Summary
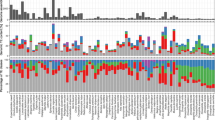
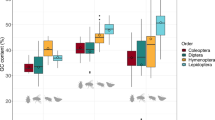
An electrophoretic analysis of histone H1 and its fragments was carried out for several orders of insects. A total of more than 500 histone H1 variants were examined. For some of them a study of general molecular structure was performed by the method of incomplete succinylation. The molecular length of the fragment containing the C-terminal domain presumably responsible for chromatin condensation was found to be highly variable. The variance of the logarithm of the electrophoretic mobility of H1, which reflects its molecular length, was estimated for seven insect orders. There was no relationship between this variance and the evolutionary age of an order. On the other hand, the variance turned out to correlate strongly with the recent species number in the order, indicating that the accumulation of variation in H1 molecular length was in line with the general intensity of adaptive processes in the orders. This result seems to provide evidence for an adaptive mode of the evolution of the molecular length of H1. The possible role of H1 variability in adaptive evolution is discussed.
Article PDF
Similar content being viewed by others
Avoid common mistakes on your manuscript.
References
Allan J, Hartman PG, Crane-Robinson C, Aviles FX (1980) The structure of histone H1 and its location in chromatin. Nature 288:675–679
Berdnikov VA, Gorel' FL, Zlochevskii VA (1973) Histones of someInvertebrata. Biokhimia 38:1208–1214 (in Russian)
Berdnikov VA, Gorel' FL (1975) A study of ratios between histone fractions. Mol Biol (USSR) 9:699–705 (in Russian)
Brown DD (1984) The role of stable complexes that repress and activate eukaryotic genes. Cell 37:359–385
Bucot AJ (1975) Evolution and extinction rate controls. Elsevier, Amsterdam
Cisne JL (1974) Evolution of the world fauna of aquatic free living arthropods. Evolution 28:337–366
Coen E, Strachan T, Dover G (1982) Dynamics of concerted evolution of ribosomal RNA and histone gene families in themelanogaster species group ofDrosophila. J Mol Biol 158:17–35
Dover GA (1989) Slips, strings and species. TIG 5:100–102
Feller W (1970) An introduction to probability theory and its applications (third edition). J Wiley & Sons, New York
Giller PS (1984) Community structure and the niche. Chapman and Hall, London, New York
Hohmann P, Tobey RA, Gurley LR (1976) Phosphorylation of distinct regions of f1 histone. Relationship to the cell cycle. J Biol Chem 251:3685
Isengerg I (1979) Histones. Annu Rev Biochem 48:159–191
Johns EW (1964) Studies of histones. 7. Preparative methods for histone fractions from calf thymus. Biochem J 92:55–59
Kimura M (1983) The neutral theory of molecular evolution. Cambridge University Press, Cambridge
Murphy JT, Blumenfeld M (1986) Nucleotide sequence of aDrosophila melanogaster H1 histone gene. Nucl Acids Res 14:5563–5564
Odum EP (1983) Basic ecology. Saunders College Publishing, Philadelphia
Panyim S, Chalkley R (1969) High resolution in acrilamide gel electrophoresis of histones. Arch Biochem Biophys 130:336–346
Reeves R (1984) Transcriptionally active chromatin. Biochim Biophys Acta 782:343–393
Rodendorf BB, Rasnitsyn AP (eds) (1980) Historical development of theInsecta class. Nauka Moscow (in Russian)
Rodin SN, Berdnikov VA, Zharkikh AA (1985) A model of genome size dynamics during speciation. Biom J 27:807–821
Rozov SM, Berdnikov VA (1982) Determination of lysine residue number, positive charge and molecular lengths of histones H1 and H5 by an incomplete succinylation method. Biokhimia 47:1378–1385 (in Russian)
Sherod D, Johnson G, Chalkley R (1974) Studies on the heterogeneity of lysine-rich histones in dividing cells. J Biol Chem 249:3923–3931
Stanley SM (1975) A theory of evolution above the species level. Proc Natl Acad Sci USA 72:646–650
Thoma F, Koller T, Klug A (1979) Involvement of histone H1 in the organization of the nucleosome and of the salt-dependent superstructures of chromatin. J Cell Biol 83:403–421
Von Holt C, Strickland WM, Brandt WF, Strickland MS (1979) More histone structures. FEBS Lett 100:201–218
Weintraub H (1985) Assembly and propagation of repressed and derepressed chromatin states. Cell 42:705–711
Wells DE, McBridle CA (1989) Comprehensive compilation and alignment of histones and histone genes. Nucl Acid Res 17:Seq Suppl r311–346
Wilson AC, Maxon LR, Sarich VM (1975) Two types of molecular evolution. Proc Natl Acad Sci USA 72:5061
Author information
Authors and Affiliations
Rights and permissions
About this article
Cite this article
Berdnikov, V.A., Rozov, S.M., Temnykh, S.V. et al. Adaptive nature of interspecies variation of histone H1 in insects. J Mol Evol 36, 497–507 (1993). https://doi.org/10.1007/BF02406725
Received:
Revised:
Accepted:
Issue Date:
DOI: https://doi.org/10.1007/BF02406725