Summary
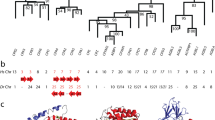
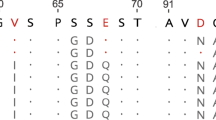
The amino acid sequences of the pancreatic ribonuclease from capybara (Hydrochoerus hydrochaeris) and cuis (Galea musteloides) were determined. Both species belong to the same superfamily of the hystricomorph rodents as the guinea-pig. In guineapig pancreas two ribonucleases are present as a result of a rencent gene duplication, but in capybara and cuis pancreas only one single ribonuclease has been found. A most parsimonious tree of ribonucleases indicates that the gene duplication leading to both guinea-pig ribonucleases occurred before the divergence of guineapig from capybara and cuis. This would mean that changes in expression of the ribonuclease genes have occurred in these taxa.
Cuis and capybara ribonuclease have no Asn-X-Ser/Thr sequences and are carbohydrate-free proteins. Capybara ribonculease has leucine at position 114, a position occupied by proline in thecis-configuration in bovine pancreatic ribonuclease.
Article PDF
Similar content being viewed by others
Avoid common mistakes on your manuscript.
References
Beintema JJ (1980) Ribonuclease variation within the species. In: Peeters H (ed) Proc. 28th coll. protides biol. fluids. Pergamon Press, Oxford, pp 139–141
Beintema JJ, Lenstra JA (1982) Evolution of mammalian pancreatic ribonucleases. In: Goodman M (ed) Macromolecular sequences in systematic and evolutionary biology. Plenum, New York, pp 43–73
Beintema JJ, Gaastra W, Lenstra JA, Welling GW, Fitch WM (1977) The molecular evolution of pancreatic ribonuclease. J Mol Evol 10:49–71
Beintema JJ, Lenstra JA, Fitch WM (1980) The molecular evolution of mammalian pancreatic ribonuclease. In: Peeters H (ed) Proc. 28th coll. protides biol. fluids. Pergamon Press, Oxford, pp 133–137
Beintema JJ, Knol G, Martena B (1982) The primary structures of pancreatic ribonucleases from African porcupine and casiragua, two hystricomorph rodent species. Biochim Biophys Acta 705:102–110
Eaker DL, King TP, Craig LC (1965) Des-lysyl glutamyl and deslysyl pyroglutamyl ribonucleases. II Structural studies. Biochemistry 4:1479–1486
Fitch WM (1971) Toward defining the course of evolution: minimum change for a specific tree topology. Syst Zool 20:406–416
Gaastra W, Welling GW, Beintema JJ (1978) The amino acid sequence of kangaroo pancreatic ribonuclease. Eur J Biochem 86:209–217
Goodman M, Czelusniak J, Moore GW, Romero-Herrara AE, Matsuda G (1979) Fitting the gene lineage into its species lineage, A parsimony strategy illustrated by cladograms constructed from globin sequences. Syst Zool 28:132–163
Gray WR (1967) Dansyl chloride procedure. Methods Enzymol 11:139–151
Hartley BS (1970) Strategy and tactics in protein chemistry. First BDH lecture. Biochem J 119:805–822
Horuk R, Goodwin P, O'Connor K, Neville RWJ, Lazarus NR, Stone D (1979) Evolutionary change in the insulin receptors of hystricomorph rodents. Nature 279:439–440
Horuk R, Blundell TL, Lazarus NR, Neville RWJ, Stone D, Wollmer A (1980) A monomeric insulin from the porcupine (Hystrix cristata), an old world hystricomorph. Nature 286:822–824
Jekel PA, Sips HJ, Lenstra JA, Beintema JJ (1979) The amino acid sequence of hamster pancreatic ribonuclease. Biochimie 61:827–839
Morris D (1965) The Mammals. Hodder and Stoughton, London
Offord RE (1966) Electrophoretic mobilities of peptides on paper and their use in the determination, of amide groups. Nature 211:591–593
Richards FM, Wyckoff HW (1973) In: Philips DC, Richards FM (eds) Atlas of molecular structures in biology. Vol 1, Ribonuclease-S. Clarendon, Oxford
Schmid FX, Blaschek H (1981) A native-like intermediate on the ribonuclease A folding pathway. 2. Comparison of its properties to native ribonuclease A. Eur J Biochem 114:111–117
Shapira R (1962) A spectrophotometric method for the measurement of ribonuclease activity. Anal Biochem 3:308–320
Suzuki H, Greco L, Parente A, Farina B, La Montagna R, Leone E (1976) Bovine seminal ribonuclease. In: Dayhoff MO (ed) Atlas of protein sequence and structure. Vol 5, Suppl. 2. National Biomedical Research Foundation, Washington DC, p 93
Van den Berg A, Van den Hende-Timmer L, Beintema JJ (1976) Isolation, properties and primary structure of coypu and chinchilla pancreatic ribonuclease. Biochim Biophys Acta 453:400–409
Van den Berg A, Van den Hende-Timmer L, Hofsteenge J, Gaastra W, Beintema JJ (1977) Guinea-pig pancreatic ribonucleases. Isolation, properties, primary structure and glycosidation. Eur J Biochem 75:91–100
Vereijken JM, Hofsteenge, J, Bak HJ, Beintema JJ (1980) The amino acid sequence of the three smallest CNBr peptides from p-hydroxybenzoate hydroxylase fromPseudomonas fluorescens. Eur J Biochem 113:151–157
Vereijken JM, Schwander EH, Soeter NM, Beintema JJ (1982) Limited proteolysis of the 94000-Dalton subunit ofPanulirus interruptus hemocyanin; the carbohydrate attachment site. Eur J Biochem 123:283–289
Welling GW, Leijenaar-van den Berg G, van Dijk B, van den Berg A, Groen G, Gaastra W, Emmens M, Beintema JJ (1975) Evolution of mammalian pancreatic ribonucleases. Biosystems 6:239–245
Wierenga RK, Huizinga JD, Gaastra W, Welling GW, Beintema JJ (1973) Affinity chromatography of porcine pancreatic ribonuclease and reinvestigation of the N-terminal amino acid sequence. FEBS Lett 31:181–185
Author information
Authors and Affiliations
Additional information
Data supplementary to this article are deposited with, and can be obtained from Elsevier Biomedical Press B.V., BBA Data Deposition, P.O. Box 1345, 1000 BH Amsterdam, The Netherlands. Reference should be made to BBA/DD/246. The supplementary information included Tables 3 and 4 (procedures used for purification of peptides from capybara and cuis ribonuclease and charge determinations of peptides at pH 6.5), Tables 5 and 6 (amino acid compositions of peptides) and Figs. 5–9 (gel filtration patterns of CNBr and enzymic digests).
Rights and permissions
About this article
Cite this article
Beintema, J.J., Neuteboom, B. Origin of the duplicated ribonuclease gene in guinea-pig: Comparison of the amino acid sequences with those of two close relatives: Capybara and cuis ribonuclease. J Mol Evol 19, 145–152 (1983). https://doi.org/10.1007/BF02300752
Received:
Revised:
Issue Date:
DOI: https://doi.org/10.1007/BF02300752