Summary
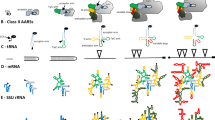
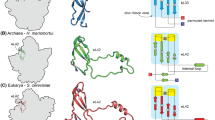
Several sequences flanking the large rRNA genes of several transcripts from extreme thermophiles, extreme halophiles, and methanogens were aligned and analyzed for the presence of common primary and secondary structural features, which would bear on the concept of monphyletic archaebacteria. Few sequences were common to all the archaebacterial transcripts, and these were confined to short regions generally flanking putative double helices. At a secondary structural level, however, in addition to the previously characterized processing stems of the 16S and 23S RNAs, four helices were detected that were common to the archaebacterial transcripts: two in the 16S RNA leader sequence and two in the 16S-23S RNA spacer. Although all of these helices vary in size and form from organism to organism, three of them contain double helical segments that are strongly supported by compensating base changes among the three archaebacterial groups. Three extreme halophiles exhibited two additional helices in their relatively large spacers and a further helix preceding the 5S RNA, which are also supported by compensating base changes. Ribosomal RNA transcripts from eubateria/chloroplasts and eukaryotes were also examined for secondary structural features with locations and forms corresponding to those of the archaebacteria, but none were detected. The analysis provides support for the monophyletic nature of the archaebacteria and reinforces their differences from eubacteria/chloroplasts and eukaryotes.
Article PDF
Similar content being viewed by others
Avoid common mistakes on your manuscript.
References
Achenbach-Richter L, Woese CR (1988) The ribosomal RNA gene spacer in archaebacteria. Syst Appl Microbiol 10:211–214
Brosius J, Dull TJ, Sleeter DD, Noller HF (1981) Gene organization and primary structure of a rRNA operon fromEscherichia coli. J Mol Biol 148:107–127
Fox GE, Stackebrandt E, Hespell RB, Gibson J, Maniloff J, Dyer TA, Wolfe RS, Balch WE, Tanner R, Magrum L, Zablen LB, Blakemore R, Gupta R, Bonen L, Lewis BJ, Stahl DA, Luehrsen KR, Chen KN, Woese CR (1980) The phylogeny of prokaryotes. Science 209:457–463
Green CJ, Stewart GC, Hollis MA, Vold BS, Bott KF (1985) Nucleotide sequence of theBacillus subtilus rRNA operonrrnB. Gene 37:261–266
Hassouna N, Michot B, Bachellerie JP (1984) The complete nucleotide sequence of mouse 28S rRNA gene. Implications for the process of size increase of the large subunit rRNA in higher eukaryotes. Nucleic Acids Res 12:3563–3583
Hui I, Dennis PP (1985) Characterization of the rRNA gene clusters inHalobacterium cutirubrum. J Biol Chem 260:899–906
Jarsch M, Böck A (1985) Sequence of the 23S rRNA gene from the archaebacteriumMethanococcus vannielii: evolutionary and functional implications. Mol Gen Genet 200:305–312
Kjems J, Garrett RA (1987) Novel expression of the rRNA genes in the extreme thermophile and archaebacteriumDesulfurococcus mobilis. EMBO J 6:3521–3530
Kjems J, Leffers H, Garrett RA, Wich G, Leinfelder W, Böck A (1987) Gene organization, transcription signals and processing of the single rRNA operon of the archaebacteriumThermoproteus tenax. Nucleic Acids Res 15:4821–4835
Kjems J, Leffers H, Olesen T, Holz I, Garrett RA (1990) Sequence, organization and transcription of the ribosomal RNA operon and the downstream tRNA and protein genes in the archaebacteriumThermofilum pendens. Syst Appl Microbiol (in press)
Kössel H, Edwards K, Koch W, Langridge P, Schiefermayer E, Schwarz Z, Strittmatter G, Zenke G (1982) Structural and functional analysis of an rRNA operon and its flanking tRNA genes fromZea mays chloroplasts. Nucleic Acids Res Symp 11:117–120
Lake JA (1988) Origin of the eukaryotic nucleus determined by rate-invariant analysis of rRNA sequences. Nature 331:184–186
Lake JA (1989) Origin of the eukaryotic nucleus: eukaryotes and eocytes are genetically related. Can J Microbiol 35:109–118
Lake JA, Hendersen E, Clark MW, Scheinman A, Oakes MI (1986) Mapping evolution with three dimensional ribosome structure. Syst Appl Microbiol 7:131–136
Larsen N, Leffers H, Kjems J, Garrett RA (1986) Evolutionary divergence between the rRNA operons ofHalococcus morrhuae andDesulfurococcus mobilis. Syst Appl Microbiol 7:49–57
Leffers H, Kjems J, Østergaard L, Larsen N, Garrett RA (1987) Evolutionary relationships amongst archaebacteria: a comparative study of 23S rRNAs of a sulfur-dependent extreme themophile, extreme halophile and a thermophilic methanogen. J Mol Biol 195:43–61
Mankin AS, Kagramanova VK (1986) Complete nucleotide sequence of the rRNA operon ofHalobacterium halobium: a secondary structure of the archaebacterial 23S rRNA. Mol Gen Genet 202:152–161
Mankin AS, Teterina NL, Baratova LA, Kagramanova VK (1984) Putative promoter region of rRNA operon from archaebacteriumHalobacterium halobium. Nucleic Acids Res 12:6537–6546
Mankin AS, Skripkin EA, Kagramanova VK (1987) A putative internal promoter in the 16S/23S intergenic spacer of the rRNA operons of archaebacteria and eubacteria. FEBS Lett 219:269–273
Massenet O, Martinez P, Seyer P, Briat J-F (1987) Sequence organization of the chloroplast ribosomal spacer ofSpinacia oleracea including the 3′-end of the 16S RNA and the 5′-end of the 23S RNA. Plant Mol Biol 10:53–63
Neumann H, Gierl A, Tu J, Leibrock J, Staiger D, Zillig W (1983) Organization of the genes for ribosomal RNA in archaebacteria. Mol Gen Genet 192:66–72
Olsen GJ, Pace NR, Nuel M, Kaine BP, Gupta R, Woese CR (1985) Sequence of the 16S rRNA gene from the thermophilic archaebacteriumSulfolobus solfataricus and its evolutionary implications. J Mol Evol 22:301–307
Østergaard L, Larsen N, Leffers H, Kjems J, Garrett RA (1987) A rRNA operon and its flanking region from the archaebacteriumMethanobacterium thermoautotrophicum, Marburg strain: transcirption signals, rRNA structure, and evolutionary implications. Syst Appl Microbiol 9:199–209
Planta RJ, Klootwijk J, Raué HA, Brand RC, Veldman GM, Stiekema WJ, de Jonge P (1980) Transcription and processing of ribosomal RNA in yeast and bacilli. In: Osawa S, Ozeki H, Uchida H, Yura T (eds) Genetics and evolution of RNA polymerase tRNA ribosomes, Elsevier, Amsterdam, pp 451–470
Ree HK, Cas K, Thurlow D, Zimmermann R (1989) Structure and organization of the 16S rRNA gene from the archaebacteriumThermoplasma acidophilum. Can J Microbiol 35:124–133
Reiter WD, Palm P, Voos W, Kaniecki J, Grampp B, Schulz W, Zillig W (1987) Putative promoter elements for the rRNA genes of the thermoacidophilic archaebacteriumSulfolobus sp. strain B12. Nucleic Acids Res 15:5581–5595
Staden R (1982) An interactive graphics program for comparing and aligning nucleic acid and amino acid sequences (DIA-GON). Nucleic Acids Res 10:2650–2696
Wich G, Leinfelder W, Böck A (1987) Genes for stable RNA in the extreme thermophileThermoproteus tenax: introns and transcription signals. EMBO J 6:523–528
Woese CR, Olsen G (1986) Archaebacterial phylogeny: perspectives on the urkingdoms. Syst Appl Microbiol 7:161–177
Author information
Authors and Affiliations
Rights and permissions
About this article
Cite this article
Kjems, J., Garrett, R.A. Secondary structural elements exclusive to the sequences flanking ribosomal RNAs lend support to the monophyletic nature of the archaebacteria. J Mol Evol 31, 25–32 (1990). https://doi.org/10.1007/BF02101789
Received:
Revised:
Issue Date:
DOI: https://doi.org/10.1007/BF02101789