and B. W. O’Malley, Mol. Cell., 15, 937 (2004).
S. Ali and R. C. Coombes, Nat. Rev. Cancer, 2, 101 (2002).
J. Jiang, N. Sarwar, D. Peston, E. Kulinskaya, S. Shousha, R. C. Coombes and S. Ali, Clin. Cancer Res., 13, 5769 (2007).
G. Giamas, L. Castellano, Q. Feng, U. Knippschild, J. Jacob, R. S. Thomas, R. C. Commbes, C. L. Smith, L. R. Jiao and J. Stebbing, Nucleic Acids Res., 37, 3110 (2009).
G. Giamas, J. Stebbing, C. E. Vorgias and U. Knippschild, Pharmacogenomics, 8, 1005 (2007).
A. B. Johnson and B. W. O’Malley, Nat. Med., 17, 660 (2011).
V. Vella, G. Giamas and A. Ditsiou, Cancer Gene Ther., 24, 1 (2021).
C. Cilibrasi, A. Ditsiou, A. Papakyriakou, G. Mavridis, M. Eravci, J. Stebbing, T. Gagliano and G. Giamas, Mol. Cancer, 20(53), 1 (2021).
K. Anbarasu and S. Jayanthi, Mol. Biosyst., 10, 1139 (2014).
K. Anbarasu and S. Jayanthi, J. Recept. Signal Transduct, 37, 51 (2017).
A. Lin, Z. Cai, G. Hu and Q. Li, J. Recept. Signal Transduct, 35, 559 (2015).
M. Rambabu and S. Jayanthi, J. Cell. Biochem., 120, 8588 (2019).
Schrodinger Suite 2012: Protein Preparation Wizard. Schrodinger, LLC; New York, NY.
Schrodinger Suite. Schrodinger, LLC; New York, NY (2012).
G. M. Sastry, M. Adzhigirey, T. Day, R. Annabhimoju and W. Sherman, J. Comput Aided Mol. Des., 27, 221 (2013).
K. M. Elokely and R. J. Doerksen, J. Chem. Inf. Model., 53, 1934 (2013).
Schrodinger Suite 2012: LigPrep, version 2.5. Schrodinger, LLC; New York, NY.
J. M. Hayes, M. Stein and J. Weiser, J. Phys. Chem. A, 108, 3572 (2004).
Glide version 6.1 [software]. New York: Schrödinger LLC (2013).
R. A. Friesner, R. B. Murphy, M. P. Repasky, L. L. Frye, J. R. Greenwood, T. A. Halgren, P. C. Sanschagrin and D. T. Mainz, J. Med. Chem., 49, 6177 (2006).
Y. Shan, E. T. Kim, M. P. Eastwood, R. O. Dror, M. A. Seeliger and D. E. Shaw, J. Am. Chem. Soc., 133, 9181 (2011).
S. Vilar, J. Karpiak, B. Berk and S. Costanzi, J. Mol. Graph., 29, 809 (2011).
J. P. Ryckaert, G. Ciccotti and H. J. Berendsen, J. Comput. Phys., 23, 327 (1977).
U. Essmann, L. Perera, M. L. Berkowitz, T. Darden, H. Lee and L. G. Pedersen, J. Chem. Phys., 103, 8577 (1995).
W. Shinoda and M. Mikami, J. Comput. Chem., 24, 920 (2003).
S. Nosé, J. Chem. Phys., 81, 511 (1984).
W. Shinoda and M. Mikami, J. Comput. Chem., 24, 920 (2003).
QikProp (Version 3.8). New York, NY: Schrodinger, LLC (2013).
C. A. Lipinski, F. Lombardo, B. W. Dominy and P. J. Feeney, Adv. Drug. Deliv. Rev., 23, 3 (1997).
M. Rambabu and S. Jayanthi, J. Recept Signal Transduct, 40, 436 (2020).
W. L. Jorgensen and E. M. Duffy, Adv. Drug. Deliv. Rev., 54, 355 (2002).
Acknowledgements
This work was supported by the National Research Foundation of Korea (NRF) grant funded by the Korea government (MSIT) (No. 2021R1A2C2006888). The authors thank Schrodinger for facility and management of Saveetha School of Engineering, SIMATS.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
The authors declare that there are no conflicts of interest.
Additional information
Supporting Information
Additional information as noted in the text. This information is available via the Internet at http://www.springer.com/chemistry/journal/11814.
Rights and permissions
About this article
Cite this article
Krishnan, A., Dhamodharan, D., Sundaram, T. et al. Computational discovery of novel human LMTK3 inhibitors by high throughput virtual screening using NCI database. Korean J. Chem. Eng. 39, 1368–1374 (2022). https://doi.org/10.1007/s11814-022-1120-5
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11814-022-1120-5