Abstract
Bacterial expression systems play an indispensable role in the biosynthesis of recombinant proteins. Different proteins and the tasks associated with them may require different systems. The purpose of this work is to make an expression vector that allows switching on and off the expression of the target gene during cell incubation. Several expression vectors for use in Escherichia coli cells were developed using elements of the luxR/luxI type quorum sensing system of psychrophilic bacterium Aliivibrio logei. These vectors contain A. logei luxR2 and (optionally) luxI genes and LuxR2-regulated promoter, under the control of which a target gene is intended to be inserted. The synthesis of the target protein depends directly on the temperature: gene expression starts when the temperature drops to 22 °C and stops when it rises to 37 °C, which makes it possible to fix the desired amount of the target protein in the cell. At the same time, the expression of the target gene at a low temperature depends on the concentration of the autoinducer (L-homoserine N-(3-oxohexanoyl)-lactone, AI) in the culture medium in a wide range from 1 nM to 10 μM, which makes it possible to smoothly regulate the rate of target protein synthesis. Presence of luxI in the vector provides the possibility of autoinduction. Constructed expression vectors were tested with gfp, ardA, and ardB genes. At maximum, we obtained the target protein in an amount of up to 33% of the total cellular protein.
Key points
• A. logei quorum sensing system elements were applied in new expression vectors
• Expression of target gene is inducible at 22 °C and it is switched off at 37 °C
• Target gene expression at 22 °C is tunable by use different AI concentrations
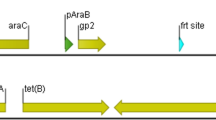
Graphical Abstract







Similar content being viewed by others
Data availability
Constructed plasmids and their nucleotide sequences are available at https://nbp.biophystech.ru/. The datasets generated during and analyzed during the current study are available from the corresponding author on reasonable request.
References
Balabanov VP, Kudryavtseva AA, Melkina OE, Pustovoit KS, Khrulnova SA, Zavilgelsky GB (2019) ArdB protective activity for unmodified λ phage against EcoKI restriction decreases in UV-treated Escherichia coli. Curr Microbiol 76:1374–1378. https://doi.org/10.1134/S0006297920030074
Bang SS, Baumann P, Nealson KH (1978) Phenotypic characterization of Photobacterium logei (sp. nov.), a species related to P. fischeri. Curr Microbiol 1:285–288. https://doi.org/10.1007/BF02601683
Bazhenov SV (2020) [LuxI/LuxR “quorum sensing” systems of bacteria from Aliivibrio genus]. Dissertation, Moscow Institute of Physics and Technology
Bazhenov SV, Kessenikh AG, Novoyatlova US, Scheglova ES, Fomin VV, Khrulnova SA, Zavilgelsky GB, Kuznetsova SB, Manukhov IV (2022a) [Bacterial lux-biosensor with increased sensitivity for detection of acyl derivatives of homoserine lactone] RU patent RU2777196C2, Moscow
Bazhenov SV, Novoyatlova US, Scheglova ES, Fomin VV, Khrulnova SA, Melkina OE, Chistyakov VA, Manukhov IV (2021a) Influence of the luxR regulatory gene dosage and expression level on the sensitivity of the whole-cell biosensor to acyl-homoserine lactone. Biosensors 11:166. https://doi.org/10.3390/bios11060166
Bazhenov SV, Melkina OE, Fomin VV, Scheglova ES, Krasnik PV, Khrulnova SA, Zavilgelsky GB, Manukhov IV (2021b) LitR directly upregulates autoinducer synthesis and luminescence in Aliivibrio logei. PeerJ 9:e12030. https://doi.org/10.7717/peerj.12030
Bazhenov SV, Scheglova ES, Fomin VV, Zavilgelsky GB, Manukhov IV (2022b) Two-stage activation of lux regulon of psychrophilic marine luminescent bacteria Aliivibrio logei. Russ J Genet 58:143–151. https://doi.org/10.1134/S1022795422020028
Blackwell HE, Geske GD, O’Neill JC (2008) Modulation of bacterial quorum sensing with synthetic ligands. US patent US7910622B2, London
Chen R (2012) Bacterial expression systems for recombinant protein production: E. coli and beyond. Biotechnol Adv 30:1102–1107. https://doi.org/10.1016/J.BIOTECHADV.2011.09.013
Choi YJ, Morel L, Le FT, Bourque D, Lucie B, Groleau D, Massie B, Miguez CB (2010) Novel, versatile, and tightly regulated expression system for Escherichia coli strains. Appl Environ Microbiol 76:5058–5066. https://doi.org/10.1128/AEM.00413-10
Colton DM, Stabb EV, Hagen SJ (2015) Modeling analysis of signal sensitivity and specificity by Vibrio fischeri LuxR variants. PLoS ONE 10:1–21. https://doi.org/10.1371/journal.pone.0126474
Du F, Liu YQ, Xu YS, Li ZJ, Wang YZ, Zhang ZX, Sun XM (2021) Regulating the T7 RNA polymerase expression in E. coli BL21 (DE3) to provide more host options for recombinant protein production. Microb Cell Fact 20:189. https://doi.org/10.1186/S12934-021-01680-6
Eberhard A, Burlingame AL, Eberhard C, Kenyon GL, Nealson KH, Oppenheimer NJ (1981) Structural identification of autoinducer of Photobacterium fischeri luciferase. Biochemistry 20:2444–2449. https://doi.org/10.1021/bi00512a013
Ferrer M, Chernikova TN, Timmis KN, Golyshin PN (2004) Expression of a temperature-sensitive esterase in a novel chaperone-based Escherichia coli strain. Appl Environ Microbiol 70:4499–4504. https://doi.org/10.1128/AEM.70.8.4499-4504.2004
Ferrer M, Chernikova TN, Yakimov MM, Golyshin PN (2003) Timmis KN (2003) Chaperonins govern growth of Escherichia coli at low temperatures. Nat Biotechnol 2111(21):1266–1267. https://doi.org/10.1038/nbt1103-1266
Gibson DG, Young L, Chuang RY, Venter JC, Hutchison CA, Smith HO (2009) Enzymatic assembly of DNA molecules up to several hundred kilobases. Nat Methods 6:343–345. https://doi.org/10.1038/nmeth.1318
Hansen H, Purohit AA, Leiros HKS, Johansen JA, Kellermann SJ, Bjelland AM, Willassen NP (2015) The autoinducer synthases LuxI and AinS are responsible for temperature-dependent AHL production in the fish pathogen Aliivibrio salmonicida. BMC Microbiol 15:1–13. https://doi.org/10.1186/s12866-015-0402-z
Hensel Z (2017) A plasmid-based Escherichia coli gene expression system with cell-to-cell variation below the extrinsic noise limit. PlosOne 12:e0187259. https://doi.org/10.1371/journal.pone.0187259
Kessenikh A, Gnuchikh E, Bazhenov S, Bermeshev M, Pevgov V, Samoilov V, Shorunov S, Maksimov A, Yaguzhinsky L, Manukhov I (2020) Genotoxic effect of 2,2’-bis(bicyclo[2.2.1] heptane) on bacterial cells. PLoS One 15(8):e0228525. https://doi.org/10.1371/journal.pone.0228525
Kessenikh AG, Novoyatlova US, Bazhenov SV, Stepanova EA, Khrulnova SA, Gnuchikh EY, Kotova VY, Kudryavtseva AA, Bermeshev MV, Manukhov IV (2021) Constructing of Bacillus subtilis-based lux-biosensors with the use of stress-inducible promoters. Int J Mol Sci 22:9571. https://doi.org/10.3390/ijms22179571
Khlebnikov A, Keasling JD (2002) Effect of lacYExpression on Homogeneity of Induction from the Ptac and Ptrc Promoters by Natural and Synthetic Inducers. Biotechnol Prog 18:672–674. https://doi.org/10.1021/BP010141K
Khrulnova SA, Baranova A, Bazhenov SV, Goryanin II, Konopleva MN, Maryshev IV, Salykhova AI, Vasilyeva AV, Manukhov IV, Zavilgelsky GB (2016) Lux-operon of the marine psychrophilic bacterium Aliivibrio logei: a comparative analysis of the LuxR1/LuxR2 regulatory activity in Escherichia coli cells. Microbiology 162:717–724. https://doi.org/10.1099/mic.0.000253
Kotova VY, Manukhov IV, Zavilgelskii GB (2010) Lux-biosensors for detection of SOS-response, heat shock, and oxidative stress. Appl Biochem Microbiol 46:781–788. https://doi.org/10.1134/S0003683810080089
Kudryavtseva AA, Okhrimenko IS, Didina VS, Zavilgelsky GB, Manukhov IV (2020) Antirestriction Protein ArdB (R64) Interacts with DNA. Biochem (mosc) 85:318–325. https://doi.org/10.1134/S0006297920030074
Larsen MH, Figurski DH (1994) Structure, expression, and regulation of the kilC operon of promiscuous IncP alpha plasmids. J Bacteriol 176:5022–5032. https://doi.org/10.1128/JB.176.16.5022-5032.1994
Lozano Terol G, Gallego-Jara J, Sola Martínez RA, Martínez Vivancos A, Cánovas Díaz M, de Diego PT (2021) Impact of the expression system on recombinant protein production in Escherichia coli BL21. Front Microbiol 12:1511. https://doi.org/10.3389/FMICB.2021.682001
Manukhov IV, Khrul’nova SA, Baranova A, Zavilgelsky GB (2011) Comparative analysis of the lux operons in Aliivibriologei KCh1 (a Kamchatka Isolate) and Aliivibriosalmonicida. J Bacteriol 193:3998–4001. https://doi.org/10.1128/JB.05320-11
Manukhov IV, Kotova VI, Zavil’gel’skiĭ GB (2006) Host factors in the regulation of the Vibrio fischeri lux operon in Escherichia coli cells. Mikrobiologiia 75:525–531
Melkina OE, Goryanin II, Bazhenov SV, Manukhov IV, Zavilgelsky GB (2019) Comparative analysis of Aliivibrio logei luxR1 and luxR2 genes regulation in Escherichia coli cells. Arch Microbiol 201:1415–1425. https://doi.org/10.1007/s00203-019-01691-3
Menart V, Jevševar S, Vilar M, Trobiš A, Pavko A (2003) Constitutive versus thermoinducible expression of heterologous proteins in Escherichia coli based on strong PR, PL promoters from phage lambda. Biotechnol Bioeng 83:181–190. https://doi.org/10.1002/BIT.10660
Nocadello S, Swennen EF (2012) The new pLAI (lux regulon based auto-inducible) expression system for recombinant protein production in Escherichia coli. Microb Cell Fact 11:3. https://doi.org/10.1186/1475-2859-11-3
Qing G, Ma LC, Khorchid A, Swapna GVT, Mal TK, Takayama MM, Xia B, Phadtare S, Ke H, Acton T, Montelione GT, Ikura M (2004) Inouye M (2004) Cold-shock induced high-yield protein production in Escherichia coli. Nat Biotechnol 227(22):877–882. https://doi.org/10.1038/nbt984
Riggs PD (2018) Overview of protein expression vectors for E. coli. Curr Protoc Essent Lab Tech 17:1–22. https://doi.org/10.1002/cpet.23
Rosano GL, Ceccarelli EA (2014) Recombinant protein expression in Escherichia coli: Advances and challenges. Front Microbiol 5:172. https://doi.org/10.3389/FMICB.2014.00172
Sambrook J, Russell DW (2001) Molecular cloning : a laboratory manual. Cold Spring Harbor Laboratory Press, New York
Shirano Y, Shibata D (1990) Low temperature cultivation of Escherichia coli carrying a rice lipoxygenase L-2 cDNA produces a soluble and active enzyme at a high level. FEBS Lett 271:128–130. https://doi.org/10.1016/0014-5793(90)80388-Y
Sitnikov DM, Schineller JB, Baldwin TO (1995) Transcriptional regulation of bioluminesence genes from Vibrio fischeri. Mol Microbiol 17:801–812. https://doi.org/10.1111/j.1365-2958.1995.mmi_17050801.x
Stargardt P, Striedner G, Mairhofer J (2021) Tunable expression rate control of a growth-decoupled T7 expression system by l-arabinose only. Microb Cell Fact 20:27. https://doi.org/10.1186/S12934-021-01512-7
Stevens AM, Dolan KM, Greenberg EP (1994) Synergistic binding of the Vibrio fischeri LuxR transcriptional activator domain and RNA polymerase to the lux promoter region. Proc Natl Acad Sci U S A 91:12619–12623. https://doi.org/10.1073/pnas.91.26.12619
Studier FW (2005) Protein production by auto-induction in high density shaking cultures. Protein Expr Purif 41:207–234. https://doi.org/10.1016/j.pep.2005.01.016
Széliová D, Krahulec J, Šafránek M, Lišková V, Turňa J (2016) Modulation of heterologous expression from PBAD promoter in Escherichia coli production strains. J Biotechnol 236:1–9. https://doi.org/10.1016/J.JBIOTEC.2016.08.004
Van Dyk TK, Majarian WR, Konstantinov KB, Young RM, Dhurjati PS, LaRossa RA (1994) Rapid and sensitive pollutant detection by induction of heat shock gene- bioluminescence gene fusions. Appl Environ Microbiol 60:1414–1420. https://doi.org/10.1128/aem.60.5.1414-1420.1994
Vera A, González-Montalbán N, Arís A, Villaverde A (2007) The conformational quality of insoluble recombinant proteins is enhanced at low growth temperatures. Biotechnol Bioeng 96:1101–1106. https://doi.org/10.1002/BIT.21218
Winson MK, Swift S, Fish L, Throup JP, Jorgensen F, Chhabra SR, Bycroft BW, Williams P, Stewart GSA (1998) Construction and analysis of luxCDABE -based plasmid sensors for investigating N -acyl homoserine lactone-mediated quorum sensing. FEMS Microbiol Lett 163:185–192. https://doi.org/10.1111/j.1574-6968.1998.tb13044.x
Funding
Expression vector constructing and testing with gfp by SVB and ESS were supported of the Ministry of Science and Higher Education of the Russian Federation (grant MK-1164.2022.1.4). Theoretical assessment of applicability in biotechnology, literature review, and revision of the text of the publication, conducted by IVM, was supported by the Ministry of Science and Higher Education of the Russian Federation (project 1022060200069–0-1.6.2;1.6.4;1.6.5;1.6.10;1.6.19, “Development of technology for rational and highly productive use of agro- and bioresources, their efficient processing and obtaining safe and high-quality sources of food and non-food products”). Cloning and expression of ardA and ardB antirestriction genes, conducted by AAU and AAK, was supported by the Russian Science Foundation (project No. 22–74-00027).
Author information
Authors and Affiliations
Contributions
SVB and IVM conceived and designed research. SVB, ESS, RAE, AAU, and AAK conducted experiments. SVB and IVM analyzed data and wrote the original draft. SVB, ESS, AAU, AAK, and IVM reviewed and edited manuscript. All authors read and approved the manuscript.
Corresponding author
Ethics declarations
Ethics approval
This article does not report any studies with human participants or animals performed by the authors.
Conflict of interest
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Bazhenov, S.V., Scheglova, E.S., Utkina, A.A. et al. New temperature-switchable acyl homoserine lactone-regulated expression vector. Appl Microbiol Biotechnol 107, 807–818 (2023). https://doi.org/10.1007/s00253-022-12341-y
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00253-022-12341-y