Abstract
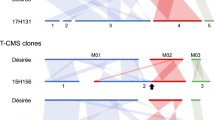
We have characterized two related regions of twoPetunia mitochondrial genomes in order to understand how plant mt genomes from a cytoplasmic male sterile (cms) line and a fertile line diverge from one another. Restriction maps of these regions indicate that a sequence arrangement shared by the two genomes adjoins sequences which are not shared at the corresponding locations in the two genomes. A point where the mt genomes from the cms line and the fertile lines diverge from each other was identified and mapped.
Previously we had observed that somatic hybrids constructed from the cms and the fertile line contained mt genomes carrying new combinations of parental mtDNA restriction fragments (3). Using the restriction maps of the two related mtDNA regions, a mtDNA arrangement unique to the cms parent could be shown to be present in all 17 stable sterile somatic hybrids tested and none of the 24 stable fertile somatic hybrids tested. This data does not exclude the possibility that additional, as yet unidentified, mtDNA arrangements unique to the cms parent might also be found exclusively in sterile somatic hybrids. Whether or not the sterile parental mtDNA arrangement reported here is functionally related to cms, it apparently segregates with cms in somatic hybrids.
Article PDF
Similar content being viewed by others
Avoid common mistakes on your manuscript.
References
Belliard G, Pelletier G, Vedel F, Quetier F: Morphological characteristics and chloroplast DNA distribution in different cytoplasmic parasexual hybrids ofNicotiana tabacum. Mol Gen Genet 165:231–237, 1978.
Belliard G, Vedel F, Pelletier G: Mitochondrial recombination in cytoplasmic hybrids ofNicotiana tabacum by protoplast fusion. Nature 281:401–403, 1979.
Boeshore ML, Lifshitz I, Hanson MR, Izhar S: Novel composition of mitochondrial genomes inPetunia somatic hybrids derived from cytoplasmic male sterile and fertile plants. Mol Gen Genet 190:459–467, 1983.
Clark EM, Hanson MR: Analysis of chloroplast genomes in small amounts of tissue. Plant Molecular Biology Reporter 1:77–79, 1983.
Dixon LK, Leaver CJ: Mitochondrial gene expression and cytoplasmic male sterility in sorghum. Pl Mol Biol 1:89–102, 1982.
Forde BG, Leaver CJ: Nuclear and cytoplasmic genes controlling synthesis of variant mitochondrial polypepides in male-sterile maize. Proc Natl Acad Sci USA 77:418–422, 1980.
Galun E, Arzee-Gonen P, Fluhr R, Edelman M, Aviv D: Cytoplasmic hybridization inNicotiana: mitochondrial DNA analysis in progenies resulting from fusion between protoplasts having different organelle constitutions. Mol Gen Genet 186:50–56, 1982.
Grunstein M, Hogness DS: Colony hybridization: a method for the isolation of cloned DNAs that contain a specific gene. Proc Natl Acad Sci USA 72:3961–3965, 1975.
Hanson M, Conde M: Functioning and variation of cytoplasmic genomes: lessons from cytoplasmic-nuclear interactions affecting male fertility in plants. Intl Review of Cytology (in press).
Izhar S, Schlicter M, Swartzberg D: Sorting out of cytoplasmic elements in somatic hybrids ofPetunia and the prevalence of the heteroplasmon through several meiotic cycles. Mol Gen Genet 190:468–474, 1983.
Karn J, Brenner S, Barnett L, Cesareni G: Novel bacteriophage λ cloning vector. Proc Natl Acad Sci 77:5172–5176, 1980.
Kemble RJ, Mans RJ: Examination of the mitochondrial genome of revertant progeny from S cms maize with cloned S-1 and S-2 hybridization probes. J Mol Appl Genet 2:161–172, 1983.
Leaver CJ, Gray MW: Mitochondrial genome organization and expression in higher plants. Ann Rev Pl Physiol 65:785–791, 1982.
Levings CS III: The plant mitochondrial genome and its mutants. Cell 32:659–661, 1983.
Levings CS III, Kin BD, Pring DR, Conde MF, Mans RJ, Laughnan JR, Gabay-Laughnan SJ: Cytoplasmic reversion of cms-S in maize: association with a transpositional event. Science 209:1021–1023, 1980.
Maniatis T, Hardison RC, Lacy E, Lauer J, O'Connell C, Quon D, Sim GK, Efstratiadis A: The isolation of structural genes from libraries of eucaryotic DNA. Cell 15:687–701, 1978.
McNay JW, Pring DR, Lonsdale DM: Polymorphism of mitochondrial DNA ‘S’ regions among normal cytoplasms of maize. Pl Mol Biol 2:177–187, 1983.
Murray MG, Thompson WF: Rapid isolation of high molecular weight plant DNA. Nuc Acids Res 8:4321–4325, 1980.
Nagao RT, Shah DM, Eckenrode VK, Meagher RB: Multigene family of actin-related sequences isolated from a soybean genomic library. DNA 2:1–9, 1981.
Palmer JD, Shields CR, Cohen CB, Orton TJ: An unusual mitochondrial DNA plasmid in the genusBrassica. Nature 301:725–728, 1983.
Pelletier G, Primard C, Vedel F, Chetrit P, Remy R, Rousselle P, Renard M: Intergeneric cytoplasmic hybridization in Cruciferea by protoplast fusion. MGG 191:244–250, 1983.
Pring DR, Levings CS III: Heterogeneity of maize cytoplasmic genomes among male sterile cytoplasms. Genetics 89:121–136, 1978.
Thompson RD, Kemble RJ, Flavell RB: Variation in mitochondrial DNA organization between normal and male-sterile cytoplasms of maize. Nucleic Acids Res 8:1999–2008, 1980.
Umbeck P, Gengenbach B: Reversion of male-sterile T cytoplasm maize to male fertility in tissue culture. Crop Sci 23:584–588, 1983.
Vieira J, Messing J: The pUC plasmids, and M13mp7-derived system for insertion mutagenesis and sequencing with synthetic universal primers. Gene 19:259–268, 1982.
Warmke HE, Lees SLJ: Mitochondrial degeneration in T cytoplasmic male-sterile corn anthers. J Hered 68:213–22, 1977.
Watraud, LS, Hooker AL, Koeppe DE: The effects of nuclear restorer genes of Texas male-sterile cytoplasm on host response toHelminthosporium mydis Race T. Phytopathology 65:178–182, 1975.
Zelcer A, Aviv D, Galun E: Interspecific transfer of cytoplasmic male sterility by fusion between protoplasts of normalNicotiana sylvestris and X-ray irradiated protoplasts of male-sterileN. tabacum. Z Pflanzenphysiol 90:397–407, 1978.
Author information
Authors and Affiliations
Rights and permissions
About this article
Cite this article
Boeshore, M.L., Hanson, M.R. & Izhar, S. A variant mitochondrial DNA arrangement specific toPetunia stable sterile somatic hybrids. Plant Mol Biol 4, 125–132 (1985). https://doi.org/10.1007/BF02418759
Received:
Revised:
Accepted:
Issue Date:
DOI: https://doi.org/10.1007/BF02418759