Summary.
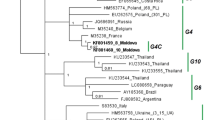
Isolates of bovine immunodeficiency virus (BIV) exhibit a striking genomic diversity, most of which are located in the viral envelope gene. Since this property of the BIV group of viruses may play an important role in the pathobiology of the virus, the surface envelope gene, particularly the conserved (C) 2, hypervariable (V)1, V2 and C3 regions, of eleven different isolates from different environments with different bovine breeds naturally infected with BIV, including dairy cows in Japan, buffaloes in Pakistan and draught animals in Cambodia, were sequenced. When compared to the nucleotide sequence of American BIV isolates, all Asian BIV field isolates seem to be smaller, several base substitutions were observed in the V1 region, and deletions were also found in the V2 region of env gene in these samples. However, deduced amino acid sequences were not so different among isolates from different bovine breeds, suggesting that bovine susceptibility to BIV infection may not depend upon bovine breed or buffaloes. Moreover, phylogenetic analysis revealed that genotypes were distinct between Asian and American BIV isolates and these results also provide an information on the molecular epidemiology of naturally occurring BIV infection in cattle and buffaloes
Article PDF
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Author information
Authors and Affiliations
Additional information
Accepted October 7, 2000
Rights and permissions
About this article
Cite this article
Meas, S., Ohashi, K., Sugimoto, C. et al. Phylogenetic relationships of bovine immunodeficiency virus in cattle and buffaloes based on surface envelope gene sequences. Arch. Virol. 146, 1037–1045 (2001). https://doi.org/10.1007/s007050170134
Issue Date:
DOI: https://doi.org/10.1007/s007050170134