Abstract.
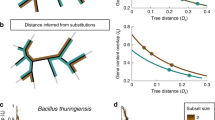
One of the main causes of bacterial chromosome asymmetry is replication-associated mutational pressure. Different rates of nucleotide substitution accumulation on leading and lagging strands implicate qualitative and quantitative differences in the accumulation of mutations in protein coding sequences lying on different DNA strands. We show that the divergence rate of orthologs situated on leading strands is lower than the divergence rate of those situated on lagging strands. The ratio of the mutation accumulation rate for sequences lying on lagging strands to that of sequences lying on leading strands is rather stable and time-independent. The divergence rate of sequences which changed their positions, with respect to the direction of replication fork movement, is not stable—sequences which have recently changed their positions are the most prone to mutation accumulation. This effect may influence estimations of evolutionary distances between species and the topology of phylogenetic trees.
Article PDF
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Author information
Authors and Affiliations
Additional information
Received: 24 July 2000 / Accepted: 16 January 2001
Rights and permissions
About this article
Cite this article
Szczepanik, D., Mackiewicz, P., Kowalczuk, M. et al. Evolution Rates of Genes on Leading and Lagging DNA Strands. J Mol Evol 52, 426–433 (2001). https://doi.org/10.1007/s002390010172
Issue Date:
DOI: https://doi.org/10.1007/s002390010172