Abstract.
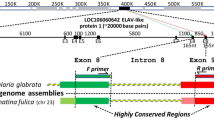
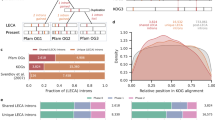
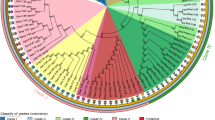
To test the validity of intron–exon structure as a phylogenetic marker, the intron–exon structure of EF-1α genes was investigated for starfish, acornworms, ascidians, larvaceans, and amphioxus and compared with that of vertebrates. Of the 11 distinct intron insertion sites found within the coding regions of the deuterostome EF-1α genes, 7 are shared by several taxa, while the remainder are unique to certain taxa. Examination of the shared introns of the deuterostome EF-1α gene revealed that independent intron loss or intron insertion must have occurred in separate lineages of the deuterostome taxa. Maximum parsimony analysis of the intron–exon data matrix recovered five parsimonious trees (consistency index = 0.867). From this result, we concluded that the intron–exon structure of deuterostome EF-1α has evolved more dynamically than previously thought, rendering it unsuitable as a phylogenetic marker. We also reconstructed an evolutionary history of intron insertion–deletion events on the deuterostome phylogeny, based on several molecular phylogenetic studies. These analyses revealed that the deuterostome EF-1α gene has lost individual introns more frequently than all introns simultaneously.
Article PDF
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Author information
Authors and Affiliations
Rights and permissions
About this article
Cite this article
Wada, H., Kobayashi, M., Sato, R. et al. Dynamic Insertion–Deletion of Introns in Deuterostome EF-1α Genes. J Mol Evol 54, 118–128 (2002). https://doi.org/10.1007/s00239-001-0024-y
Issue Date:
DOI: https://doi.org/10.1007/s00239-001-0024-y