Abstract.
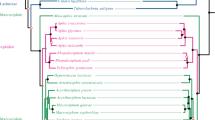
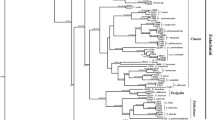
A+T content, phylogenetic relationships, codon usage, evolutionary rates, and ratio of synonymous versus non-synonymous substitutions have been studied in partial sequences of the atpD and aroQ/pheA genes of primary (Buchnera) and secondary symbionts of aphids and a set of selected non-symbiotic bacteria, belonging to the five subdivisions of the Proteobacteria. Compared to the homologous genes of the last group, both genes belonging to Buchnera behave in a similar way, showing a higher A+T content, forming a monophyletic group, a loss in codon bias, especially in third base position, an evolutionary acceleration and an increase in the number of non-synonymous substitutions, confirming previous results reported elsewhere for other genes. When available, these properties have been partly observed with the secondary symbionts, but with values that are intermediate between Buchnera and free living Proteobacteria. They show high A+T content, but not as high as Buchnera, a non-solved phylogenetic position between Buchnera, and the other γ-Proteobacteria, a loss in codon bias, again not as high as in Buchnera and a significant evolutionary acceleration in the case of the three atpD genes, but not when considering aroQ/pheA genes. These results give support to the hypothesis that they are symbionts at different stages of the symbiotic accommodation to the host.
Article PDF
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Author information
Authors and Affiliations
Rights and permissions
About this article
Cite this article
Moya, A., Latorre, A., Sabater-Muñoz, B. et al. Comparative Molecular Evolution of Primary (Buchnera) and Secondary Symbionts of Aphids Based on Two Protein-Coding Genes. J Mol Evol 55, 127–137 (2002). https://doi.org/10.1007/s00239-001-2307-8
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1007/s00239-001-2307-8