Summary
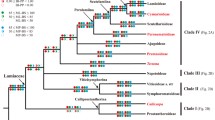
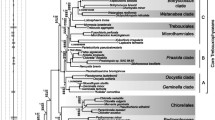
The molecular evolution of cytochrome c from angiosperms is compared to that from vertebrates. On the basis of a cladistic analysis from 26 plant species, compared to that from 27 vertebrate species, we find that although the vertebrate sequences yield reasonably well-defined minimal trees that are congruent with the biological tree, the plant sequences yield multiple minimal trees that are not only highly incongruent with each other, but none of which is congruent with any reasonably biological tree. That is, the plant sequence set is much more homoplastic than that of the animal. However, as judged by the relative rate test, the extent of divergence, and degree of functional constraint, cytochrome c evolution in plants does not appear to differ from that of vertebrates.
Article PDF
Similar content being viewed by others
Avoid common mistakes on your manuscript.
References
Boulter D (1974) The evolution of plant proteins with special reference to higher plant cytochrome c. Curr Adv Plant Sci 4(Commentaries in Plant Sci No 8):1–16
Boulter D, Peacock D, Guise A, Gleaves JT, Estabrook G (1979) Relationship between the partial amino acid sequences of plastocyanin from members of ten families of flowering plants. Phytochemistry 18:603–608
Dahlgren R, Bremer K (1986) Major clades of angiosperms. Cladistics 1:349–368
Dayhoff MO (1978) Atlas of protein sequence and structure— supplement 3. National Biomedical Research Foundation, Washington DC
Doyle JA, Donoghue MJ (1986) Seed plant phylogeny and the origin of angiosperms: an experimental cladistic approach. Bot Rev 52:34–431
Fitch WM (1967) A method for estimating the number of invariant amino acid coding positions in a gene; use of cytochrome c as a model case. Biochem Genet 1:65–71
Langley CH, Fitch WM (1974) An examination of, the constancy of the rate of evolution. J Mol Evol 3:161–177
Martin PG, Boulter D, Penny D (1985) Angiosperm phylogeny studied using sequences of five macromolecules. Taxon 34: 393–400
Peacock D (1981) Data handling for phylogenetic trees. In: Gutfreund H (ed) Biochemical evolution. Cambridge University Press, pp 88–115
Peacock K, Boulter D (1975) Use of amino acid sequence data in phylogeny and evaluation of methods using computer simulation. J Mol Biol 95:513–527
Rothwell R (1982) New interpretations of the earliest conifers. Rev Palaeobot Palynol 37:7–28
Sarich VM, Wilson AC (1967) Rates of albumin evolution in primates. Proc Natl Acad Sci USA 58:142–148
Stevens PF (1984) Metaphors and typology in the development of botanical systematics 1690–1960 or the art of putting new wine in old bottles. Taxon 33:169–211
Stevens PF (1986) Evolutionary classification in botany, 1960–1985. J. Arnold Arbor 67:313–339
Vavilov NI (1922) The law of homologous series in variation. J Genet 12:47–89
Waterman M, Smith T (1978) On the similarity of dendrograms. J Theor Biol 73:789–800
Went FW (1971) Parallel evolution. Taxon 20:197–226
Author information
Authors and Affiliations
Rights and permissions
About this article
Cite this article
Syvanen, M., Hartman, H. & Stevens, P.F. Classical plant taxonomic ambiguities extend to the molecular level. J Mol Evol 28, 536–544 (1989). https://doi.org/10.1007/BF02602934
Received:
Revised:
Issue Date:
DOI: https://doi.org/10.1007/BF02602934