Summary
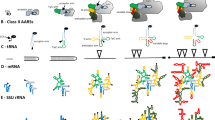
We have determined the nucleotide sequence of the 26S large subunit (LSU) rRNA genes for twoTetrahymena species,T. thermophila andT. pyriformis. The inferred rRNA sequences are presented in their most probable secondary structures based on compensatory mutations, energy, and conservation criteria. The majority of the nucleotide changes between the twoTetrahymena LSU rRNAs and the positions of a relatively large deletion and of the processing cleavage sites resulting in the generation of the hidden break are all located within the so-called divergent domains or expansion segments. These are regions within the common core of secondary structure where expansions have taken place during the evolution of the rRNA of higher eukaryotes.
The dispensable nature of some of the expansion segments has been taken as evidence of their non-functionality. However, our data show that a considerable selective constraint has operated to presesrve the secondary structure of these segments. Especially in the case of the D2 and D8 segments, the presence of a considerable number of compensatory base changes suggests that the secondary structure of these regions is of functional importance. Alternatively, these expansion segments may have maintained characteristic folding patterns because only such structures are being tolerated within otherwise functionally important regions.
Article PDF
Similar content being viewed by others
Avoid common mistakes on your manuscript.
References
Baroin A, Perasso R, Qu LH, Brugerolle G, Bachellerie J-P, Adoutte A (1988) Partial phylogeny of unicellular eukaryotes based on rapid sequencing of a portion of the 28S ribosomal RNA. Proc Natl Acad Sci USA 85:3474–3478
Boer PH, Gray MW (1988) Scrambled ribosomal RNA gene pieces inChlamydomonas reinhardtii mitochondrial DNA. Cell 55:399–411
Cech TR (1986) Ribosomal RNA gene expression inTetrahymena: transcription and RNA splicing. In: Gall JG (ed) The molecular biology of ciliated protozoa. Academic Press, New York, pp 203–225
Clark CG (1987) On the evolution of ribosomal RNA. J Mol Evol 25:343–350
Din N, Engberg J, Gall JG (1982) The nucleotide sequence at the transcription termination site of the ribosomal RNA gene inTetrahymena thermophila. Nucleic Acids Res 10:1503–1513
Eckert WA, Kaffenberger W, Krohne G, Franke WW (1978) Introduction of hidden breaks during rRNA maturation and ageing inTetrahymena pyriformis. Eur J Biochem 87:607–616
Engberg J (1985) The ribosomal RNA genes ofTetrahymena. Structure and function. Eur J Cell Biol 36:133–151
Engberg J, Din N, Eckert WA, Kaffenberger W, Pearlman RE (1980) Detailed transcription map of extrachromosomal ribosomal RNA genes inTetrahymena thermophila. J Mol Biol 142:289–313
Geliebter J (1987) Dideoxy nucleotide sequencing of RNA and uncloned cDNA. Focus 9:5–8
Gerbi SA (1985) Evolution of ribosomal RNA. In: MacIntyre RJ (ed) Molecular evolutionary genetics. Plenum, New York, pp 419–517
Gouy M, Li W-H (1989) Molecular phylogeny of kingdoms Animalia, Plantae, and Fungi. Mol Biol Evol 6:109–122
Gutell RR, Fox EF (1988) A compilation of large subunit RNA sequences presented in a structural format. Nucleic Acids Res [suppl] 16:r175-r269
Hancock JM, Dover GA (1988) Molecular coevolution among cryptically simple expansion segments of eukaryotic 26S/28S rRNAs. Mol Biol Evol 5:377–391
Hancock JM, Tautz D, Dover GA (1988) Evolution of the secondary structures and compensatory mutations of the ribosomal RNAs ofDrosophila melanogaster. Mol Biol Evol 5:393–414
Heinonen TYK, Schnarre MN, Young PG, Gray MW (1987) Rearranged coding segments, separated by a transfer RNA gene, specify the two parts of a discontinuous large subunit ribosomal RNA inTetrahymena pyriformis mitochondria. J Biol Chem 262:2879–2887
Higashinakagawa T, Saiga H, Shintani N, Narushima-Iio M, Mita T (1981) Localization of putative transcription initiation site on the cloned rDNA fragment ofTetrahymena pyriformis. Nucleic Acids Res 9:5905–5916
Karrer KM (1986) The nuclear DNA's af holotrichous ciliates. In: Gall JG (ed) The molecular biology of ciliated protozoa. Academic Press, New York, pp 85–110
Leffers H, Kjems J, Østergaard L, Lassen N, Garrett RA (1987) Evolutionary relationships amongst Archebacteria. J Mol Biol 195:43–61
Lenaers G, Nielsen H, Engberg J, Herzog M (1988) The secondary structure of large-subunit rRNA divergent domains, a marker for protist evolution. BioSystems 21:215–222
Lenaers G, Maroteaux L, Michot B, Herzog M (1989) Dinoflagellates in evolution. A molecular phylogeny analysis of large subunit ribosomal RNA. J Mol Evol 29:40–51
Michot B, Bachellerie J-P (1987) Comparisons of large subunit rRNA's reveal some eukaryotic-specific elements of secondary structure. Biochimie 69:11–23
Michot B, Hassouna N, Bachellerie J-P (1984) Secondary structure of mouse 28S rRNA and general model for the folding of the large rRNA in eukaryotes. Nucleic Acids Res 12:4259–4279
Niles EG, Cunningham K, Jain R (1981) Structure of theTetrahymena pyriformis rRNA gene. J Biol Chem 256:12857–12860
Noller HF, Asire M, Barta A, Douthwaite S, Goldstein T, Gutell RR, Moazed D, Normaly J, Prince JB, Stern S, Triman K, Turner S, Van Stolk B, Weaton V, Weises B, Woese CR (1986) Studies on the structure and function of ribosomal RNA. In: Hardesty B, Kramer G (eds) Structure, function and genetics of ribosomes. Springer Verlag, Berlin, pp 143–163
Pedersen N, Hellung-Larsen P, Engberg J (1985) Small nuclear RNA's in the ciliateTetrahymena. Nucleic Acids Res 13:4203–4224
Preparata RM, Meyer EB, Preparata FP, Simon EM, Vossbrinck CR, Nanney DL (1989) The ribosomal phylogenies of the tetrahymenine ciliates. J Mol Evol 28:427–441
Raué HA, Klootwijk J, Musters W (1988) Evolutionary conservation of structure and function of high molecular weight ribosomal RNA. Progr Biophys Mol Biol 51:77–129
Sogin ML, Elwood HJ, Gunderson JH (1986a) Evolutionary diversity of eukaryotic small-subunit rRNA genes. Proc Natl Acad Sci USA 83:1383–1387
Sogin ML, Ingold A, Karlok M, Nielsen H, Engberg J (1986b) Phylogenetic evidence for the acquisition of ribosomal RNA introns subsequent to the divergence of some of the majorTetrahymena groups. EMBO J 5:3625–3630
Sollner-Webb B, Reeder R (1979) The nucleotide sequence of the initiation and termination sites for ribosomal RNA transcription inX. laevis. Cell 18:485–499
Spencer DF, Collings JC, Schnarre MN, Gray MW (1987) Multiple spacer sequences in the nuclear large subunit ribosomal RNA gene ofCrithidia fasciculata. EMBO J 6:1063–1071
Sweeney R, Yao M-C (1989) Identifying functional regions of rRNA by insertion mutagenesis and complete gene replacement inTetrahymena thermophild. EMBO J 8:933–938
Ware VC, Renkawitz R, Gerbi SA (1985) rRNA processing: removal of only nineteen bases at the gap between 28 α and 28 β rRNAs inSciara coprophila. Nucleic Acids Res 13:3581–3597
Wild MA, Sommer R (1980) Sequence of a ribosomal RNA gene intron fromTetrahymena. Nature 283:693–694
Woese CR (1987) Bacterial evolution. Microbiol Rev 51:221–271
Yao M-C (1986) Amplification of ribosomal RNA genes. In: Gall JG (ed) The molecular biology of ciliated protozoa. Academic Press, New York, pp 179–201
Zuker M, Stiegler P (1981) Optimal computer folding of large RNA sequences using thermodynamics and auxiliary information. Nucleic Acids Res 9:133–148
Author information
Authors and Affiliations
Rights and permissions
About this article
Cite this article
Engberg, J., Nielsen, H., Lenaers, G. et al. Comparison of primary and secondary 26S rRNA structures in twoTetrahymena species: Evidence for a strong evolutionary and structural constraint in expansion segments. J Mol Evol 30, 514–521 (1990). https://doi.org/10.1007/BF02101107
Received:
Revised:
Issue Date:
DOI: https://doi.org/10.1007/BF02101107