Abstract
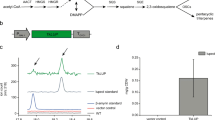
TheERG6 gene fromSaccharomyces cerevisiae has been functionally expressed inEscherichia coli, for the first time, yielding a protein that catalyzes the bisubstrate transfer reaction whereby the reactive methyl group from (S)-adenosyl-l-methionine is transferred stereoselectively to C-24 of the sterol side chain. The structural requirements of sterol in binding and catalysis were similar to the native protein fromS. cerevisiae. Inhibition of biomethylation was observed with fecosterol and ergosterol which suggests that ergosterol may function in wild-type yeast as a feedback regulator of sterol biosynthesis.
Article PDF
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Abbreviations
- AdoMet:
-
(S)-adenosyl-l-methionine
- HPLC:
-
high-performance liquid chromatography
- ORF:
-
open reading frame
- SMT:
-
(S)-adenosyl-l-methionine:Δ24(25)
References
Nes, W.R., and McKean, M.L. (1977)Biochemistry of Steroids and Other Isopentenoids, University Park Press, Baltimore, pp. 326–410.
Arigoni, D. (1978) Stereochemical Studies of Enzymic C-Methylations,Chiba Found. Symp. 103, 243–261.
Akhtar, M., and Jones, C. (1978) Some Biological Transformations Involving Unsaturated Linkages: The Importance of Charge Separation and Charge Neutralization in Enzyme Catalysis.Tetrahedron 34, 813–832.
Oehlschlager, A.C., Angus, R.H., Pierce, A.M., Pierce, H.D., and Srinvasan, R. (1984) Azasterol Inhibition of Δ24-Sterol Methyl Transferase inSaccharomyces cerevisae, Biochemistry 23, 3582–3589.
Ator, M.A., Schmidt, S.J., Adam, J.L., and Dolle, R.E. (1989) Mechanism and Inhibition of Δ24-sterol Methyl Transferase fromCandida albicans andCandida tropicalis, Biochemistry 28, 9633–9640.
Mercer, E.I. (1993) Inhibitors of Sterol Biosynthesis and Their Applications,Prog. Lipid Res. 32, 357–416.
Gaber, R.F., Copple, D.M., Kennedy, B.K., Vidal, M., and Bard, M. (1989) The Yeast GeneERG6 Is Required for Normal Membrane Functioning But Is Not Essential for Biosynthesis of the Cell-Cycle Sparking Sterol,Mole. Cell. Biol. 9, 3447–3456.
Hardwick, K.G., and Pelham, H.R.B. (1994)SED6 Is Identical toERG6 and Encodes a Putative Methyl Transferase Required for Ergosterol Biosynthesis,Yeast 10, 265–269.
Welihinda, A.A., Beavis, A.D., and Trumbly, R.J. (1994) Mutations in L1S1 (ERG6) Gene Confer Increased Sodium and Lithium Uptake inSaccharomyces cerevisiae, Biochim. Biophys. Acta 1193, 107–117.
Bolivar, F., Rodrigues, R.L., Greene, P.J., Beltach, M.C., Heyneker, H.L., and Boyer, H.W. (1977) Construction and Characterization of New Cloning Vehicles: II. A Multipurpose Cloning System,Gene 2, 95–113.
Ausubel, F.M., Brent, R., Kingston, R.E., Moore, D.D., Smith, J.A., Seidman, J.G., and Struhl, K. (1983)Current Protocols in Molecular Biology, Wiley, New York, 1988, pp. 1–100.
Hanahan, D. (1983) Studies on Transformation ofEscherichia coli with Plasmids,J. Mol. Biol. 166, 557–580.
Sanger, F., Nicklen, S., and Coulson, A.R. (1977) DNA Sequencing with Chain-Terminating Inhibitors,Proc. Natl. Acad. Sci. USA 74, 5463–5467.
McCammon, M.T., Hartmann, M.-A., Bottema, C.D.K., and Parks, L.W. (1984) Sterol Methylation inSaccharomyces cerevisiae, J. Bacteriol. 157, 475–483.
Nes, W.D., Janssen, G.G., and Bergenstrahle, A. (1991) Structural Requirements for Transformation of Substrates by the (S)-Adenosyl-l-Methionine: Δ24(25)-Sterol Methyl Transferase,J. Biol. Chem. 266, 15202–15212.
Janssen, G.G., and Nes, W.D. (1992) Structural Requirements for Transformation of Substrates by the (S)-Adenosyl-l-Methionine: Δ24(25)-Sterol Methyl Transferase. II. Inhibition by Analogs of the Transition State Coordinate,J. Biol. Chem. 267, 25856–25863.
Schachterle, G.R., and Pollack, R.L. (1973) A Simplified Method for the Quantitative Assay of Small Amounts of Protein in Biological Material,Anal. Biochem. 51, 654–655.
Nes, W.R., Adler, J.H., Frasinel, C., Nes, W.D., Young, M., and Joseph, J.M. (1980) The Independence of Photosynthesis and Aerobiosis from Sterol Biosynthesis in Bacteria,Phytochemistry 19, 1439–1443.
Guo, D., Venkatramesh, M., and Nes, W.D. (1995) Developmental Regulation ofZea mays, Lipids 30, 203–219.
Moore, J.T., and Gaylor, J.L. (1970) Investigation of anS-Adenosylmethionine: Δ24-Sterol Methyl Transferase in Ergosterol Biosynthesis: Specificity of Sterol Substrates and Inhibitors,J. Biol. Chem. 18, 4684–4688.
Venkatramesh, M., Guo, D., Jia, Z., and Nes, W.D. (1996) Mechanism and Structural Requirements for Transformation of Substrates by the (S)-Adenosyl-l-methionine: Δ24(25)-Sterol Methyl Transferase fromSaccharomyces cerevisae, Biochim. Biophys. Acta 1299, 313–324.
Zinser, E., Paltauf, F., and Daum, G. (1993) Sterol Composition of Yeast Organelle Membranes and Subcellular Distribution of Enzymes Involved in Sterol Metabolism,J. Bacteriol. 175, 2853–2856.
Author information
Authors and Affiliations
About this article
Cite this article
Venkatramesh, M., Guo, Da., Harman, J.G. et al. Sterol specificity of theSaccharomyces cerevisiae ERG6 gene product expressed inEscherichia coli . Lipids 31, 373–377 (1996). https://doi.org/10.1007/BF02522922
Received:
Revised:
Accepted:
Issue Date:
DOI: https://doi.org/10.1007/BF02522922