Abstract
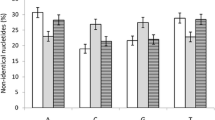
Analysis of DNA sequences of 132 introns and 140 exons from 42 pairs of orthologous genes of mouse and rat was used to compare patterns of evolutionary change between introns and exons. The mean of the absolute difference in length (measured in base pairs) between the two species was nearly five times as high in the case of introns as in the case of exons. The average rate of nucleotide substitution in introns was very similar to the rate of synonymous substitution in exons, and both were about three times the rate of substitution at nonsyn-onymous sites in exons. G+C content of introns and exons of the same gene were correlated; but mean G+C content at the third positions of exons was significantly higher than that of introns or positions 1-2 of exons from the same gene. G+C content was conserved over evolutionary time, as indicated by strong correlations between mouse and rat; but the change in G+C content was greatest at position 3 of exons, intermediate in introns, and lowest at positions 1-2 in introns.
Article PDF
Similar content being viewed by others
Avoid common mistakes on your manuscript.
References
Bernardi G (1993) The vertebrate genome: isochores and evolution. Mol Biol Evol 10:186–204
Bird AP (1987) CpG islands as gene markers in the vertebrate nucleus. Trends Genet 3:342–347
Bulmer M (1987) A statistical analysis of nucleotide sequences of introns and exons in human genes. Mol Biol Evol 4:395–405
Cereb N, Hughes AL, Yang, SY (1997) Locus-specific conservation of the HLA class I introns by intra-locus homogenization. Immunogenetics (in press)
Duret L, Mouchiroud D, Gouy M (1994) HOVERGEN: a database of homologous vertebrate genes. Nucl Acids Res 22:2360–2365
Filipski J (1987) Correlation between molecular clock ticking, codon usage, fidelity of DNA repair, chromosome banding and chromatin compactness in germline cells. FEBS Lett 217:184–186
Filipski J (1988) Why the rate of silent codon substitutions is variable within a vertebrate’s genome. J Theor Biol 134:159–164
Gillespie J (1991) The causes of molecular evolution. Oxford University Press, New York
Graur D (1985) Amino acid composition and the evolutionary rates of protein-coding genes. J Mol Evol 22:53–62
Higgins DG, Bleasby AJ, Fuchs R (1992) Clustal V: improved software for multiple sequence alignment. Comput Appl Biosci 8:189–191
Hughes AL (1997) Rapid evolution of immunoglobulin superfamily C2 domains expressed in immune system cells. Mol Biol Evol 14:1–5
Hughes AL, Nei M (1988) Pattern of nucleotide substitution at MHC class I loci reveals overdominant selection. Nature 335:167–170
Hughes AL, Hughes MK (1995) Natural selection on the peptidebinding regions of major histocompatibility complex molecules. Immunogenetics 42:233–243
Janke A, Feldmaier-Fuchs, Thomas WK, von Haeseler A, Paabo S (1994) The marsupial mitochondrial genome and the evolution of placental mammals. Genetics 137:243–256
Jukes TH, Cantor CR (1969) Evolution of protein molecules. In: Munro HN (ed) Mammalian protein metabolism. Academic Press, New York, pp 21–132
Kimura M (1977) Preponderance of synonymous changes as evidence for the neutral theory of molecular evolution. Nature 267:275–276
Kimura M (1983) The neutral theory of molecular evolution. Cambridge University Press, Cambridge
Kimura M, Crow JF (1978) Effect of overall phenotypic selection on genetic change at individual loci. Proc Natl Acad Sci USA 75: 6188–6171
Kumar SA, Tamura K, Nei M (1993) MEGA: molecular evolutionary genetic analysis, version 1.0. Pennsylvania State University, University Park
Lewin B (1994) Genes V. Oxford University Press, Oxford
Li W-H, Graur D (1991) Fundamentals of molecular evolution. Sinauer Associates, Sunderland, MA
Mayer WE, Jonker D, Klein D, Ivanyi O, van Seventer G, Klein J (1988) Nucleotide sequence of chimpanzee MHC class I alleles: evidence for trans-species mode of evolution. EMBO J 7:2765–2774
Milkman R (1978) Selection differentials and selection coefficients. Genetics 88:391–403
Mouchiroud D, Gautier C, Bernardi G (1995) Frequencies of synonymous substitutions in mammals are gene-specific and correlated with frequencies of nonsynonymous substitutions. J Mol Evol 40: 107–113
Nei M (1987) Molecular evolutionary genetics. Columbia University Press, New York
Nei M, Gojobori T (1986) Simple methods for estimating the numbers of synonymous and nonsynonymous nucleotide substitutions. Mol Biol Evol 3:418–426
Ogata H, Fujibachi W, Kanehisa M (1996) The size differences among mammalian introns are due to the accumulation of small deletions. FEBS Lett 390:99–103
Ohta T, Ina Y (1995) Variation in synonymous substitution rates among mammalian genes and the correlation between synonymous and nonsynonymous divergence. J Mol Evol 41:717–720
Parker PH (1996) An improved estimate of the mouse-rat divergence time and rates of amino acid substitution in mammals and birds. Unpublished M.S. thesis, Department of Biology, The Pennsylvania State University, University Park.
Shields DC, Sharp PM, Higgins DG, Wright F (1988) “Silent” sites in Drosophila genes are not neutral: evidence of selection among synonymous codons. Mol Biol Evol 5:704–716
Sueoka N (1988) Directional mutation pressure and neutral molecular evolution. Proc Natl Acad Sci USA 85:2653–2657
Wolfe KE (1991) Mammalian DNA replication: mutation biases and the mutation rate. J Theor Biol 149:441–451
Wolfe KH, Sharp PM (1993) Mammalian gene evolution: nucleotide sequence divergence between mouse and rat. J Mol Evol 37:441–456
Wolfe KH, Sharp PM, Li W-H (1989) Mutation rates differ among regions of the mammalian genome. Nature 337:283–285
Author information
Authors and Affiliations
Additional information
An erratum to this article is available at http://dx.doi.org/10.1007/PL00006330.
Rights and permissions
About this article
Cite this article
Hughes, A.L., Yeager, M. Comparative evolutionary rates of introns and exons in murine rodents. J Mol Evol 45, 125–130 (1997). https://doi.org/10.1007/PL00006211
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1007/PL00006211