Abstract
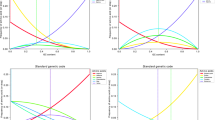
Since base composition of translational stop codons (TAG, TAA, and TGA) is biased toward a low G+C content, a differential density for these termination signals is expected in random DNA sequences of different base compositions. The expected length of reading frames (DNA segments of sense codons flanked by in-phase stop codons) in random sequences is thus a function of GC content. The analysis of DNA sequences from several genome databases stratified according to GC content reveals that the longest coding sequences—exons in vertebrates and genes in prokaryotes—are GC-rich, while the shortest ones are GC-poor. Exon lengthening in GC-rich vertebrate regions does not result, however, in longer vertebrate proteins, perhaps because of the lower number of exons in the genes located in these regions. The effects on coding-sequence lengths constitute a new evolutionary meaning for compositional variations in DNA GC content.
Article PDF
Similar content being viewed by others
Avoid common mistakes on your manuscript.
References
Bernardi G (1989) The isochore organization of the human genome. Annu Rev Genet 23:637–661
Bernardi G (1993) The isochore organization of the human genome and its evolutionary history—a review. Gene 135:57–66
Bernardi G (1995) The human genome: organization and evolutionary story. Annu Rev Genet 29:445–476
Bernardi G, Olofsson B, Filipski J, Zerial M, Salinas J, Cuny G, Meunier-Rotival M, Rodier F (1985) The mosaic genome of warm-blooded vertebrates. Science 228:953–958
Blake C (1983) Exons—present from the beginning? Nature 306:535–537
Blake C (1985) Exons and the evolution of proteins. Int Rev Cytol 93:149–185
Boldögkoi Z, Murvai J, Fodor I (1995) G and C accumulation at silent positions of codons produces additional ORFs. Trends Genet 11: 125–126
Cebrat S, Dudek MR (1996) Generation of overlapping reading frames. Trends Genet 12:12
D'Onofrio G, Bernardi G (1992) A universal compositional correlation among codon positions. Gene 110:81–88
Duret L, Mouchiroud D, Gautier C (1995) Statistical analysis of vertebrate sequences reveals that long genes are scarce in GC-rich isochores. J Mol Evol 40:308–317
Fleischmann RD et al. (1995) Whole-genome random sequencing and assembly ofHaemophilus influenzae Rd. Science 269:496–512
Fraser CM et al. (1995) The minimal gene complement ofMycoplasma genitalium. Science 270:397–403
Guigó R, Fickett JW (1995) Distinctive sequence features in protein coding, genic non-coding, and intergenic human DNA. J Mol Biol 253:51–60
Hawkins JD (1988) A survey on imron and exon lengths. Nucleic Acids Res 16:9893–9908
Höglund M, Säll T, Röhme D (1990) On the origin of coding sequences from random open reading frames. J Mol Evol 30:104–108
Holland SK, Blake CCF (1990) Proteins, exons, and molecular evolution. In: Stone EM, Schwartz RJ (eds) Intervening sequences in evolution and development. Oxford University Press, New York, p 32
Holmquist GP (1989) Evolution of chromosome bands: molecular ecology of noncoding DNA. J Mol Evol 28:469–486
Hughes AL, Hughes MK (1995) Small genomes for better flyers. Nature 377:391
Merino E, Balbás P, Puente JL, Bolivar F (1994) Antisense overlapping open reading frames in genes from bacteria to humans. Nucleic Acids Res 22:1903–1908
Naora H, Miyahara K, Curnow RN (1987) Origin of non coding DNA sequences: molecular fossils of genome evolution. Proc Natl Acad Sci USA 84:6195–6199
Nomura M, Sor F, Yamagishi M, Lawson M (1987) Heterogeneity of GC content within a single bacterial genome and its implications for evolution. Cold Spring Harb Symp Quant Biol 52:658–663
Perrière G, Gouy M, Gojobori T (1994) NRSub: a non-redundant data base for theBacillus subtilis genome. Nucleic Acids Res 22:5525–5529
Poole ES, Brown CM, Tate WP (1995) The identity of the base following the stop codon determines the efficiency ofin vitro translational termination inEscherichia coli. EMBO J 14:151–158
Seidel HM, Pompliano DL, Knowles JR (1992) Exons as microgenes? Science 257:1489–1490
Senapathy P (1986) Origin of eukaryotic introns: a hypothesis, based on codon distribution statistics in genes, and its implications. Proc Natl Acad Sci USA 83:2133–2137
Senapathy P (1988) Possible evolution of splice-junction signals in eukaryotic genes from stop codons. Proc Natl Acad Sci USA 85: 1129–1133
Senapathy P (1995) Introns and the origin of protein-coding genes. Science 268:1366–1367
Senapathy P, Shapiro MB, Harris NL (1990) Splice junctions, branch point sites, and exons: sequence statistics, identification, and applications to genome project. Methods Enzymol 183:252–278
Sharp PM, Burgess CJ, Lloyd AT, Mitchell KJ (1992) Selective use of termination codons and variations in codon choice. In: Hatfield DL, Lee BL Pirtle RM (eds) Transfer RNA in protein synthesis. CRC Press, Boca Raton, pp 398–425
Smith MW (1988) Structure of vertebrate genes: a statistical analysis implicating selection. J Mol Evol 27:45–55
Stoehr PJ, Cameron ON (1991) The EMBL data library. Nucleic Acids Res (Suppl) 19:2227–2230
Stoltzfus A, Spencer DF, Zuker M, Logsdon JM, Doolittle WF (1995) Introns and the origin of protein-coding genes (response). Science 268:1367–1369
Sueoka N (1992) Directional mutation pressure, selection constraints, and genetic equilibria. J Mol Evol 34:95–114
Traut TW (1988) Do exons code for structural or functional units in proteins? Proc Natl Acad Sci USA 85:2944–2948
Wahl R, Rice P, Rice CM, Kröger M (1994) ECD—a totally integrated database ofEscherichia coli K12. Nucleic Acids Res 22:3450–3455
White SH, Jacobs RE (1993) The evolution of proteins from random amino acid sequences. 1. Evidence from the lengthwise distribution of amino acids in modem protein sequences. J Mol Evol 36:79–95
Author information
Authors and Affiliations
Additional information
Correspondence to: J. L. Oliver
Rights and permissions
About this article
Cite this article
Oliver, J.L., Marín, A. A relationship between GC content and coding-sequence length. J Mol Evol 43, 216–223 (1996). https://doi.org/10.1007/BF02338829
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1007/BF02338829