Abstract
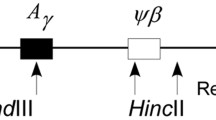
We have analyzed allelic sequence variation in sixty-one 3-kb β-globin sequences from the Melanesian population of Vanuatu to demonstrate the value of (1) turning to the autosomal nuclear genome for studies on the evolution of modern humans and (2) using new analytical methods based on a coalescent model. After excluding recombination events, β-globin sequence variants were connected in a unique gene tree. A gene tree provides more information for inferences on the population genealogy than simple summary statistics such as the average pairwise sequence difference. Estimates of the time to the most recent common ancestor (MRCA) and of the ages of each mutation, conditional on the gene tree, were made using new maximum likelihood methods assuming a coalescent model. We found that allelic β-globin variation coalesces to a single shared ancestral haplotype over a time scale of approximately 900,000 years. Three major haplotypes (A1, B1, C3) that are older than 200,000 years identify ancestral diversity contemporaneous with the single MRCA for mitochondrial variation.
Article PDF
Similar content being viewed by others
Avoid common mistakes on your manuscript.
References
Antonarakis SE, Boehm CD, Giardina PJV, Kazazian HH Jr (1982) Nonrandom association of polymorphic restriction sites in the β-globin gene cluster. Proc Natl Acad Sci USA 79:137–141
Armour JAL, Anttinen T, May CA, Vega EE, Sajantila A, Kidd JR, Kidd KK, Bertranpetit J, Pääbo S, Jeffreys AJ (1996) Minisatellite diversity supports a recent African origin for modern humans. Nat Genet 13:154–160
Ayala FJ (1995) The myth of Eve: molecular biology and human origins. Science 270:1930–1936
Cann RL, Stoneking M, Wilson AC (1987) Mitochondrial DNA and human evolution. Nature 325:31–36
Donnelly P (1996) Interpreting genetic variability: the effects of shared evolutionary history. In: Chadwick D, Cardew G (eds) Variation in the human genome. John Wiley, Chichester, p 25
Donnelly P, Tavaré S (1995) Coalescents and genealogical structure under neutrality. Annu Rev Genet 29:401–421
Ewens WJ (1972) The sampling theory of selectively neutral alleles. Theor Pop Biol 3:87–112
Fullerton SM, Harding RM, Boyce AJ, Clegg JB (1994) Molecular and population genetic analysis of allelic sequence diversity at the human β-globin locus. Proc Natl Acad Sci USA 91:1805–1809
Griffiths RC (1987) An algorithm for constructing genealogical trees. Statistics Research Report 163, Department of Mathematics, Monash University
Griffiths RC, Tavaré S (1994a) Ancestral inference in population genetics. Stat Sci 9:307–319
Griffiths RC, Tavaré S (1994b) Sampling theory for neutral alleles in a varying environment. Philos Trans R Soc Lond Biol 344:403–410
Gusfield D (1991) Efficient algorithms for inferring evolutionary trees. Networks 21:19–28
Hammer MF (1995) A recent common ancestry for human Y chromosomes. Nature 378:376–378
Harding RM (1996) New phylogenies: an introductory look at the coalescent. In: Harvey PH, Leigh Brown AJ, Maynard Smith J, Nee S (eds) New uses for new phylogenies. Oxford University Press, Oxford, p 15
Horai S, Hayasaka K, Kondo R, Tsugane K, Takahata N (1995) Recent African origin of modern humans revealed by complete sequences of hominoid mitochondrial DNAs. Proc Natl Acad Sci USA 92: 532–536
Jobling MA, Tyler-Smith C (1995) Fathers and sons: the Y chromosome and human evolution. Trends Genet 11:449–456
Kimura M (1983) The neutral theory of molecular evolution. Cambridge University Press, Cambridge
Kingman JFC (1982a) On the genealogy of large populations. J Appl Prob 19A:27–43
Kingman JFC (1982b) The coalescent. Stochastic Proc Appl 13:235–248
Kingman JFC (1982c) Exchangeability and the evolution of large populations. In: Koch G, Spizzichino F (eds) Exchangeability in probability and statistics. North-Holland, Amsterdam, p 97
Knight A, Batzer MA, Stoneking M, Tiwari HK, Scheer WD, Herrera RJ, Deininger PL (1996) DNA sequences of Alu elements indicate a recent replacement of the human autosomal genetic complement. Proc Natl Acad Sci USA 93:4360–4364
Kreitman M (1983) Nucleotide polymorphism at the alcohol dehydrogenase locus of Drosophila melanogaster. Nature 304:412–417
Li W-H, Sadler LA (1991) Low nucleotide diversity in Man. Genetics 129:513–523
Marjoram P, Donnelly P (1994) Pairwise comparisons of mitochondrial DNA sequences in subdivided populations and implications for early human evolution. Genetics 136:673–683
Orkin SH, Kazazian HH Jr, Antonarakis SE, Goff SC, Boehm CD, Sexton JP, Waber PG, Giardina PJV (1982) Linkage of β-thalassaemia mutations and β-globin gene polymorphisms with DNA polymorphisms in the human β-globin gene cluster. Nature 296: 627–631
Pluzhnikov A, Donnelly P (1996) Optimal sequencing strategies for surveying molecular genetic diversity. Genetics (in press)
Ruvolo M, Zehr S, von Dornum M, Pan D, Chang B, Lin J (1993) Mitochondrial COII sequences and modern human origins. Mol Biol Evol 10:1115–1135
Tajima F (1983) Evolutionary relationship of DNA sequences in finite populations. Genetics 105:437–460
Tajima F (1989) Statistical method for testing the neutral mutation hypothesis by DNA polymorphism. Genetics 123:585–595
Takahata N, Satta Y, Klein J (1995) Divergence time and population size in the lineage leading to modern humans. Theor Popul Biol 48:198–221
Tishkoff SA, Dietzsch E, Speed W, Pakstis AJ, Kidd JR, Cheung K, Bonné-Tamir B, Santachiara-Benerecetti AS, Moral P, Krings M, Pääbo S, Watson E, Risch N, Jenkins T, Kidd KK (1996) Global patterns of linkage disequilibrium at the CD4 locus and modern human origins. Science 271:1380–1387
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Harding, R.M., Fullerton, S.M., Griffiths, R.C. et al. A gene tree for β-globin sequences from melanesia. J Mol Evol 44 (Suppl 1), S133–S138 (1997). https://doi.org/10.1007/PL00000063
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1007/PL00000063