Summary
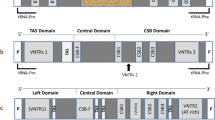
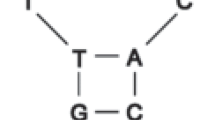
A detailed comparative study of the regions surrounding the origin of replication in vertebrate mitochondrial DNA (mtDNA) has revealed a number of interesting properties. This region, called the D-loop-containing region, can be divided into three domains. The left (L) and right (R) domains, which have a low G content and contain the 5′ and the 3′ D-loop ends, respectively, are highly variable for both base sequence and length. They, however, contain thermodynamically stable secondary structures which include the conserved sequence blocks called CSB-1 and TAS which are associated with the start and stop sites, respectively, for D-loop strand synthesis. We have found that a “mirror symmetry” exists between the CSB-1 and TAS elements, which suggests that they can act as specific recognition sites for regulatory, probably dimeric, proteins. Long, statistically significant repeats are found in the L and R domains.
Between the L and R domains we observed in all mtDNA sequences a region with a higher G content which was apparently free of complex secondary structure. This central domain, well preserved in mammals, contains an open reading frame of variable length in the organisms considered.
The identification of common features well preserved in evolution despite the high primary structural divergence of the D-loop-containing region of vertebrate mtDNA suggests that these properties are of prime importance for the mitochondrial processes that occur in this region and may be useful for singling out the sites on which one should operate experimentally in order to discover functionally important elements.
Article PDF
Similar content being viewed by others
Avoid common mistakes on your manuscript.
References
Anderson S, Bankier AT, Barrell GB, de Bruijn MHL, Coulson AR, Drouin J, Eperon IC, Nierlich DP, Roe BA, Sanger F, Schreier PH, Smith AJH, Staden R, Young IG (1981) Sequence and organization of the human mitochondrial genome. Nature 290:457–464
Ano T, Imanaka T, Aiba S (1986) The copy number ofBacillus plasmid pRBH1 is negatively controlled by RepB protein. Mol Gen Genet 202:416–420
Attardi G (1985) Animal mitochondrial DNA: an extreme example of genetic economy. Int Rev Cytol 93:93–145
Brown GG, Gadaleta G, Pepe G, Saccone C, Sbisà E (1986) Structural conservation and variation in the D-loop containing region of vertebrate mitochondrial DNA. J Mol Biol 192: 503–511
Brown WM, George M Jr, Wilson AC (1979) Rapid evolution of animal mitochondrial DNA. Proc Natl Acad Sci USA 76: 1967–1971
Chang DD, Hausweirth WW, Clayton DA (1985) Replication priming and transcription initiate from precisely the same site in mouse mitochondrial DNA. EMBO J 4:1559–1567
Clayton DA (1982) Replication of animal mitochondrial DNA. Cell 28:693–705
Clayton DA (1984) Transcription of the mammalian mitochondrial genome. Annu Rev Biochem 53:573–594
Dawid IB (1972) Evolution of mitochondrial DNA sequences inXenopus. Dev Biol 29:139–151
De Zamaroczy M, Faugeron-Fonty G, Baldacci G, Goursot R, Bernardi G (1984) The ori sequences of the mitochondrial genome of a wild-type yeast strain: number, location, orientation and structure. Gene 32:439–457
Doda JN, Wright CT, Clayton DA (1981) Elongation of displacement-loop strands in human and mouse mitochondrial DNA is arrested near specific template sequences. Proc Natl Acad Sci USA 78:6116–6120
Ferris S, Brown WM, Davidson WS, Wilson AC (1981) Extensive polymorphism in the mitochondrial DNA of apes. Proc Natl Acad Sci USA 78:6319–6323
Gouy M, Gautier C, Attimonelli M, Lanave C, Di Paola G (1985) ACNUC-a portable retrieval system for nucleic acid sequence data-bases: logical and physical designs and usage. CABIOS 1:167–172
Greenberg BD, Newbold JE, Sugino A (1983) Intraspecific nucleotide sequence variability surrounding the origin of replication in human mitochondrial DNA. Gene 21:33–49
Higgins CF, Ferro-Luzzi Ames G (1982) Regulatory regions of two transport operons under nitrogen control: nucleotide sequences. Proc Natl Acad Sci USA 79:1083–1087
Lacatena RM, Cesareni G (1981) Base pairing of RNA I with its complementary sequence in the primer precursor inhibits Co1 E1 replication. Nature 294:623–626
Lanave C, Preparata G, Saccone C, Serio G (1984) A new method for calculating evolutionary substitution rates. J Mol Evol 20:86–93
Lanave C, Preparata G, Saccone C (1985) Mammalian genes as molecular clocksm J Mol Evol 21:346–350
Lewin B (1983) Genes. Wiley J and Sons (eds) New York
Masukata H, Tomizawa J (1984) Effects of point mutations on formation and structure of the RNA primer for Co1 E1 DNA replication. Cell 36:513–522
Ogata RT, Gilbert W (1979) DNA-binding site of lac repressor probed by dimethylsulfate methylation of lac operator. J Mol Biol 132:709–728
Ovchinnikov Yu A, Monastyrskaya GS, Gubanov VV, Guryer SO, Salomatina IS, Shuvaeva TM, Lipkin VM, Sverdlov ED (1982) The primary structure ofE. coli RNA polymerase. Nucleotide sequence of the rpoC gene and amino acid sequence of the B′ subunit. Nucleic Acids Res 10:4035–4044
Pepe G, Holtrop M, Gadaleta G, Kroon AM, Cantatore P, Gallerani R, de Benedetto C, Quagliariello C, Sbisà E, Saccone C (1983) Non-random patterns of nucleotide substitutions and codon strategy in the mammalian mitochondrial genes coding for identified and unidentified reading frames. Biochem Int 6:553–563
Pritchard AE, Seilhamer JJ, Cummings DJ (1986)Paramecium mitochondrial DNA sequences and RNA transcripts for cytochrome oxidase subunit I, URF1, and three ORFs adjacent to the replication origin. Gene 44:243–253
Saccone C, Attimonelli M, Sbisà E (1985) Primary and higher order structural analysis of animal mitochondrial DNA. In: Quagliariello E, Slater EC, Palmieri F, Saccone C, Kroon AM (eds) Achievements and perspectives of mitochondrial research, vol II. Biogenesis. Elsevier BV (Biomedical Division), Amsterdam, p 37
Shuey DJ, Attardi G (1985) Characterization of an RNA polymerase activity from HeLa cell mitochondria, which initiates transcription at the heavy strand rRNA promoter and the light strand promoter in human mitochondrial DNA. J Biol Chem 260:1952–1958
Smith TF, Waterman MS, Burks C (1985) The statistical distribution of nucleic acid similarities. Nucleic Acids Res 13: 645–655
Stenberg RM, Thomsen RD, Stinski MF (1984) Structural analysis of the major immediate early gene of human cytomegalovirus. J Virology 49:190–198
Tomizawa J, Itoh T (1981) Plasmid Col E1 incompatibility determined by interaction of RNA I with primer transcript. Proc Natl Acad Sci USA 78:6096–6100
Tomizawa J, Som T (1984) Control of Col E1 plasmid replication: enhancement of binding of RNA I to the primer transcript by the Rom protein. Cell 38:871–878
Walberg MW, Clayton DA (1981) Sequence and properties of the human KB cell and mouse L cell D-loop regions of mitochondrial DNA. Nucleic Acids Res 9:5411–5421
Walberg MW, Clayton DA (1983)In vitro transcription of human mitochondrial DNA. Identification of specific light strand transcripts from the displacement loop region. J Biol Chem 258:1268–1275
Wong TW, Clayton DA (1985)In vitro replication of human mitochondrial DNA: accurate initiation at the origin of lightstrand synthesis. Cell 42:951–958
Author information
Authors and Affiliations
Rights and permissions
About this article
Cite this article
Saccone, C., Attimonelli, M. & Sbisà, E. Structural elements highly preserved during the evolution of the D-loop-containing region in vertebrate mitochondrial DNA. J Mol Evol 26, 205–211 (1987). https://doi.org/10.1007/BF02099853
Issue Date:
DOI: https://doi.org/10.1007/BF02099853