Abstract
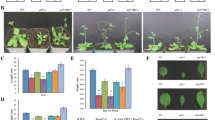
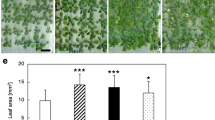
Formins (FH2 proteins) are implicated in F-actin nucleation and other aspects of cytoskeletal organization. Plants possess two formin clades, relatively well-described Class I formins and so far poorly characterized Class II formins. Comparison of Class II formin genes of two Arabidopsis species, A. thaliana and A. lyrata, indicates dynamic evolution within the Class II formin clade. Disruption of an outlier A. thaliana Class II formin gene, AtFH12 (At1g42980), whose expression is induced by NaCl, produced only negligible phenotypic effects under a variety of conditions, including salt stress, suggesting functional redundancy among Class II formins. However, the same mutation massively aggravated toxic effects of the expression of a fluorescent actin marker, GFP-tagged mouse talin (GFP-mTalin), known to interfere with normal actin dynamics. Abnormal actin structures were observed in atfh12 mutants expressing GFP-mTalin as compared to wild type. This not only demonstrates an actin-associated function for AtFH12, but also documents the feasibility of using the heterologous actin marker to “stress-test” the actin cytoskeleton in phenotyping “weak“ actin related mutant alleles.
Article PDF
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Abbreviations
- FH2:
-
formin homology 2
- LatB:
-
latrunculin B
References
Abramoff, M.D., Magelhaes, P.J., Ram, S. J.: Image processing with ImageJ. — Biophotonics Int. 11: 36–42, 2004.
Alonso, J.M., Stepanova, A.N., Leisse, T.J., Kim, C.J., Chen, H., Shinn, P., Stevenson, D.K., Zimmerman, J., Barajas, P., Cheuk, R., Gadrinab, C., Heller, C., Jeske, A., Koesema, E., Meyers, C.C., Parker, H., Prednis, L., Ansari, Y., Choy, N., Deen, H., Geralt, M., Hazari, N., Hom, E., Karnes, M., Mulholland, C., Ndubaku, R., Schmidt, I., Guzman, P., Aguilar-Henonin, L., Schmid, M., Weigel, D., Carter, D.E., Marchand, T., Risseeuw, E., Brogden, D., Zeko, A., Crosby, W.L., Berry, C.C., Ecker, J.R.: Genome-wide insertional mutagenesis of Arabidopsis thaliana. — Science 301: 653–657, 2003.
Anisimova, M., Gascuel, O.: Approximate likelihood ratio test for branches: a fast, accurate and powerful alternative. — Syst. Biol. 55: 539–552, 2006.
Banno, H., Chua, N.H.: Characterization of the Arabidopsis formin-like protein AFH1 and its interacting protein. — Plant Cell Physiol. 41: 617–626, 2000.
Bartolini, F., Gundersen, G.G.: Formins and microtubules. — Biochim. biophys. Acta 1803: 164–173, 2010.
Blanchoin, L., Staiger, C.J.: Plant formins: diverse isoforms and unique molecular mechanism. — Biochim. biophys. Acta 1803: 201–206, 2010.
Chalkia, D., Nikolaidis, N., Makalowski, W., Klein, J., Nei, M.: Origins and evolution of the formin multigene family that is involved in the formation of actin filaments. — Mol. Biol. Evol. 25: 2717–2733, 2008.
Cheung, A.Y., Wu, H.: Overexpression of an Arabidopsis formin stimulates supernumerary actin cable formation from pollen tube cell membrane. — Plant Cell 16: 257–269, 2004.
Cole, R.A., Synek, L., Žárský, V., Fowler, J.E.: SEC8, a subunit of the putative Arabidopsis exocyst complex, facilitates pollen germination and competitive pollen tube growth. — Plant Physiol. 138: 2005–2018, 2005.
Collings, D.A., Lill, A.W., Himmelspach, R., Wasteneys, G.O.: Hypersensitivity to cytoskeletal antagonists demonstrates microtubule-microfilament cross-talk in the control of root elongation in Arabidopsis thaliana. — New Phytol. 170: 275–290, 2006.
Cvrčková, F., Novotný, M., Pícková, D., Žárský, V.: Formin homology 2 domains occur in multiple contexts in angiosperms. — BMC Genomics 5: 44–--, 2004.
Deeks, M.J., Cvrčková, F., Machesky, L.M., Mikitová, V., Ketelaar, T., Žárský, V., Davies, B., Hussey, P.J.: Arabidopsis group Ie formins localize to specific cell membrane domains, interact with actin-binding proteins and cause defects in cell expansion upon aberrant expression. — New Phytol. 168: 529–540, 2005.
Deeks, M.J., Fendrych, M., Smertenko, A., Bell, K.S., Oparka, K., Cvrčková, F., Žárský, V., Hussey, P.J.: The plant formin AtFH4 interacts with both actin and microtubules, and contains a newly identified microtubule-binding domain. — J. cell. Sci. 123: 1209–1215, 2010.
Deeks, M J., Hussey, P., Davies, B.: Formins: intermediates in signal transduction cascades that affect cytoskeletal reorganization. — Trends Plant Sci. 7: 492–498, 2002.
Dereeper, A., Guignon, V., Blanc, G., Audic, S., Buffet, S., Chevenet, F., Dufayard, J.F., Guindon, S., Lefort, V., Lescot, M., Claverie, J.M., Gascuel, O: Phylogeny.fr: robust phylogenetic analysis for the non-specialist. — Nucl. Acids Res. 36: W465–W469, 2008.
Dvořáková, L., Cvrčková, F., Fischer, L.: Analysis of the hybrid proline-rich protein families from seven plant species suggests rapid diversification of their sequences and expression patterns. — BMC Genomics 8: 412, 2007.
Favery, B., Chelysheva, L. A., Lebris, M., Jammes, F., Marmagne, A., De Almeida-Engler, J., Lecomte, P., Vaury, C., Arkowitz, R. A., Abad, P.: Arabidopsis formin AtFH6 is a plasma membrane-associated protein upregulated in giant cells induced by parasitic nematodes. — Plant Cell 16: 2529–2540, 2004.
Frost, A.O., Roberts, A.W.: Cortical actin filaments fragment and aggregate to form chloroplast-associated and free Factin rings in mechanically isolated Zinnia mesophyll cells. — Protoplasma 164: 195–207, 1996.
Grunt, M., Žárský, V., Cvrčková, F.: Roots of angiosperm formins: the evolutionary history of plant FH2 domaincontaining proteins. — BMC Evol. Biol. 8: 1–19, 2008.
Guindon, S., Gascuel, O.: A simple, fast an accurate algorithm to estimate large phylogenies by maximum likelihood. — Syst. Biol. 52: 696–704, 2003.
Hartman, J.L., Garvik, B., Hartwell, L.H.: Principles for the buffering of genetic variation. — Science 291: 1001–1004, 2001.
Higgs, H.N., Peterson, K.J.: Phylogenetic analysis of the formin homology 2 domain. — Mol. Biol. Cell 16: 1–13, 2005.
Hruz, T., Laule, O., Szabo, G., Wessendorp, F., Bleuler, S., Oertle, L., Widmayer, P., Gruissem, W., Zimmermann, P.: Genevestigator v3: a reference expression database for the meta-analysis of transcriptomes. — Adv. Bioinformat. 2008: 420747–--, 2008.
Ingouff, M., Fitzgerald, J.N., Guerin, C., Robert, H., Sorensen, M.B., Van Damme, D., Geelen, D., Blanchoin, L., Berger, F.: Plant formin AtFH5 is an evolutionarily conserved actin nucleator involved in cytokinesis. — Nat. cell. Biol. 7: 374–380, 2005.
Ketelaar, T., Anthony, R.G., Hussey, P.J.: Green fluorescent protein-mTalin causes defects in actin organization and cell expansion in Arabidopsis and inhibits actin depolymerizing factor’s actin depolymerizing activity in vitro. — Plant Physiol. 136: 3990–3998, 2004.
Koch, M.A., Haubold, B., Mitchell-Olds, T.: Comparative evolutionary analysis of chalcone synthase and alcohol dehydrogenase loci in Arabidopsis, Arabis, and related genera (Brassicaceae). — Mol. Biol. Evol. 17: 1483–1498, 2000.
Kurata, T., Yamamoto, K.T.: Light-stimulated root elongation in Arabidopsis thaliana. — J. Plant Physiol. 151: 346–351, 1997.
Li, Y., Shen, Y., Cai, C., Zhong, C., Zhu, L., Yuan, M., Ren, H.: The type II Arabidopsis formin14 interacts with microtubules and microfilaments to regulate cell division. — Plant Cell 22: 2710–2726, 2010.
Ma, S., Gong, Q., Bohnert, H.J.: Dissecting salt stress pathways. — J. exp. Bot. 57: 1097–1107, 2006.
Martinière, A., Gayral, P., Hawes, C., Runions, J.: Building bridges: formin1 of Arabidopsis forms a connection between the cell wall and the actin cytoskeleton. — Plant J. 66: 354–365, 2011.
McGinnis, S., Madden, T.L.: BLAST: at the core of a powerful and diverse set of sequence analysis tools. — Nucl. Acids Res. 32: W20–W25, 2004.
Michelot, A., Guerin, C., Huang, S., Ingouff, M., Richard, S., Rodiuc, N., Staiger, C.J., Blanchoin, L.: The Formin Homology 1 domain modulates the actin nucleation and bundling activity of Arabidopsis FORMIN1. — Plant Cell 17: 2296–2313, 2005.
Mondragon-Palomino, M., Gaunt, B.S.: Gene conversion and the evolution of three leucine-rich repeat gene families in Arabidopsis thaliana. — Mol. Biol. Evol. 22: 2444–2456, 2005.
Nakhamchik, A., Zhao, Z., Provart, N.J., Shiu, S.H., Keatley, S.K., Cameron, R.K., Goring, D.R.: A comprehensive expression analysis of the Arabidopsis proline-rich extensin-like receptor kinase gene family using bioinformatic and experimental approaches. — Plant Cell Physiol. 45: 1875–1881, 2004.
Nijman, S.M.: Synthetic lethality: general principles, utility and detection using genetic screens in human cells. — FEBS Lett. 585:1-6, 2011.
Ooi, S.L., Pan, X., Peyser, B.D., Ye, P., Meluh, P.B., Yuan, D.S., Irizarry, R.A., Bader, J.S., Spencer, F.A., Boeke, J.D.: Global synthetic-lethality analysis and yeast functional profiling. — Trends Genet. 22: 56–63, 2006.
Paul, A.S., Pollard, T.D.: Review of the mechanism of processive actin filament elongation by formins. — Cell Motility Cytoskeleton 66: 606–617, 2009.
Stern, A., Doron-Faigenboim, A., Erez, E., Martz, E., Bacharach, E., Pupko, T.: Selecton 2007: advanced models for detecting positive and purifying selection using a Bayesian inference approach. — Nucl. Acids Res. 35: W506–W511, 2007.
Vidali, L., Van Gisbergen, P.A., Guerin, C., Franco, P., Li, M., Burkart, G.M., Augustine, R.C., Blanchoin, L., Bezanilla, M.: Rapid formin-mediated actin-filament elongation is essential for polarized plant cell growth. — Proc. nat. Acad. Sci. USA 106: 13341–13346, 2009.
Wernersson, R., Pedersen, A.G.: RevTrans — constructing alignments of coding DNA from aligned amino acid sequences. — Nucl. Acids Res. 31: 3537–3539, 2003.
Winter, D., Vinegar, B., Nahal, H., Ammar, R., Wilson, G.V., Provart, N.J.: An “Electronic Fluorescent Pictograph” browser for exploring and analyzing large-scale biological data sets. — PLOS One 2: e718, 2007.
Xu, Y., Moseley, J.B., Sagot, I., Poy, F., Pellman, D., Goode, B.L., Eck, M.J.: Crystal structures of a formin homology-2 domain reveal a tethered dimer architecture. — Cell 116: 711–723, 2004.
Xue, X.H., Guo, C.Q., Du, F., Lu, Q.L., Zhang, C.M., Ren, H.Y.: AtFH8 is involved in root development under effect of lowdose Latrunculin B in dividing cells. — Mol. Plant 4: 264–278, 2011.
Yamada, K., Lim, J., Dale, J.M., Chen, H., Shinn, P., Palm, C.J., Southwick, A.M., Wu, H.C., Kim, C., Nguyen, M., Pham, P., Cheuk, R., Karlin-Newmann, G., Liu, S.X., Lam, B., Sakano, H., Wu, T., Yu, G., Miranda, M., Quach, H.L., Tripp, M., Chang, C.H., Lee, J.M., Toriumi, M., Chan, M.M., Tang, C.C., Onodera, C.S., Deng, J.M., Akiyama, K., Ansari, Y., Arakawa, T., Banh, J., Banno, F., Bowser, L., Brooks, S., Carninci, P., Chao, Q., Choy, N., Enju, A., Goldsmith, A.D., Gurjal, M., Hansen, N.F., Hayashizaki, Y., Johnson-Hopson, C., Hsuan, V.W., Iida, K., Karnes, M., Khan, S., Koesema, E., Ishida, J., Jiang, P.X., Jones, T., Kawai, J., Kamiya, A., Meyers, C., Nakajima, M., Narusaka, M., Seki, M., Sakurai, T., Satou, M., Tamse, R., Vaysberg, M., Wallender, E.K., Wong, C., Yamamura, Y., Yuan, S., Shinozaki, K., Davis, R.W., Theologis, A., Ecker, J.R.: Empirical analysis of transcriptional activity in the Arabidopsis genome. — Science 302: 842–846, 2003.
Yang, W., Ren, S., Zhang, X., Gao, M., Ye, S., Qi, Y., Zheng, Y., Wang, J., Zeng, L., Li, Q., Huang, S., He, Z.: BENT UPPERMOST INTERNODE1 encodes the class II formin FH5 crucial for actin organization and rice development. — Plant Cell 23: 661–680, 2011.
Ye, J., Zheng, Y., Yan, A., Chen, N., Wang, Z., Huang, S., Yang, Z.: Arabidopsis formin3 directs the formation of actin cables and polarized growth in pollen tubes. — Plant Cell 21: 3868–3884, 2009.
Yi, K., Guo, C., Chen, D., Zhao, B., Yang, B., Ren, H.: Cloning and functional characterization of a formin-like protein (AtFH8) from Arabidopsis. — Plant Physiol. 138: 1071–1082, 2005.
Zhang, Z., Zhang, Y., Tan, H., Wang, Y., Li, G., Liang, W., Yuan, Z., Hu, J., Ren, H., Zhang, D.: RICE MORPHOLOGY DETERMINANT encodes the type II formin FH5 and regulates rice morphogenesis. — Plant Cell 23: 681–700, 2011.
Zimmermann, P., Hirsch-Hoffmann, M., Hennig, L., Gruissem, W.: GENEVESTIGATOR. Arabidopsis microarray database and analysis toolbox. — Plant Physiol. 136: 2621–2632, 2004.
Author information
Authors and Affiliations
Corresponding author
Additional information
Acknowledgements: We appreciate the possibility to use Arabidopsis lyrata sequence data produced by the US Department of Energy Joint Genome Institute in collaboration with the user community. We thank Dr. Tijs Ketelaar for the GFP:mTalin line and Marta Čadyová for technical assistance. This work has been financially supported by the Ministry of Education of the Czech Republic MSM 0021620858, Charles University SVV 263210 and the Grant Agency of the Czech Republic P305/10/0433 projects.
Rights and permissions
About this article
Cite this article
Cvrčková, F., Grunt, M. & Žárský, V. Expression of GFP-mTalin reveals an actin-related role for the Arabidopsis Class II formin AtFH12. Biol Plant 56, 431–440 (2012). https://doi.org/10.1007/s10535-012-0071-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10535-012-0071-9