Abstract
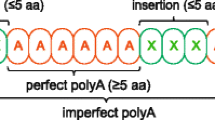
Nucleotide sequence analysis has demonstrated that interspecific size variation in the YP2 yolk protein among HawaiianDrosophila is due to in-frame insertions and deletions in two repetitive segments of the coding region of the Yp2 gene. Sequence comparisons of the complex repetitive region close to the 5′ end of this gene across 34 endemic Hawaiian taxa revealed five length morphs, spanning a length difference of 21 nucleotides (nt). A phylogenetic character reconstruction of the length mutations on an independently derived molecular phylogeny showed clade-specific length variants arising from six ancient events: two identical insertions of 6 nt, and four deletions, one of 6 nt, one of 12 nt, and two identical but independent deletions of 15 nt. These mutations can be attributed to replication slippage with nontandem trinucleotide repeats playing a major role in the slipped-strand mispairing. Geographic analysis suggests that the 15 nt deletion which distinguishes theplanitibia subgroup from thecyrtoloma subgroup occurred on Oahu about 3 million years ago. The homoplasies observed caution against relying too heavily on nucleotide insertions/deletions for phylogenetic inference. In contrast to the extensive repeat polymorphisms within otherDrosophila and the human species, the more complex 5′Yp2 repetitive region analyzed here appears to lack polymorphism among HawaiianDrosophila, perhaps due to founder effects, low population sizes, and hitchhiking effects of selection on the immediately adjacent 5′ region.
Article PDF
Similar content being viewed by others
Avoid common mistakes on your manuscript.
References
Bownes M (1982) Hormonal and genetic regulation of vitellogenesis inDrosophila. Q Rev Biol 57:247–274
Carson HL (1992) Inversions in HawaiianDrosophila. In: Krimbas CB, Powell JR (eds) Drosophila inversion polymorphism. CRC Press, Boca Raton, pp 407–439
Carson HL, Clague DA (1995) Geology and biogeography of the Hawaiian Islands. In: Wagner WL, Funk VA (eds) Hawaiian biogeography. Evolution on a hot spot Archipelago. Smithsonian Institution Press, Washington, DC, pp 14–29
Carson HL, Kaneshiro KY (1976) Drosophila of Hawaii: systematics and ecological genetics. Annu Rev Ecol Syst 7:311–345
Carson HL, Templeton AR (1984) Genetic revolutions in relation to speciation phenomena: the founding of new populations. Annu Rev Ecol Syst 15:97–131
Carson HL, Lockwood JP, Craddock EM (1990) Extinction and recolonization of local populations on a growing shield volcano. Proc Natl Acad Sci USA 87:7055–7057
Caskey CT, Pizzuti A, Fu Y-H, Fenwick RG, Nelson DL (1992) Triplet repeat mutations in human disease. Science 256:784–789
Costa R, Peixoto AA, Thackeray JR, Dalgleish R, Kyriacou CP (1991) Length polymorphism in the threonine-glycine encoding repeat region of the period gene inDrosophila. J Mol Evol 32238–246
Craddock EM, Kambysellis MP (1990) Vitellogenin protein diversity in the HawaiianDrosophila. Biochem Genet 28:415–432
Donoghue MJ, Olmstead RG, Smith JF, Palmer JD (1992) Phylogenetic relationships of Dipsacales based on rbcL sequences. Ann Mo Bot Gardens 79:333–345
Felsenstein J (1985) Confidence limits on phylogenies: an approach using the bootstrap. Evolution 39:783–791
Hey J, Kliman RM (1993) Population genetics and phylogenetics of DNA sequence variation at multiple loci within theDrosophila melanogaster species complex. Mol Biol Evol 10:804–822
Higgins DG, Bleasby AJ, Fuchs R (1992) CLUSTAL V: improved software for multiple sequence alignment. Comput Appl Biosci 8:189–191
Hung M-C, Wensink PC (1983) Sequence and structure conservation in yolk proteins and their genes. J Mol Biol 164:481–492
Huntington's Disease Collaborative Research Group (1993) A novel gene containing a trinucleotide repeat that is expanded and unstable on Huntington's disease chromosomes. Cell 72:971–983
Jeffreys AJ, Wilson V, Them SL (1985) Hypervariable “minisatellite’ regions in human DNA. Nature 314:67–73
Jones CW, Kafatos FC (1982) Accepted mutations in a gene family: evolutionary diversification of duplicated DNA. J Mot Evol 19:87–103
Kambysellis MP, Ho K-F, Craddock EM, Piano F, Parisi M, Cohen J (1995) Pattern of ecological shifts in the diversification of HawaiianDrosophila inferred from a molecular phylogeny. Curr Biol 5:1129–1139
Kaplan NL, Hudson RR, Langley CH (1989) The “hitchhiking effect” revisited. Genetics 123:887–899
La Spada AR, Wilson EM, Lubahn DB, Harding AE, Fischbeck KH (1991) Androgen receptor gene mutations in X-linked spinal and bulbar muscular atrophy. Nature 352:77–79
Levinson G, Gutman GA (1987) Slipped-strand mispairing: a major mechanism for DNA sequence evolution. Mot Biol Evol 4(3):203–221
Litt M, Luty JA (1989) A hypervariable microsatellite revealed by in vitro amplification of a dinucleotide repeat within the cardiac muscle actin gene. Am J Hum Genet 44:397–401
Logan SK, Garabedian MJ, Wensink PC (1989) DNA regions that regulate the ovarian transcriptional specificity ofDrosophila yolk protein genes. Genes Dev 3:1453–1461
Maddison WP, Maddison DR (1992) MacClade. Analysis of phylogeny and character evolution, version 3. Sinauer, Sunderland, MA
Martinez-Cruzado JC (1990) Evolution of the amosomal chorion cluster inDrosophila. IV. The HawaiianDrosophila: rapid protein evolution and constancy in the rate of DNA divergence. J Mol Evol 31:402–423
Maynard Smith J, Haigh J (1974) The hitch-hiking effect of a favourable gene. Genet Res 23:23–35
Nakamura Y, Leppert M, O'Connell P, Wolff R, Holm T, Culver M, Martin C, Fujimoto E, Hoff M, Kumlin E, WhiteR (1987) Variable number of tandem repeat (VNTR) markers for human gene mapping. Science 235:1616–1622
Parisi M (1994) Nucleotide sequence of theDrosophila grimshawi Yolk protein genes and analysis of the regulation of theYp1–Yp2 gene cluster. PhD thesis, New York University, New York
Reue K, Leete TH (1991) Genetic variation in mouse apolipoprotein A-IV due to insertion and deletion in a region of tandem repeats. J Biol Chem 266:12715–12721
Saiki R, Scharf S, Faloona F, Mullis K, Horn G, Erlich H, Arnheim N (1985) Enzymatic amplification of β-globin genomic sequences and restriction site analysis for diagnosis of sickle-cell anemia. Science 230:1350–1354
Smith GP (1976) Evolution of repeated DNA sequences by unequal crossover. Science 191:528–535
Streisinger G, Okada Y, Emrich J, Newton J, Tsugita A, Terzaghi E, Inouye M (1966) Frameshift mutations and the genetic code. Cold Spring Harb Symp Quant Biol 31:77–84
Swofford DL (1993) Phylogenetic analysis using parsimony (PAUP), version 3.1.1. University of Illinois, Champaign, IL
Tautz D (1989) Hypervariability of simple sequences as a general source for polymorphic DNA markers. Nucleic Acids Res 17:6463–6471
Tautz D, Trick M, Dover GA (1986) Cryptic simplicity in DNA is a major source of genetic variation. Nature 322:652–656
Weber JL, May PE (1989) Abundant class of human DNA polymorphisms which can be typed using the polymerase chain reaction. Am J Hum Genet 44:388–396
Wharton KA, Yedvobnick B, Finnerty VG, Artavanis-Tsakonas S (1985)opa: a novel family of transcribed repeats shared by theNotch locus and other developmentally regulated loci inD. melanogaster. Cell 40:55–62
Author information
Authors and Affiliations
Additional information
Correspondence to: M.P. Kambysellis
Rights and permissions
About this article
Cite this article
Ho, KF., Craddock, E.M., Piano, F. et al. Phylogenetic analysis of DNA length mutations in a repetitive region of the Hawaiiandrosophila yolk protein geneYp2 . J Mol Evol 43, 116–124 (1996). https://doi.org/10.1007/BF02337356
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1007/BF02337356