Abstract
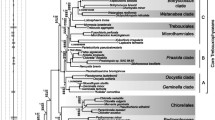
Several algae that were previously classified in the phylum Xanthophyta (yellow-green algae) were assigned in 1971 to a new phylum, Eustigmatophyta. It was anticipated that the number of algae reclassified to Eustigmatophyta would increase. However, due to the fact that the morphological characteristics that segregate eustigmatophytes from other closely related algae can be only obtained through laborious electron microscopic techniques, the number of members in this phylum have increased rather slowly. We attempted, therefore, to segregate two closely related groups of algae, eustigmatophytes and yellow-green algae, on the basis of a molecular phylogenetic tree as a means of providing an alternative method of distinguishing these phyla. We analyzed the mitochondrial cyto-chrome oxidase subunit I (COXI) gene sequences of eight algae classified as xanthophyceans and found that six manifested the expected deviant genetic code where AUA codes for methionine (AUA/Met), but not for isoleucine (AUA/Ile) as in the universal genetic code. The other two, Monodus sp. (CCMP 505) and Ophiocytium majus (CCAP 855/1), which were presumed to be yellow-green algae, and all the examined eustigmatophytes utilized AUA for Ile. In addition, the phylogenetic tree of COXI gene sequences showed that the six yellow-green algae bearing the AUA/Met deviant code composed a tight clade with a bootstrap value of 100%. The phylogenetic tree of the corresponding sequences from Monodus sp. and Ophiocytium majus and the eustigmatophytes also composed a tight cluster, but with a bootstrap value of 92%. These results strongly suggest that two previously classified members of yellow-green algae belong to the phylum Eustigmatophyta. Therefore, examination of the mitochondrial genetic code in algae appears to be a potentially very useful genetic marker for classifying these organisms, especially when it is considered with the results obtained through a molecular phylogenetic tree.
Article PDF
Similar content being viewed by others
Avoid common mistakes on your manuscript.
References
Antia NJ, Bisalputra T, Cheng JY, Kalley JP (1975) Pigment and cytological evidence for reclassification of Nannochloris oculata and Monallantus salina in Eustigmatophyceae. J Phycol 11:339–343
Bessho Y, Ohama T, Osawa S (1991) Planarian mitochondria II: The unique genetic code as deduced from COI gene sequences. J Mol Evol 34:324–330
Boyen C, Leblanc C, Bonnard G, Grienenberger J-M, Kloareg B (1994) Nucleotide sequence of the COX3 gene from Chondrus crispus: evidence that UGA encodes tryptophan and evolutionary implications. Nucleic Acids Res 22:1400–1403
Cantatore P, Roberti M, Rainaldi G, Gadaleta MN, Saccone C (1989) The complete nucleotide sequence, gene organization, and genetic code of the mitochondrial genome of Paracentrotus lividus. J Biol Chem 264:10965–10975
Douglas SE, Murphy CA, Spencer DF, Gray MW (1991) Cryptomonad algae are evolutionary chimeras of two phylogenetically distinct unicellular eukaryotes. Nature 350:148–151
Douglas SE (1992) Eukaryote-eukaryote endosymbioses: insights from studies of a cryptomonad alga. BioSystems 28:57–68
Eschbach S, Hofmann CJB, Maier G-U, Sitte P, Hansmann P (1991) A eukaryotic genome of 660 kb: electrophoretic karyotype of nucleo-morph and cell nucleus of the cryptomonad alga, Pyrenomonas salina. Nucleic Acids Res 19:1779–1781
Felsenstein J (1985) Confidence limits on phylogenies: an approach using the bootstrap. Evolution 39:783–791
Gibbs SP (1981) The chloroplasts of some algal groups may have evolved from endosymbiotic eukaryotic algae. Ann New York Acad Sci 361:193–208
Gibbs SP (1993) The evolution of algal chloroplasts. In: Lewin RA (ed) Origin of plastids. Chapman and Hall, New York, pp 107–121
H-Ishimaru Y, Ohama T, Kawatsu Y, Nakamura K, Osawa S (1996) UAG is a sense codon in several chlorophycean mitochondria. Curr Genet 30:29–33
Hibberd DJ, Leedale GF (1971) A new algal class-the Eustigmatophyceae. Taxon 20:523–525
Hibberd DJ (1981) Notes on the taxonomy and nomenclature of the algal classes Eustigmatophytes and Tribophyceae (synonym Xan-thophyceae). Botanical J Linnean Society 82:93–119
Hibberd DJ (1989a) Phylum Eustigmatophyta. In: Margulis L, Corliss JO, Melkonian M, Chapman DJ (eds) Handbook of protoctista. Jones and Bartlett Publishers, Boston, pp 326–332
Hibberd DJ (1989b) Phylum Xanthophyta. In: Margulis L, Corliss JO, Melkonian M, Chapman DJ (eds) Handbook of protoctista. Jones and Bartlett Publishers, Boston, pp 686–697
Himeno H, Masaki H, Ohta T, Kumagai I, Miura KI, Watanabe K (1987) Unusual genetic codes and a novel genome structure for tRNASer AGY in starfish mitochondrial DNA. Gene 56:219–230
Hoffmann RJ, Boore JL, Brown WM (1992) A novel mitochondrial genome organization for the blue mussel, Mytilus edulis. Genetics 131:397–412
Jacobs HT, Elliott DJ, Math VB, Farquharson A (1988) Nucleotide sequence and gene organization of sea-urchin mitochondrial DNA. J Mol Biol 202:185–217
Kimura M (1980) A simple method for estimating evolutionary rate of base substitutions through comparative studies of nucleotide sequences. J Mol Evol 16:111–120
Kumar S, Rzhetsky A (1996) Evolutionary relationships of eukaryotic kingdoms. J Mol Evol 42:183–193
Lee KW, Bold HC (1973) Pseudocharaciopsis texensis gen. et sp. nov., a new member of the Eustigmatophyceae. British Phycol J 8:31–37
Manhart JR, Palmer JD (1990) The gain of two chloroplast tRNA introns marks the green algal ancestors. Nature 345:268–270
McFadden GI, Gilson PR, Hofmann CJB, Adcock GJ, Maier U-G (1994) Evidence that an amoebae acquired a chloroplast by retaining part of an engulfed eukaryotic alga. Proc Natl Acad Sci USA 91:3690–3694
Morden CW, Delwiche CF, Kuhsel M, Palmer JD (1992) Gene phylogenies and the endosymbiotic origin of plastids. BioSystems 28: 75–90
Moriya J, Yokogawa T, Wakita K, Ueda T, Nishikawa K, Crain PF, Hashizume T, Pomerantz SC, Mccloskey JA, Kawai G, Hayashi N, Yokoyama S, Watanabe K (1994) A novel modified nucleotide found at the position of the anticodon of methionine tRNA from bovine liver mitochondria. Biochem 33:2234–2239
Muramatsu T, Yokoyama S, Horie N, Matsuda A, Ueda T, Yamaizumi Z, Kuchino Y, Nishimura S, Miyazawa T (1988a) A novel lysine-substituted nucleoside in the first position of the anticodon of minor isoleucine tRNA from Escherichia coli. J Bio Chem 263:9261–9267
Muramatsu T, Nishikawa K, Nemoto F, Kuchino Y, Nishimura S, Miyazawa T, Yokoyama S (1988b) Codon and amino acid specificities of a transfer RNA are both converted by a single post-transcriptional modification. Nature 336:179–181
Osawa S, Jukes TH, Watanabe K, Muto A (1992) Recent evidence for evolution of the genetic code. Microbiol Rev 56:229–264
Palmer JD, Delwiche CF (1996) Second-hand chloroplasts and the case of the disappearing nucleus. Proc Natl Acad Sci USA 93:7432–7435
Saitou N, Nei M (1987) The neighbor-joining method: a new method for reconstructing phylogenetic trees. Mol Biol Evol 4:406–425
Sibler AP, Dirheimer G, Martin RP (1985) Yeast mitochondrial tRNAIle and tRNAMetm: nucleotide sequence and codon recognition patterns. Nucleic Acids Res 13:1341–1345
Takasaki N, Murata S, Saitoh M, Kobayashi T, Park L, Okada N (1994) Species-specific amplification of tRNA-derived short interspersed repetitive elements (SINEs) by retroposition: a process of parasitization of entire genomes during the evolution of salmonids. Proc Natl Acad Sci USA 91:10153–10157
van dePeer Y, Rensing SA, Maier U-G, De Wachter R (1996) Substitution rate calibration of small subunit ribosomal RNA identifies chlorarachniophyte endosymbionts as remnants of green algae. Proc Natl Acad Sci USA 93:7732–7736
Weber F, Dietrich A, Weil JH, M-Drouard L (1990) A potato mitochondrial isoleucine tRNA is coded for by a mitochondrial gene possessing a methionine anticodon. Nucleic Acids Res 18:5027–5030
Author information
Authors and Affiliations
Rights and permissions
About this article
Cite this article
Ehara, M., Hayashi-Ishimaru, Y., Inagaki, Y. et al. Use of a deviant mitochondrial genetic code in yellow-green algae as a landmark for segregating members within the phylum. J Mol Evol 45, 119–124 (1997). https://doi.org/10.1007/PL00006210
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1007/PL00006210