Abstract.
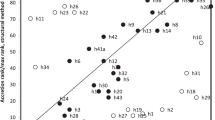
The secondary structure of rRNA internal transcribed spacer 2 is important in the process of ribosomal biogenesis. Trematode ITS sequences are poorly conserved and difficult to align for phylogenetic comparisons above a family level. If a conserved secondary structure can be identified, it can be used to guide primary sequence alignments. ITS2 sequences from 39 species were compared. These species span four orders of trematodes (Echinostomiformes, Plagiorchiformes, Strigeiformes, and Paramphistomiformes) and one monogenean (Gyrodactyliformes). The sequences vary in length from 251 to 431 bases, with an average GC content of 48%. The monogenean sequence could not be aligned with confidence to the trematodes. Above the family level trematode sequences were alignable from the 5′ end for 139 bases. Secondary structure foldings predicted a four-domain model. Three folding patterns were required for the apex of domain B. The folding pattern of domains C and D varies for each family. The structures display a high GC content within stems. Bases A and U are favored in unpaired regions and variable sites cluster. This produces a mosaic of conserved and variable regions with a structural conformation resistant to change. Two conserved strings were identified, one in domain B and the other in domain C. The first site can be aligned to a processing site identified in yeast and rat. The second site has been found in plants, and structural location appears to be important. A phylogenetic tree of the trematode sequences, aligned with the aid of secondary structures, distinguishes the four recognized orders.
Article PDF
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Author information
Authors and Affiliations
Additional information
Received: 21 November 1997 / Accepted: 9 February 1998
Rights and permissions
About this article
Cite this article
Morgan, J., Blair, D. Trematode and Monogenean rRNA ITS2 Secondary Structures Support a Four-Domain Model. J Mol Evol 47, 406–419 (1998). https://doi.org/10.1007/PL00006398
Issue Date:
DOI: https://doi.org/10.1007/PL00006398