Abstract
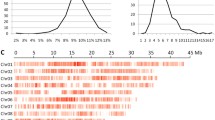
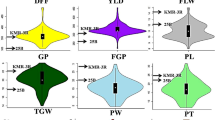
Understanding genetic characteristics in rice populations will facilitate exploring evolutionary mechanisms and gene cloning. Numerous molecular markers have been utilized in linkage map construction and quantitative trait locus (QTL) mappings. However, segregation-distorted markers were rarely considered, which prevented understanding genetic characteristics in many populations. In this study, we designed a 384-marker GoldenGate SNP array to genotype 283 recombination inbred lines (RILs) derived from 93-11 and Nipponbare Oryza sativa crosses. Using 294 markers that were highly polymorphic between parents, a linkage map with a total genetic distance of 1,583.2 cM was constructed, including 231 segregation-distorted markers. This linkage map was consistent with maps generated by other methods in previous studies. In total, 85 significant quantitative trait loci (QTLs) with phenotypic variation explained (PVE) values≥5% were identified. Among them, 34 QTLs were overlapped with reported genes/QTLs relevant to corresponding traits, and 17 QTLs were overlapped with reported sterility-related genes/QTLs. Our study provides evidence that segregation-distorted markers can be used in linkage map construction and QTL mapping. Moreover, genetic information resulting from this study will help us to understand recombination events and segregation distortion. Furthermore, this study will facilitate gene cloning and understanding mechanism of inter-subspecies hybrid sterility and correlations with important agronomic traits in rice.
Article PDF
Similar content being viewed by others
Avoid common mistakes on your manuscript.
References
Bao, J., Jin, L., Xiao, P., Shen, S., Sun, M., and Corke, H. (2008). Starch physicochemical properties and their associations with microsatellite alleles of starch-synthesizing genes in a rice RIL population. J Arg Food Chem 56, 1589–1594.
Chen, H., He, H., Zhou, F., Yu, H., and Deng, X.W. (2013). Development of genomics-based genotyping platforms and their applications in rice breeding. Curr Opin Plant Biol 16, 247–254.
Chen, H., He, H., Zou, Y., Chen, W., Yu, R., Liu, X., Yang, Y., Gao, Y.M., Xu, J.L., Fan, L.M., Li, Y., Li, Z.K., and Deng, X.W. (2011). Development and application of a set of breeder-friendly SNP markers for genetic analyses and molecular breeding of rice (Oryza sativa L.). Theor Appl Genet 123, 869–879.
Chen, H., Xie, W., He, H., Yu, H., Chen, W., Li, J., Yu, R., Yao, Y., Zhang, W., He, Y., Tang, X., Zhou, F., Deng, X.W., and Zhang, Q. (2014). A high-density SNP genotyping array for rice biology and molecular breeding. Mol Plant 7, 541–553.
Cui, Y., Zhang, F., Xu, J., Li, Z., and Xu, S. (2015). Mapping quantitative trait loci in selected breeding populations: A segregation distortion approach. Heredity 115, 538–546.
Hackett, C.A., and Broadfoot, L.B. (2003). Effects of genotyping errors, missing values and segregation distortion in molecular marker data on the construction of linkage maps. Heredity 90, 33–38.
Huang, X., Wei, X., Sang, T., Zhao, Q., Feng, Q., Zhao, Y., Li, C., Zhu, C., Lu, T., Zhang, Z., Li, M., Fan, D., Guo, Y., Wang, A., Wang, L., Deng, L., Li, W., Lu, Y., Weng, Q., Liu, K., Huang, T., Zhou, T., Jing, Y., Li, W., Lin, Z., Buckler, E.S., Qian, Q., Zhang, Q.F., Li, J., and Han, B. (2010). Genome-wide association studies of 14 agronomic traits in rice landraces. Nat Genet 42, 961–967.
Huang, X., Zhao, Y., Wei, X., Li, C., Wang, A., Zhao, Q., Li, W., Guo, Y., Deng, L., Zhu, C., Fan, D., Lu, Y., Weng, Q., Liu, K., Zhou, T., Jing, Y., Si, L., Dong, G., Huang, T., Lu, T., Feng, Q., Qian, Q., Li, J., and Han, B. (2012). Genome-wide association study of flowering time and grain yield traits in a worldwide collection of rice germplasm. Nat Genet 44, 32–39.
Jia, Y., Jia, M.H., Wang, X., and Liu, G. (2012). Indica and japonica crosses resulting in linkage block and recombination suppression on rice chromosome 12. PloS One 7, e43066.
Jiang, W., Lee, J., Jin, Y.M., Qiao, Y., Piao, R., Jang, S.M., Woo, M.O., Kwon, S.W., Liu, X., Pan, H.Y., Du, X., and Koh, H.J. (2011). Identification of QTLs for seed germination capability after various storage periods using two RIL populations in rice. Mol Cells 31, 385–392.
Lorieux, M. (2012). MapDisto: fast and efficient computation of genetic linkage maps. Mol Breeding 30, 1231–1235.
McNally, K.L., Childs, K.L., Bohnert, R., Davidson, R.M., Zhao, K., Ulat, V.J., Zeller, G., Clark, R.M., Hoen, D.R., Bureau, T.E., Stokowski, R., Ballinger, D.G., Frazer, K.A., Cox, D.R., Padhukasahasram, B., Bustamante, C.D., Weigel, D., Mackill, D.J., Bruskiewich, R.M., Ratsch, G., Buell, C.R., Leung, H., and Leach, J.E. (2009). Genomewide SNP variation reveals relationships among landraces and modern varieties of rice. Proc Natl Acad Sci USA 106, 12273–12278.
Ouyang, Y., Chen, J., Ding, J., and Zhang, Q. (2009). Advances in the understanding of inter-subspecific hybrid sterility and widecompatibility in rice. Chines Sci Bull 54, 2332–2341.
Reflinur Kim, B., Jang, S.M., Chu, S.H., Bordiya, Y., Akter, M.B., Lee, J., Chin, J.H., and Koh, H.J. (2014). Analysis of segregation distortion and its relationship to hybrid barriers in rice. Rice 7, 3.
Shah, R., Cavanagh, C.R., and Huang, B.E. (2014). Computationally efficient map construction in the presence of segregation distortion. Theor Appl Genet 127, 2585–2597.
Singh, N., Choudhury, D.R., Singh, A.K., Kumar, S., Srinivasan, K., Tyagi, R.K., Singh, N.K., and Singh, R. (2013). Comparison of SSR and SNP markers in estimation of genetic diversity and population structure of Indian rice varieties. PloS One 8, e84136.
Voorrips, R.E. (2002). MapChart: software for the graphical presentation of linkage maps and QTLs. J Hered 93, 77–78.
Wang, L., Wang, A., Huang, X., Zhao, Q., Dong, G., Qian, Q., Sang, T., and Han, B. (2011). Mapping 49 quantitative trait loci at high resolution through sequencing-based genotyping of rice recombinant inbred lines. Theor Appl Genet 122, 327–340.
Wang, S., Tan, Y., Tan, X., Zhang, Z., Wen, J., and Kou, S. (2009). Segregation distortion detected in six rice F2 populations generated from reciprocal hybrids at three altitudes. Genet Res 91, 345–353.
Xu, S. (2008). Quantitative trait locus mapping can benefit from segregation distortion. Genetics 180, 2201–2208.
Xu, S., and Hu, Z. (2009). Mapping quantitative trait Loci using distorted markers. Int J Genomics 2009, 410825.
Yang, J., Zhao, X., Cheng, K., Du, H., Ouyang, Y., Chen, J., Qiu, S., Huang, J., Jiang, Y., Jiang, L., Ding, J., Wang, J., Xu, C., Li, X., and Zhang, Q. (2012). A killer-protector system regulates both hybrid sterility and segregation distortion in rice. Science 337, 1336–1340.
Yonemaru, J.-i., Yamamoto, T., Fukuoka, S., Uga, Y., Hori, K., and Yano, M. (2010). Q-TARO: QTL Annotation Rice Online Database. Rice 3, 194–203.
Zhang, L., Wang, S., Li, H., Deng, Q., Zheng, A., Li, S., Li, P., Li, Z., and Wang, J. (2010). Effects of missing marker and segregation distortion on QTL mapping in F2 populations. Theor Appl Genet 121, 1071–1082.
Author information
Authors and Affiliations
Corresponding authors
Additional information
This article is published with open access at springerlink.bibliotecabuap.elogim.com
Electronic supplementary material
Rights and permissions
Open Access This article is distributed under the terms of the Creative Commons Attribution 4.0 International License (https://creativecommons.org/licenses/by/4.0), which permits use, duplication, adaptation, distribution, and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.
About this article
Cite this article
Yu, R., Yan, W., Liang, M. et al. Exploring the genetic characteristics of 93-11 and Nipponbare recombination inbred lines based on the GoldenGate SNP assay. Sci. China Life Sci. 59, 700–708 (2016). https://doi.org/10.1007/s11427-016-5082-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11427-016-5082-x