Summary
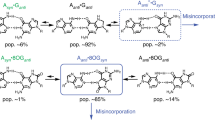
An inherent feature of double-stranded DNA is the possible replacement of any base pair by another one upon replication. A replication-dependent substitution mutation of a matched base pair requires the temporary formation of a mismatched base pair (mispair). A functionally complementary pair of mispairs is ascribed to each of the four types of substitution mutations. Provided that all types of mispairs can be formed, a dynamic biological equilibrium between the four matched base pairs must exist in all DNA, which is directly related to the formation and stability of the corresponding eight mispairs in vivo. Each nucleotide position in a genome can therefore be described as a system of six dynamic equilibria between the four matched base pairs. After a sufficient number of replications, these equilibrium states will express an overall mutation-selection balance for each individual base pair. In a thermodynamic context, the mispairs represent intermediate states on the transformation pathway between the matched base pairs. Catalysts change the stability and probability of formation of intermediate states. Mutagenic proteins are proposed as hypothetical substitution mutation catalysts in vivo. Functionally, they would be capable of recognizing a particular DNA sequence, tautomerizing a nucleotide base thereof, and hence efficiently inducing a specific misincorporation. Phenomenologically such catalysts would accelerate the rates of substitution mutations and provide pathways for directional mutation pressure.
Article PDF
Similar content being viewed by others
Avoid common mistakes on your manuscript.
References
Beutler E, Gelbart T, Han J, Koziol JA, Beutler B (1989) Evolution of the genome and the genetic code: selection at the dinucleotide level by methylation and polyribonucleotide cleavage. Proc Natl Acad Sci USA 86:192–196
Cairns J, Overbaugh J, Miller S (1988) The origin of mutants. Nature 335:142–145
Charczuk R, Tamm C, Suri B, Bickle T (1986) An unusual base pairing between pyrimidine and pyridine nucleotides. Nucleic Acids Res 14:9530
Dawkins R (1982) The extended phenotype. The gene as the unit of selection, ed 2 (1987). Oxford University Press, New York, pp 81–96
Discussions about Cairns et al. (1988/1989) in Nature (1988) 335:112–113, 336:21–22, 336:525–528, in Nature (1989) 337: 119–120, 337:123–124
Dohet C, Wagner R, Radman M (1985) Repair of defined single base-pair mismatches inEscherichia coli. Proc Natl Acad Sci USA 82:503–505
Dover GA (1987) DNA turnover and the molecular clock. J Mol Evol 26:47–58
Eigen M (1971) Selforganization of matter and the evolution of biological macromolecules. Naturwissenschaften 10:465–523
Eigen M (1978) The hypercycle. Part B: the abstract hypercycle. Naturwissenschaften 65:7–41
Eigen M, Schuster P (1977) The hypercycle. Part A: emergence of the hypercycle. Naturwissenschaften 11:541–565
Eigen M, Schuster P (1978) The hypercycle. Part C: the realistic hypercycle. Naturwissenschaften 7:341–369
Fowler RG, Schaaper RM, Glickman BW (1986) Characterization of mutational specificity within the lac-1 gene for a mutD5 mutator strain ofEscherichia coli defective in 3′-5′ exonuclease proofreading activity. J Bacteriol 167:130–137
French DL, Laskov R, Scharff MD (1989) The role of somatic hypermutation in the generation of antibody diversity. Science 244:1152–1157
Goodman MF, Branscomb EW (1986) DNA replication fidelity and base mispairing mutagenesis. In: Kirkwood TBL, Rosenberger RF, Galas DJ (eds) Accuracy in molecular processes. Chapman & Hall, London, New York, pp 191–232
Gutman GA, Hatfield GW (1989) Nonrandom utilization of codon pairs inEscherichia coli. Proc Natl Acad Sci USA 86: 3699–3703
Ikemura T (1985) Codon usage, tRNA content, and rate of synonymous substitution. In: Ohta T, Aoki K (eds) Population genetics and molecular evolution. Springer-Verlag, Berlin, pp 385–406
Jukes T, Bhushan V (1986) Silent nucleotide substitutions and G+C content of some mitochondrial and bacterial genes. J Mol Evol 24:39–44
Kimura M (1983) The neutral theory of molecular evolution. Cambridge University Press, London
Kuchta RD, Benkovic P, Benkovic SJ (1988) Kinetic mechanism whereby DNA polymerase I (Klenow) replicates with high fidelity. Biochemistry 27:6716–6725
Kunkel TA, Bebenek K (1988) Recent studies of the fidelity of DNA synthesis. Biochim Biophys Acta 951:1–15
Ohno S (1988a) Codon preference is but an illusion created by the construction principle of coding sequences. Proc Natl Acad Sci USA 85:4378–4382
Ohno S (1988b) Universal rule for coding sequence construction: TA/CG deficiency-TG/CT excess. Proc Natl Acad Sci USA 85:9630–9634
Ollis DL, Brick P, Hamlin R, Xuong NG, Steitz TA (1985) Structure of large fragment ofEscherichia coli DNA polymerase I complexed with dTMP. Nature 313:762–766
Schaaper RM, Danforth BN, Glickman BW (1986) Mechanisms of spontaneous mutagenesis. An analysis of the spectrum of spontaneous mutation in theEscherichia coli lac-1 gene. J Mol Biol 189:273–284
Schaaper RM, Dunn RL (1987) Spectra of spontaneous mutations inEscherichia coli strains defective in mismatch correction. The nature of in-vivo DNA replication errors. Proc Natl Acad Sci USA 84:6220–6224
Strazewski P (1988) Mispair formation in DNA can involve rare tautomeric forms in the template. Nucleic Acids Res 16: 9377–9398
Strazewski P, Tamm C (1990) Replication experiments with nucleotide base analogues. Angew Chem 102 (in press)
Sueoka N (1988) Directional mutation pressure and neutral molecular evolution. Proc Natl Acad Sci USA 85:2653–2657
Tonegawa S (1988) How does an organism manage during its lifetime to respond to a huge number of different antigens. Angew Chem Int Ed Engl 27:1028–1039
Topal MD, Fresco JR (1976) Complementary base pairing and the origin of substitution mutations. Nature 263:285–289
Vogel H, Zuckerkandl E (1971) Randomness and “thermodynamics” of molecular evolution. In: Schoffeniels E (ed) Biochemical evolution and the origin of life. North-Holland. Amsterdam, pp 352–365
Wolfe KH, Sharp PM, Li W-H (1989) Mutation rates differ among regions of the mammalian genome. Nature 337:283–285
Zckerkandl E (1965) Remarques sur l'évolution des polynucléotides comparée à celle des polypeptides. Bull Soc Chim Biol 47:1729–1730
Zckerkandl E (1987) On the molecular evolutionary clock. J Mol Evol 26:34–46
Zuckerkandl E, Pauling L (1965) Molecules as documents of evolutionary history. J theoret Biol 8:357–366
Author information
Authors and Affiliations
Additional information
Present address until March 31 1990: University Chemical Laboratory, Lensfield Road, Cambridge CB2 1EW, Great Britain
Rights and permissions
About this article
Cite this article
Strazewski, P. The biological equilibrium of base pairs. J Mol Evol 30, 116–124 (1990). https://doi.org/10.1007/BF02099938
Received:
Revised:
Accepted:
Issue Date:
DOI: https://doi.org/10.1007/BF02099938