Abstract.
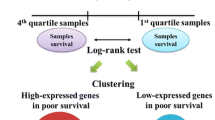
We analyzed associations between gene expression in breast cancer and patient survival for 8024 genes from a previously published microarray data set. Analysis of survival, by using the logrank test, was performed automatically for each gene. After correcting for multiple testing, we identified 95 genes whose expression was significantly associated with patient survival. The independent prognostic value of the genes ranking the highest in univariate analysis, together with clinical parameters, was assessed by Cox multivariate regression anaysis. The P-values from these logrank tests were also mapped to chromosomal positions and compared with previously reported amplicon regions. We used PubGene web tools to identify groups of genes that had co-occurred in the literature and whose expression patterns were associated with survival. Our analyses demonstrate the comprehensiveness of the microarray technology with respect to measuring gene expression and indicate that the technology may be used to screen for potential clinical markers.
Article PDF
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Author information
Authors and Affiliations
Additional information
Electronic Publication
Rights and permissions
About this article
Cite this article
Jenssen, .TK., Kuo, .W., Stokke, .T. et al. Associations between gene expressions in breast cancer and patient survival. Hum Genet 111, 411–420 (2002). https://doi.org/10.1007/s00439-002-0804-5
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1007/s00439-002-0804-5