Summary
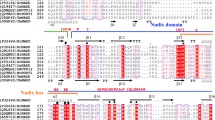
The nucleotide sequences of the two T-DNA-encoded crown gall imino acid dehydrogenases octopine dehydrogenase and nopaline dehydrogenase were compared with each other and with the sequences of other dehydrogenases. A multistep strategy comprising computer sequence analysis and secondary- and antigenic-structure predictions was used. An alignment of octopine and nopaline dehydrogenase was obtained in which a 20-amino-acid N-terminal arm and six fairly long gaps in the C-terminal moiety were introduced. The aligned sequences have identities of 26% at the amino acid level and 38% at the nucleotide level. They appear to contain two domains. The N-terminal coenzyme-binding domains are similar to those of the well-characterized NAD(P) dehydrogenases. Conserved fragments were found in the C-terminal catalytic domains that likely contain essential residues for catalysis. Comparison of the sequences with those of two other 2-keto acid dehydrogenases, lactate and malate dehydrogenase, suggests that as in those enzymes, histidine, aspartic acid, and arginine residues are located at the octopine and nopaline dehydrogenase active sites. The crown gall enzymes could not be classified with any known family of dehydrogenases. Their evolutionary origin remains unknown. However, predictions concerning their internal organization may provide new insight into protein evolution.
Article PDF
Similar content being viewed by others
Avoid common mistakes on your manuscript.
References
Barker RF, Idler KB, Thompson DV, Kemp JD (1983) Nucleotide sequence of the T-DNA region fromAgrobacterium tumefaciens octopine Ti plasmid pTi15955. Plant Mol Biol 2:335–350
Bevan M, Barnes WM, Chilton M-D (1983) Structure and transcription of the nopaline synthase gene region of the T-DNA. Nucleic Acids Res 11:369–385
Birktoft JJ, Banaszak LJ (1983) The presence of a histidine-aspartic acid pair in the active site of 2-hydroxyacid dehydrogenases. X ray refinement of cytoplasmic malate dehydrogenase. J Biol Chem 258:472–482
Blake C (1983) Exons-present from the beginning? Nature 306:535–537
Bränden C-I (1980) Relation between structure and function of α/β proteins. Q Rev Biophys 13:317–338
Chang C-C, Chen C-M (1983) Evidence for the presence of N2-(1,3-dicarboxypropyl)-l-amino acids in crown gall tumors induced byAgrobacterium tumefaciens strains 181 and EU6. FEBS Lett 162:432–435
Chang C-C, Chen C-M, Adams BR, Trost BM (1983) Leucinopine, a characteristic compound of some crown gall tumors. Proc Natl Acad Sci USA 80:3573–3576
Chou PY, Fasman GD (1978) Empirical predictions of protein conformation. Annu Rev Biochem 47:251–276
Dayhoff MO (ed) (1978) Atlas of protein sequence and structure, vol 5, suppl 3. National Biomedical Research Foundation, Washington, DC
De Greve H, Dhaese P, Seurinck J, Lemmers M, Van Montagu M, Schell J (1982) Nucleotide sequence and transcript map of theAgrobacterium tumefaciens Ti plasmid-encoded octopine synthase gene. J Mol Appl Genet 1:499–511
Dennis ES, Gerlach WL, Pryor AJ, Bennetzen JL, Inglis A, Llewellyn D, Sachs MM, Ferl RJ, Peacock WJ (1984) Molecular analysis of the alcohol dehydrogenase (Adhi) gene of maize. Nucleic Acids Res 12:3983–4000
Depicker A, Stachel S, Dhaese P, Zambryski P, Goodman HM (1982) Nopaline synthase: transcript mapping and DNA sequence. J Mol Appl Genet 1:561–573
de Zwaan A, Zurburg W (1981) The formation of strombine in the adductor muscle of the sea musselMytilus edulis L. Marine Biol Lett 2:179–192
Doolittle RF (1981) Similar amino acid sequences: change or common ancestry? Science 214:149–159
Fields JHA, Hochachka PW (1981) Purification and properties of alanopine dehydrogenase from the adductor muscle of the oysterCrassostrea gigas (Mollusca, Bivalvia). Eur J Biochem 114:615–621
Fort L, Dando PR, Rouzé P, Monneuse M-O, Olomucki A (1982) Immunological comparative studies of octopine dehydrogenase and other “pyruvate reductases” from different species. Comp Biochem Physiol [B] 73:865–871
Gäde G (1980) Biological role of octopine formation in marine molluscs. Marine Biol Lett 1:121–135
Garnier J, Osguthorpe DJ, Robson B (1978) Analysis of the accuracy and implications of simple methods for predicting the secondary structure of globular proteins. J Mol Biol 120:97–120
Go M (1983) Modular structural units, exons, and function in chicken lysozyme. Proc Natl Acad Sci USA 80:1964–1968
Goad WB, Kanehisa MI (1982) Pattern recognition in nucleic acid sequences. I. A general method for finding local homologies and symmetries. Nucleic Acids Res 10:247–263
Goldmann A (1977) Octopine and nopaline dehydrogenases in crown gall tumors. Plant Sci Lett 10:49–58
Goldmann A, Moureaux T, Rouzé P (1981) Antibody against octopine dehydrogenase from crown gall tumor tissue, a tool in studies of plant cell transformation. FEBS Lett 130:213–216
Hol WGJ (1985) The role of the α-helix dipole in protein function and structure. Prog Biophys Mol Biol 45:149–195
Hopp TP, Woods KR (1981) Prediction of protein antigenic determinants from amino acid sequences. Proc Natl Acad Sci USA 78:3824–3828
Jany K-D, Ulmer W, Fröschle M, Pfleiderer G (1984) Complete amino acid sequence of glucose dehydrogenase fromBacillus megaterium. FEBS Lett 165:6–10
Jeffery J, Cederlund E, Jörnvall H, (1984) Sorbitol dehydrogenase. The primary structure of the sheep-liver enzyme. Eur J Biochem 140:7–16
McLachlan AD (1971) Tests for comparing related amino acid sequences. Cytochrome c and cytochrome c551. J Mol Biol 61:409–424
Monneuse-Doublet M-O, Olomucki A, Buc J (1978) Investigation on the kinetic mechanism of octopine dehydrogenase. A regulatory behavior. Eur J Biochem 84:441–448
Nester EW, Gordon MP, Amasino RM, Yanafsky MF (1984) Crown gall: a molecular and physiological analysis. Annu Rev Plant Physiol 35:387–413
Ohno S, Epplen JT (1983) The primitive code and repeats of base oligomers as the primordial protein-encoding sequence. Proc Natl Acad Sci USA 80:3391–3395
Olomucki A (1981) Structure and function of octopine dehydrogenase ofPecten maximus (great scallop). Biochem Soc Trans 9:278–279
Otto J, Argos P, Rossmann MG (1980) Prediction of secondary structural elements in glycerol-3-phosphate dehydrogenase by comparison with other dehydrogenases. Eur J Biochem 109:325–330
Ralston R, Bishop JM (1983) The protein products of the myc and myb oncogenes and adenovirus Eia are structurally related. Nature 306:803–806
Rose GD, Geselowitz AR, Lesser GJ, Lee RH, Zehfus MH (1985) Hydrophobicity of amino acid residues in globular proteins. Science 229:834–838
Rossman MG, Garavito RM, Eventoff W (1977) Conformational adaptations among dehydrogenases. In Sund H (ed) Pyridine-nucleotide-dependent dehydrogenases, FEBS symposium Konstanz. Walter de Gruyter, Berlin, pp 3–30
Schell J (1978) The use of the Ti-plasmid as a vector for the introduction of foreign DNA into plants. In: Ridé M (ed) Proceedings of the fourth international conference on plant pathog bact, INRA, Angers, pp 115–126
Schröder J, Schröder G, Huisman H, Schilperoort RA, Schell J (1981) The mRNA for lysopine dehydrogenase in plant tumor cells is complementary to a Ti plasmid fragment. FEBS Lett 129:166–168
Staden R (1982) An interactive graphics program for comparing and aligning nucleic acid and amino acid sequences. Nucleic Acids Res 10:2951–2961
Tempé J (1983) Chemistry and biochemistry of open-chain imino acids. In: Weinstein B (ed) Chemistry of amino acids, peptides and proteins: chemistry and biochemistry, vol 7. Marcel Dekker, New York, pp 113–204
Tepfer D (1983) The biology of the genetic transformation of higher plants byAgrobacterium tumefaciens. In Pühler A (ed) Molecular genetics of the bacteria plant interaction. Springer-Verlag, New York, pp 248–258
Thoai Nv, Huc C, Dang Ba Pho, Olomucki A (1969) Octopine déshydrogénase. Purification et propriétés catalytiques. Biochim Biophys Acta 191:46–57
Thornton JM, Sibanda BL (1983) Amino and carboxy-terminal regions in globular proteins. J Mol Biol 167:443–460
Walker JE, Sarate M, Runswick MJ, Gay NJ (1982) Distantly related sequences in the α- and β-subunits of ATP synthase, myosin, kinases and other ATP-requiring enzymes and a common nucleotide binding fold. EMBO J 1:945–951
Ward ER, Barnes WM (1983) Marine molluscan genomes contain sequences homologous to the octopine synthase gene ofAgrobacterium tumefaciens. Biol Bull 165:498
Welling GW, Weijer WJ, von der Zee R, Welling-Wester S (1985) Prediction of sequential antigenic regions in proteins. FEBS Lett 188:215–218
White FF, Garfinkel DJ, Huffman GA, Gordon MP, Nester EW (1983) Sequences homologous toAgrobacterium rhizogenes T-DNA in the genomes of uninfected plants. Nature 301:348–350
Wierenga RK, Hol WGJ (1983) Predicted nucleotide-binding properties of p21 protein and its cancer-associated variant. Nature 302:842–844
Wierenga RK, Terpstra P, Hol WGJ (1986) Prediction of the occurrence of the ADP-binding βαβ-fold in proteins, using an amino acid sequence fingerprint. J Mol Biol 187:101–107
Willmitzer L, Dhaese P, Schreier PH, Schmalenbach W, Van Montagu M, Schell J (1983) Size, location and polarity of T-DNA-encoded transcripts in nopaline crown gall tumors; common transcripts in octopine and nopaline tumors. Cell 32:1045–1056
Author information
Authors and Affiliations
Rights and permissions
About this article
Cite this article
Monneuse, M.O., Rouzé, P. Sequence comparisons betweenAgrobacterium tumefaciens T-DNA-encoded octopine and nopaline dehydrogenases and other nucleotide-requiring enzymes: Structural and evolutionary implications. J Mol Evol 25, 46–57 (1987). https://doi.org/10.1007/BF02100040
Received:
Revised:
Issue Date:
DOI: https://doi.org/10.1007/BF02100040