Abstract
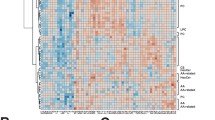
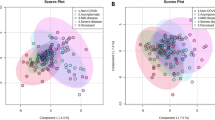
Coronavirus disease 2019 (COVID-19), caused by severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2), is characterized by a strong production of inflammatory cytokines such as TNF and IL-6, which underlie the severity of the disease. However, the molecular mechanisms responsible for such a strong immune response remains unclear. Here, utilizing targeted tandem mass spectrometry to analyze serum metabolome and lipidome in COVID-19 patients at different temporal stages, we identified that 611 metabolites (of 1,039) were significantly altered in COVID-19 patients. Among them, two metabolites, agmatine and putrescine, were prominently elevated in the serum of patients; and 2-quinolinecarboxylate was changed in a biphasic manner, elevated during early COVID-19 infection but levelled off. When tested in mouse embryonic fibroblasts (MEFs) and macrophages, these 3 metabolites were found to activate the NF-κB pathway that plays a pivotal role in governing cytokine production. Importantly, these metabolites were each able to cause strong increase of TNF and IL-6 levels when administered to wildtype mice, but not in the mice lacking NF-κB. Intriguingly, these metabolites have little effects on the activation of interferon regulatory factors (IRFs) for the production of type I interferons (IFNs) for antiviral defenses. These data suggest that circulating metabolites resulting from COVID-19 infection may act as effectors to elicit the peculiar systemic inflammatory responses, exhibiting severely strong proinflammatory cytokine production with limited induction of the interferons. Our study may provide a rationale for development of drugs to alleviate inflammation in COVID-19 patients.
Article PDF
Similar content being viewed by others
Avoid common mistakes on your manuscript.
References
Arunachalam, P.S., Wimmers, F., Mok, C.K.P., Perera, R.A.P.M., Scott, M., Hagan, T., Sigal, N., Feng, Y., Bristow, L., Tak-Yin Tsang, O., et al. (2020). Systems biological assessment of immunity to mild versus severe COVID-19 infection in humans. Science 369, 1210–1220.
Banerjee, A.K., Blanco, M.R., Bruce, E.A., Honson, D.D., Chen, L.M., Chow, A., Bhat, P., Ollikainen, N., Quinodoz, S.A., Loney, C., et al. (2020). SARS-CoV-2 disrupts splicing, translation, and protein trafficking to suppress host defenses. Cell 183, 1325–1339.e21.
Berry, C.T., May, M.J., and Freedman, B.D. (2018). STIM- and Orai-mediated calcium entry controls NF-κB activity and function in lymphocytes. Cell Calcium 74, 131–143.
Blanco-Melo, D., Nilsson-Payant, B.E., Liu, W.C., Uhl, S., Hoagland, D., Møller, R., Jordan, T.X., Oishi, K., Panis, M., Sachs, D., et al. (2020). Imbalanced host response to SARS-CoV-2 drives development of COVID-19. Cell 181, 1036–1045.e9.
Chen, N., Zhou, M., Dong, X., Qu, J., Gong, F., Han, Y., Qiu, Y., Wang, J., Liu, Y., Wei, Y., et al. (2020). Epidemiological and clinical characteristics of 99 cases of 2019 novel coronavirus pneumonia in Wuhan, China: a descriptive study. Lancet 395, 507–513.
Cheng, M.X., Cao, D., Chen, Y., Li, J.Z., Tu, B., and Gong, J.P. (2019). α-ketoglutarate attenuates ischemia-reperfusion injury of liver graft in rats. Biomed pharmacother 111, 1141–1146.
Del Valle, D.M., Kim-Schulze, S., Huang, H.H., Beckmann, N.D., Nirenberg, S., Wang, B., Lavin, Y., Swartz, T.H., Madduri, D., Stock, A., et al. (2020). An inflammatory cytokine signature predicts COVID-19 severity and survival. Nat Med 26, 1636–1643.
Gassen, N.C., Papies, J., Bajaj, T., Emanuel, J., Dethloff, F., Chua, R.L., Trimpert, J., Heinemann, N., Niemeyer, C., Weege, F., et al. (2021). SARS-CoV-2-mediated dysregulation of metabolism and autophagy uncovers host-targeting antivirals. Nat Commun 12, 3818.
Gordon, D.E., Jang, G.M., Bouhaddou, M., Xu, J., Obernier, K., White, K. M., O’Meara, M.J., Rezelj, V.V., Guo, J.Z., Swaney, D.L., et al. (2020). A SARS-CoV-2 protein interaction map reveals targets for drug repurposing. Nature 583, 459–468.
Hadjadj, J., Yatim, N., Barnabei, L., Corneau, A., Boussier, J., Smith, N., Péré, H., Charbit, B., Bondet, V., Chenevier-Gobeaux, C., et al. (2020). Impaired type I interferon activity and inflammatory responses in severe COVID-19 patients. Science 369, 718–724.
Huang, C., Wang, Y., Li, X., Ren, L., Zhao, J., Hu, Y., Zhang, L., Fan, G., Xu, J., Gu, X., et al. (2020a). Clinical features of patients infected with 2019 novel coronavirus in Wuhan, China. Lancet 395, 497–506.
Huang, M., Zhang, W., Chen, H., and Zeng, J. (2020b). Targeting polyamine metabolism for control of human viral diseases. Infect Drug Resist Volume 13, 4335–4346.
Huang, W., Li, C., Wang, Z., Wang, H., Zhou, N., Jiang, J., Ni, L., Zhang, X.A., and Wang, D.W. (2020c). Decreased serum albumin level indicates poor prognosis of COVID-19 patients: hepatic injury analysis from 2,623 hospitalized cases. Sci China Life Sci 63, 1678–1687.
Kanters, E., Gijbels, M.J.J., van der Made, I., Vergouwe, M.N., Heeringa, P., Kraal, G., Hofker, M.H., and de Winther, M.P.J. (2004). Hematopoietic NF-κB1 deficiency results in small atherosclerotic lesions with an inflammatory phenotype. Blood 103, 934–940.
Karki, R., Sharma, B.R., Tuladhar, S., Williams, E.P., Zalduondo, L., Samir, P., Zheng, M., Sundaram, B., Banoth, B., Malireddi, R.K.S., et al. (2021). Synergism of TNF-α and IFN-γ triggers inflammatory cell death, tissue damage, and mortality in SARS-CoV-2 infection and cytokine shock syndromes. Cell 184, 149–168.e17.
Konno, Y., Kimura, I., Uriu, K., Fukushi, M., Irie, T., Koyanagi, Y., Sauter, D., Gifford, R.J., Nakagawa, S., and Sato, K. (2020). SARS-CoV-2 ORF3b is a potent interferon antagonist whose activity is increased by a naturally occurring elongation variant. Cell Rep 32, 108185.
Laing, A.G., Lorenc, A., Del Molino Del Barrio, I., Das, A., Fish, M., Monin, L., Muñoz-Ruiz, M., McKenzie, D.R., Hayday, T.S., Francos-Quijorna, I., et al. (2020). A dynamic COVID-19 immune signature includes associations with poor prognosis. Nat Med 26, 1623–1635.
Lam, S.M., Zhang, C., Wang, Z., Ni, Z., Zhang, S., Yang, S., Huang, X., Mo, L., Li, J., Lee, B., et al. (2021). A multi-omics investigation of the composition and function of extracellular vesicles along the temporal trajectory of COVID-19. Nat Metab 3, 909–922.
Lei, X., Dong, X., Ma, R., Wang, W., Xiao, X., Tian, Z., Wang, C., Wang, Y., Li, L., Ren, L., et al. (2020). Activation and evasion of type I interferon responses by SARS-CoV-2. Nat Commun 11, 3810.
Li, J., Liu, Y., and Zhang, X. (2010). Murine coronavirus induces type I interferon in oligodendrocytes through recognition by RIG-I and MDA5. J Virol 84, 6472–6482.
Liu, C., Martins, A.J., Lau, W.W., Rachmaninoff, N., Chen, J., Imberti, L., Mostaghimi, D., Fink, D.L., Burbelo, P.D., Dobbs, K., et al. (2021). Time-resolved systems immunology reveals a late juncture linked to fatal COVID-19. Cell 184, 1836–1857.e22.
Liu, Y., Yang, Y., Zhang, C., Huang, F., Wang, F., Yuan, J., Wang, Z., Li, J., Li, J., Feng, C., et al. (2020). Clinical and biochemical indexes from 2019-nCoV infected patients linked to viral loads and lung injury. Sci China Life Sci 63, 364–374.
Long, Q.X., Tang, X.J., Shi, Q.L., Li, Q., Deng, H.J., Yuan, J., Hu, J.L., Xu, W., Zhang, Y., Lv, F.J., et al. (2020). Clinical and immunological assessment of asymptomatic SARS-CoV-2 infections. Nat Med 26, 1200–1204.
Lu, J., Lam, S.M., Wan, Q., Shi, L., Huo, Y., Chen, L., Tang, X., Li, B., Wu, X., Peng, K., et al. (2019). High-coverage targeted lipidomics reveals novel serum lipid predictors and lipid pathway dysregulation antecedent to type 2 diabetes onset in normoglycemic Chinese adults. Diabetes Care 42, 2117–2126.
Lucas, C., Wong, P., Klein, J., Castro, T.B.R., Silva, J., Sundaram, M., Ellingson, M.K., Mao, T., Oh, J.E., Israelow, B., et al. (2020). Longitudinal analyses reveal immunological misfiring in severe COVID-19. Nature 584, 463–469.
Mazaleuskaya, L., Veltrop, R., Ikpeze, N., Martin-Garcia, J., and Navas-Martin, S. (2012). Protective role of Toll-like Receptor 3-induced type I interferon in murine coronavirus infection of macrophages. Viruses 4, 901–923.
Mills, E.L., Kelly, B., Logan, A., Costa, A.S.H., Varma, M., Bryant, C.E., Tourlomousis, P., Däbritz, J.H.M., Gottlieb, E., Latorre, I., et al. (2016). Succinate dehydrogenase supports metabolic repurposing of mitochondria to drive inflammatory macrophages. Cell 167, 457–470. e13.
Mulchandani, R., Lyngdoh, T., and Kakkar, A.K. (2021). Deciphering the COVID-19 cytokine storm: systematic review and meta-analysis. Eur J Clin Invest 51, e13429.
Quinn, S.J., Ye, C.P., Diaz, R., Kifor, O., Bai, M., Vassilev, P., and Brown, E. (1997). The Ca2+-sensing receptor: a target for polyamines. Am J Physiol Cell Physiol 273, C1315–C1323.
Rodriguez, A.E., Ducker, G.S., Billingham, L.K., Martinez, C.A., Mainolfi, N., Suri, V., Friedman, A., Manfredi, M.G., Weinberg, S.E., Rabinowitz, J.D., et al. (2019). Serine metabolism supports macrophage IL-1β production. Cell Metab 29, 1003–1011.e4.
Rogers, A.C., McDermott, F.D., Mohan, H.M., O’Connell, P.R., Winter, D. C., and Baird, A.W. (2015). The effects of polyamines on human colonic mucosal function. Eur J Pharmacol 764, 157–163.
Rosas, I.O., Bräu, N., Waters, M., Go, R.C., Hunter, B.D., Bhagani, S., Skiest, D., Aziz, M.S., Cooper, N., Douglas, I.S., et al. (2021). Tocilizumab in hospitalized patients with severe COVID-19 pneumonia. N Engl J Med 384, 1503–1516.
Sa Ribero, M., Jouvenet, N., Dreux, M., and Nisole, S. (2020). Interplay between SARS-CoV-2 and the type I interferon response. PLoS Pathog 16, e1008737.
Salama, C., Han, J., Yau, L., Reiss, W.G., Kramer, B., Neidhart, J.D., Criner, G.J., Kaplan-Lewis, E., Baden, R., Pandit, L., et al. (2021). Tocilizumab in patients hospitalized with COVID-19 pneumonia. N Engl J Med 384, 20–30.
Samuvel, D.J., Sundararaj, K.P., Nareika, A., Lopes-Virella, M.F., and Huang, Y. (2009). Lactate boosts TLR4 signaling and NF-κB pathway-mediated gene transcription in macrophages via monocarboxylate transporters and MD-2 up-regulation. J Immunol 182, 2476–2484.
Schultze, J.L., and Aschenbrenner, A.C. (2021). COVID-19 and the human innate immune system. Cell 184, 1671–1692.
Sette, A., and Crotty, S. (2021). Adaptive immunity to SARS-CoV-2 and COVID-19. Cell 184, 861–880.
Sha, W.C., Liou, H.C., Tuomanen, E.I., and Baltimore, D. (1995). Targeted disruption of the p50 subunit of NF-κB leads to multifocal defects in immune responses. Cell 80, 321–330.
Shen, B., Yi, X., Sun, Y., Bi, X., Du, J., Zhang, C., Quan, S., Zhang, F., Sun, R., Qian, L., et al. (2020). Proteomic and metabolomic characterization of COVID-19 patient sera. Cell 182, 59–72.e15.
Shui, G., Guan, X.L., Low, C.P., Chua, G.H., Goh, J.S.Y., Yang, H., and Wenk, M.R. (2010). Toward one step analysis of cellular lipidomes using liquid chromatography coupled with mass spectrometry: application to Saccharomyces cerevisiae and Schizosaccharomyces pombe lipidomics. Mol Biosyst 6, 1008–1017.
Siehler, S., Wang, Y., Fan, X., Windh, R.T., and Manning, D.R. (2001). Sphingosine 1-phosphate activates nuclear factor-κB through Edg receptors. J Biol Chem 276, 48733–48739.
Song, J.W., Lam, S.M., Fan, X., Cao, W.J., Wang, S.Y., Tian, H., Chua, G. H., Zhang, C., Meng, F.P., Xu, Z., et al. (2020). Omics-driven systems interrogation of metabolic dysregulation in COVID-19 pathogenesis. Cell Metab 32, 188–202.e5.
Thomas, T., Stefanoni, D., Reisz, J.A., Nemkov, T., Bertolone, L., Francis, R.O., Hudson, K.E., Zimring, J.C., Hansen, K.C., Hod, E.A., et al. (2020). COVID-19 infection alters kynurenine and fatty acid metabolism, correlating with IL-6 levels and renal status. JCI Insight 5.
Waggershauser, C.H., Tillack-Schreiber, C., Berchtold-Benchieb, C., Szokodi, D., Howaldt, S., and Ochsenkühn, T. (2020). Letter: immunotherapy in IBD patients in a SARS-CoV-2 endemic area. Aliment Pharmacol Ther 52, 898–899.
Wang, D., Hu, B., Hu, C., Zhu, F., Liu, X., Zhang, J., Wang, B., Xiang, H., Cheng, Z., Xiong, Y., et al. (2020a). Clinical characteristics of 138 hospitalized patients with 2019 novel coronavirus-infected pneumonia in Wuhan, China. JAMA 323, 1061–1069.
Wang, J., Jiang, M., Chen, X., and Montaner, L.J. (2020b). Cytokine storm and leukocyte changes in mild versus severe SARS-CoV-2 infection: review of 3939 COVID-19 patients in China and emerging pathogenesis and therapy concepts. J Leukoc Biol 108, 17–41.
Wu, P., Chen, D., Ding, W., Wu, P., Hou, H., Bai, Y., Zhou, Y., Li, K., Xiang, S., Liu, P., et al. (2021). The trans-omics landscape of COVID-19. Nat Commun 12, 4543.
Xia, H., Cao, Z., Xie, X., Zhang, X., Chen, J.Y.C., Wang, H., Menachery, V. D., Rajsbaum, R., and Shi, P.Y. (2020). Evasion of type I interferon by SARS-CoV-2. Cell Rep 33, 108234.
Xiong, Y., Liu, Y., Cao, L., Wang, D., Guo, M., Jiang, A., Guo, D., Hu, W., Yang, J., Tang, Z., et al. (2020). Transcriptomic characteristics of bronchoalveolar lavage fluid and peripheral blood mononuclear cells in COVID-19 patients. Emerg Microbes Infect 9, 761–770.
Zhang, Q., Lenardo, M.J., and Baltimore, D. (2017). 30 Years of NF-κB: a blossoming of relevance to human pathobiology. Cell 168, 37–57.
Zheng, M., Karki, R., Williams, E.P., Yang, D., Fitzpatrick, E., Vogel, P., Jonsson, C.B., and Kanneganti, T.D. (2021). TLR2 senses the SARS-CoV-2 envelope protein to produce inflammatory cytokines. Nat Immunol 22, 829–838.
Acknowledgements
This work was supported by the National Natural Science Foundation of China (31922034, 82088102, 91854208, 92057204, 31730058, 82072777), Science and Technology Program of Fujian Provincial Health Commission (2021ZD02006), Xiamen Science and Technology Major Project (3502Z2020YJ05), Xiamen Municipal Bureau of Science and Technology (3502Z20209005), Fundamental Research Funds for the Central Universities (20720200014, 20720200069, 20720190084), and Program of Introducing Talents of Discipline to Universities (BP2018017). We acknowledge all health care workers involved in the diagnosis and treatment of patients in the First Affiliated Hospital of Xiamen University, and all the patients, supporters, and their families for their confidence in our work. We also thank Dr. Menno de Winther (Academic Medical Center, Universiteit Utrecht) for providing p105/p50-/- mice (Jackson Laboratory, #006097).
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Compliance and ethics The author(s) declare that they have no conflict of interest.
Electronic supplementary material
11427_2021_2099_MOESM3_ESM.xlsx
Table S2 Full list of metabolites identified in this study. See Sheets 1 and 2 for polar metabolites, and Sheets 3 and 4 for lipids; also included are P values for each metabolite changed during the COVID-19
11427_2021_2099_MOESM4_ESM.xlsx
Table S3 List of polar metabolites significantly changed in COVID-19, and their effects on NF-κB signaling. See Sheet 1 for those increased, Sheet 2 for those decreased, and Sheet 3 for those fluctuated metabolites in COVID-19. See also supplier information, and the concentrations of each metabolite used in screening assays.
Rights and permissions
About this article
Cite this article
Zhang, CS., Zhang, B., Li, M. et al. Identification of serum metabolites enhancing inflammatory responses in COVID-19. Sci. China Life Sci. 65, 1971–1984 (2022). https://doi.org/10.1007/s11427-021-2099-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11427-021-2099-7