Summary
The evolution of the main regulatory region (D-loop) of the mammalian mitochondrial genome was analyzed by comparing the sequences of eight mammalian species: human, common chimpanzee, pygmy chimpanzee, dolphin, cow, rat, mouse, and rabbit. The best alignment of the sequences was obtained by optimization of the sequence similarities common to all these species.
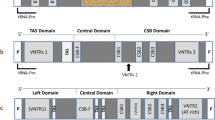
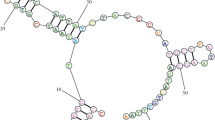
The two peripheral left and right D-loop domains, which contain the main regulatory elements so far discovered, evolved rapidly in a species-specific manner generating heterogeneity in both length and base composition. They are prone to the insertion and deletion of elements and to the generation of short repeats by replication slippage. However, the preservation of some sequence blocks and similar cloverleaf-like structures in these regions, indicates a basic similarity in the regulatory mechanisms of the mitochondrial genome in all mammalian species.
We found, particularly in the right domain, significant similarities to the telomeric sequences of the mitochondrial (mt) and nuclear DNA ofTetrahymena thermophila. These sequences may be interpreted as relics of telomeres present in ancestral linear forms of mtDNA or may simply represent efficient templates of RNA primase-like enzymes.
Due to their peculiar evolution, the two peripheral domains cannot be used to estimate in a quantitative way the genetic distances between mammalian species. On the other hand the central domain, highly conserved during evolution, behaves as a good molecular clock.
Reliable estimates of the times of divergence between closely and distantly related species were obtained from the central domain using a Markov model and assuming nonhomogeneous evolution of nucleotide sites.
Article PDF
Similar content being viewed by others
Avoid common mistakes on your manuscript.
References
Bennet JL, Clayton DA (1990) Efficient site-specific cleavage by RNase MRP requires interaction with two evolutionarily conserved mitochondrial RNA sequences. Mol Cell Biol 10: 2191–2201
Blackburn EH (1990) Telomeres: structure and synthesis. J Mol Biol 265:5919–5921
Brown GG, Gadaleta G, Pepe G, Saccone C, Sbisá E (1986) Structural conservation and variation in the D-loop-containing region of vertebrate mitochondrial DNA. J Mol Biol 192: 503–511
Doda JN, Wright CT, Clayton DA (1981) Elongation of displacement-loop strands in human and mouse mitochondrial DNA is arrested near specific template sequences. Proc Natl Acad Sci USA 78:6116–6120
Dunon-Bluteau DC, Brun GM (1987) Mapping at the nucleotide level ofXenopus laevis mitochondrial D-loop H strand: structural features of the 3′ region. Biochem Int 14:643–645
Easteal S (1987) The rates of nucleotide substitution in the human and rodent lineages: a reply to Li and Wu. Mol Biol Evol 4:78–80
Easteal S (1990) The pattern of mammalian evolution and the relative rate of molecular evolution. Genetics 124:165–173
Fisher RP, Parisi MA, Clayton DA (1989) Flexible recognition of rapidly evolving promoter sequences by mitochondrial transcription factor 1. Genes & Dev 3:2202–2217
Foran DR, Hixson JE, Brown WM (1988) Comparison of ape and human sequences that regulate mitochondrial DNA transcription and D-loop DNA synthesis. Nucleic Acids Res 16: 5841–5861
Gadaleta G, Pepe G, De Candia G, Quagliariello C, Sbisá E, Saccone C (1989) The complete nucleotide sequence of theRattus norvegicus mitochondrial genome: cryptic signals revealed by comparative analysis between vertebrates. J Mol Evol 28:497–516
Holmes EC, Pesole G, Saccone C (1989) Stochastic models of molecular evolution and the estimation of phylogeny and rates of nucleotide substitution in the hominoid primates. J Hum Evol 18:775–794
Li W-H, Wu C-I (1987) Rates of nucleotide substitution are evidently higher in rodents than in man. Mol Biol Evol 4: 74–77
MacKay SD, Olivo PD, Laipis PJ, Hauswirth WW (1986) Template-directed arrest of mammalian mitochondrial DNA synthesis. Mol Cell Biol 6:1261–1267
Mignotte F, Gueride F, Champagne AM, Mounolou JC (1990) Direct repeats in the noncoding region of rabbit mitochondrial DNA: involvement in the generation of intra- and inter-individual heterogeneity. Eur J Biochem 194:561–571
Morin GB, Cech TR (1986) The telomeres of the linear mitochondrial DNA ofTetrahymena thermophila consist of 53 bp tandem repeats. Cell 46:873–883
Murray A (1990) All's well that ends well. Nature 346:797–798
Saccone C, Attimonelli M, Sbisá E (1985) Primary and higher order structural analysis of animal mitochondrial DNA. In: Quagliariello E, Slater EC, Palmieri F, Saccone C, Kroon AM (eds) Achievements and perspectives of mitochondrial research, vol II. Biogenesis. Elsevier, Amsterdam, p 37
Saccone C, Attimonelli M, Sbisá E (1987) Structural elements highly preserved during the evolution of the D-loop-containing region in vertebrate mitochondrial DNA. J Mol Evol 25: 205–211
Saccone C, Lanave C, Pesole G, Preparata G (1990) Influence of base composition on quantitative estimates of gene evolution. Meth Enzymol 183:570–583
Sbisá E, Nardelli M, Tanzariello F, Tullo A, Saccone C (1990) The complete and symmetric transcription of the main noncoding region of rat mitochondrial genome: in vivo mapping of heavy and light transcripts. Curr Genet 17:247–253
Southern SO, Southern PJ, Dizon AE (1988) Molecular characterization of a cloned dolphin mitochondrial genome. J Mol Evol 28:32–42
Walberg MW, Clayton DA (1981) Sequence and properties of the human KB cell and mouse L cell D-loop regions of mitochondrial DNA. Nucleic Acids Res 9:5411–5421
Author information
Authors and Affiliations
Rights and permissions
About this article
Cite this article
Saccone, C., Pesole, G. & Sbisá, E. The main regulatory region of mammalian mitochondrial DNA: Structure-function model and evolutionary pattern. J Mol Evol 33, 83–91 (1991). https://doi.org/10.1007/BF02100199
Issue Date:
DOI: https://doi.org/10.1007/BF02100199